Abstract
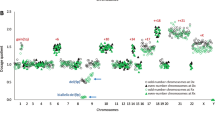
PAX5, a master regulator of B-cell development, was recently shown to be involved in several leukemia-associated rearrangements, which result in fusion genes encoding chimeric proteins that antagonize PAX5 transcriptional activity. In a population-based fluorescence in situ hybridization screening study of 446 childhood acute lymphoblastic leukemia (ALL) patients, we now show that PAX5 rearrangements occur at an incidence of about 2.5% of B-cell precursor ALL. Identification of several novel PAX5 partner genes, including POM121, BRD1, DACH1, HIPK1 and JAK2 brings the number of distinct PAX5 in-frame fusions to at least 12. Our data show that these not only comprise transcription factors but also structural proteins and genes involved in signal transduction, which at least in part have not been implicated in tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Busslinger M . Transcriptional control of early B cell development. Annu Rev Immunol 2004; 22: 55–79.
Mikkola I, Heavey B, Horcher M, Busslinger M . Reversion of B cell commitment upon loss of Pax5 expression. Science 2002; 297: 110–113.
Nutt SL, Eberhard D, Horcher M, Rolink AG, Busslinger M . Pax5 determines the identity of B cells from the beginning to the end of B-lymphopoiesis. Int Rev Immunol 2001; 20: 65–82.
Nutt SL, Heavey B, Rolink AG, Busslinger M . Commitment to the B-lymphoid lineage depends on the transcription factor Pax5. Nature 1999; 401: 556–562.
Delogu A, Schebesta A, Sun Q, Aschenbrenner K, Perlot T, Busslinger M . Gene repression by Pax5 in B cells is essential for blood cell homeostasis and is reversed in plasma cells. Immunity 2006; 24: 269–281.
Schebesta M, Heavey B, Busslinger M . Transcriptional control of B-cell development. Curr Opin Immunol 2002; 14: 216–223.
Busslinger M, Klix N, Pfeffer P, Graninger PG, Kozmik Z . Deregulation of PAX-5 by translocation of the Emu enhancer of the IgH locus adjacent to two alternative PAX-5 promoters in a diffuse large-cell lymphoma. Proc Natl Acad Sci USA 1996; 93: 6129–6134.
Iida S, Rao PH, Nallasivam P, Hibshoosh H, Butler M, Louie DC et al. The t(9;14)(p13;q32) chromosomal translocation associated with lymphoplasmacytoid lymphoma involves the PAX-5 gene. Blood 1996; 88: 4110–4117.
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 2007; 446: 758–764.
Cazzaniga G, Daniotti M, Tosi S, Giudici G, Aloisi A, Pogliani E et al. The paired box domain gene PAX5 is fused to ETV6/TEL in an acute lymphoblastic leukemia case. Cancer Res 2001; 61: 4666–4670.
Strehl S, Konig M, Dworzak MN, Kalwak K, Haas OA . PAX5/ETV6 fusion defines cytogenetic entity dic(9;12)(p13;p13). Leukemia 2003; 17: 1121–1123.
Bousquet M, Broccardo C, Quelen C, Meggetto F, Kuhlein E, Delsol G et al. A novel PAX5-ELN fusion protein identified in B-cell acute lymphoblastic leukemia acts as a dominant negative on wild-type PAX5. Blood 2007; 109: 3417–3423.
Nebral K, Konig M, Harder L, Siebert R, Haas OA, Strehl S . Identification of PML as novel PAX5 fusion partner in childhood acute lymphoblastic leukaemia. Br J Haematol 2007; 139: 269–274.
Kawamata N, Ogawa S, Zimmermann M, Sanada M, Hemminki K, Yamatomo G et al. Rearrangement and deletion of the PAX5 gene in pediatric acute B-cell lineage lymphoblastic leukemia. ASH Annu Meet Abstr 2007; 110: 981.
Pieters R, Schrappe M, De Lorenzo P, Hann I, De Rossi G, Felice M et al. A treatment protocol for infants younger than 1 year with acute lymphoblastic leukaemia (Interfant-99): an observational study and a multicentre randomised trial. Lancet 2007; 370: 240–250.
Konig M, Reichel M, Marschalek R, Haas OA, Strehl S . A highly specific and sensitive fluorescence in situ hybridization assay for the detection of t(4;11)(q21;q23) and concurrent submicroscopic deletions in acute leukaemias. Br J Haematol 2002; 116: 758–764.
Ayres JA, Shum L, Akarsu AN, Dashner R, Takahashi K, Ikura T et al. DACH: genomic characterization, evaluation as a candidate for postaxial polydactyly type A2, and developmental expression pattern of the mouse homologue. Genomics 2001; 77: 18–26.
Baker SJ, Rane SG, Reddy EP . Hematopoietic cytokine receptor signaling. Oncogene 2007; 26: 6724–6737.
Murray PJ . The JAK-STAT signaling pathway: input and output integration. J Immunol 2007; 178: 2623–2629.
Levine RL, Pardanani A, Tefferi A, Gilliland DG . Role of JAK2 in the pathogenesis and therapy of myeloproliferative disorders. Nat Rev Cancer 2007; 7: 673–683.
Ihle JN, Gilliland DG . Jak2: normal function and role in hematopoietic disorders. Curr Opin Genet Dev 2007; 17: 8–14.
Poitras JL, Cin PD, Aster JC, Deangelo DJ, Morton CC . Novel SSBP2-JAK2 fusion gene resulting from a t(5;9)(q14.1;p24.1) in pre-B acute lymphocytic leukemia. Genes Chromosomes Cancer 2008; 47: 884–889.
Schwaller J, Frantsve J, Aster J, Williams IR, Tomasson MH, Ross TS et al. Transformation of hematopoietic cell lines to growth-factor independence and induction of a fatal myelo- and lymphoproliferative disease in mice by retrovirally transduced TEL/JAK2 fusion genes. EMBO J 1998; 17: 5321–5333.
Kovac CR, Emelyanov A, Singh M, Ashouian N, Birshtein BK . BSAP (Pax5)-importin alpha 1 (Rch1) interaction identifies a nuclear localization sequence. J Biol Chem 2000; 275: 16752–16757.
McCullagh P, Chaplin T, Meerabux J, Grenzelias D, Lillington D, Poulsom R et al. The cloning, mapping and expression of a novel gene, BRL, related to the AF10 leukaemia gene. Oncogene 1999; 18: 7442–7452.
Chaplin T, Bernard O, Beverloo HB, Saha V, Hagemeijer A, Berger R et al. The t(10;11) translocation in acute myeloid leukemia (M5) consistently fuses the leucine zipper motif of AF10 onto the HRX gene. Blood 1995; 86: 2073–2076.
Prasad R, Leshkowitz D, Gu Y, Alder H, Nakamura T, Saito H et al. Leucine-zipper dimerization motif encoded by the AF17 gene fused to ALL-1 (MLL) in acute leukemia. Proc Natl Acad Sci USA 1994; 91: 8107–8111.
Ruthenburg AJ, Li H, Patel DJ, Allis CD . Multivalent engagement of chromatin modifications by linked binding modules. Nat Rev Mol Cell Biol 2007; 8: 983–994.
Taverna SD, Li H, Ruthenburg AJ, Allis CD, Patel DJ . How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat Struct Mol Biol 2007; 14: 1025–1040.
Arai S, Matsushita A, Du K, Yagi K, Okazaki Y, Kurokawa R . Novel homeodomain-interacting protein kinase family member, HIPK4, phosphorylates human p53 at serine 9. FEBS Lett 2007; 581: 5649–5657.
Kim YH, Choi CY, Lee SJ, Conti MA, Kim Y . Homeodomain-interacting protein kinases, a novel family of co-repressors for homeodomain transcription factors. J Biol Chem 1998; 273: 25875–25879.
Aikawa Y, Nguyen LA, Isono K, Takakura N, Tagata Y, Schmitz ML et al. Roles of HIPK1 and HIPK2 in AML1- and p300-dependent transcription, hematopoiesis and blood vessel formation. EMBO J 2006; 25: 3955–3965.
Ecsedy JA, Michaelson JS, Leder P . Homeodomain-interacting protein kinase 1 modulates Daxx localization, phosphorylation, and transcriptional activity. Mol Cell Biol 2003; 23: 950–960.
Kondo S, Lu Y, Debbas M, Lin AW, Sarosi I, Itie A et al. Characterization of cells and gene-targeted mice deficient for the p53-binding kinase homeodomain-interacting protein kinase 1 (HIPK1). Proc Natl Acad Sci USA 2003; 100: 5431–5436.
Li X, Zhang R, Luo D, Park SJ, Wang Q, Kim Y et al. Tumor necrosis factor alpha-induced desumoylation and cytoplasmic translocation of homeodomain-interacting protein kinase 1 are critical for apoptosis signal-regulating kinase 1-JNK/p38 activation. J Biol Chem 2005; 280: 15061–15070.
Rochat-Steiner V, Becker K, Micheau O, Schneider P, Burns K, Tschopp J . FIST/HIPK3: a Fas/FADD-interacting serine/threonine kinase that induces FADD phosphorylation and inhibits fas-mediated Jun NH(2)-terminal kinase activation. J Exp Med 2000; 192: 1165–1174.
Chen R, Amoui M, Zhang Z, Mardon G . Dachshund and eyes absent proteins form a complex and function synergistically to induce ectopic eye development in Drosophila. Cell 1997; 91: 893–903.
Davis RJ, Shen W, Heanue TA, Mardon G . Mouse Dach, a homologue of Drosophila dachshund, is expressed in the developing retina, brain and limbs. Dev Genes Evol 1999; 209: 526–536.
Kozmik Z, Pfeffer P, Kralova J, Paces J, Paces V, Kalousova A et al. Molecular cloning and expression of the human and mouse homologues of the Drosophila dachshund gene. Dev Genes Evol 1999; 209: 537–545.
Wawersik S, Maas RL . Vertebrate eye development as modeled in Drosophila. Hum Mol Genet 2000; 9: 917–925.
Hanson IM . Mammalian homologues of the Drosophila eye specification genes. Semin Cell Dev Biol 2001; 12: 475–484.
Sunde JS, Donninger H, Wu K, Johnson ME, Pestell RG, Rose GS et al. Expression profiling identifies altered expression of genes that contribute to the inhibition of transforming growth factor-beta signaling in ovarian cancer. Cancer Res 2006; 66: 8404–8412.
Wu K, Yang Y, Wang C, Davoli MA, D'Amico M, Li A et al. DACH1 inhibits transforming growth factor-beta signaling through binding Smad4. J Biol Chem 2003; 278: 51673–51684.
Wu K, Liu M, Li A, Donninger H, Rao M, Jiao X et al. Cell fate determination factor DACH1 inhibits c-Jun-induced contact-independent growth. Mol Biol Cell 2007; 18: 755–767.
Hallberg E, Wozniak RW, Blobel G . An integral membrane protein of the pore membrane domain of the nuclear envelope contains a nucleoporin-like region. J Cell Biol 1993; 122: 513–521.
Gerace L, Ottaviano Y, Kondor-Koch C . Identification of a major polypeptide of the nuclear pore complex. J Cell Biol 1982; 95: 826–837.
Antonin W, Franz C, Haselmann U, Antony C, Mattaj IW . The integral membrane nucleoporin pom121 functionally links nuclear pore complex assembly and nuclear envelope formation. Mol Cell 2005; 17: 83–92.
Stavru F, Nautrup-Pedersen G, Cordes VC, Gorlich D . Nuclear pore complex assembly and maintenance in POM121- and gp210-deficient cells. J Cell Biol 2006; 173: 477–483.
Lusk CP, Blobel G, King MC . Highway to the inner nuclear membrane: rules for the road. Nat Rev Mol Cell Biol 2007; 8: 414–420.
Romana SP, Radford-Weiss I, Ben Abdelali R, Schluth C, Petit A, Dastugue N et al. NUP98 rearrangements in hematopoietic malignancies: a study of the Groupe Francophone de Cytogenetique Hematologique. Leukemia 2006; 20: 696–706.
Slape C, Aplan PD . The role of NUP98 gene fusions in hematologic malignancy. Leuk Lymphoma 2004; 45: 1341–1350.
Graux C, Cools J, Melotte C, Quentmeier H, Ferrando A, Levine R et al. Fusion of NUP214 to ABL1 on amplified episomes in T-cell acute lymphoblastic leukemia. Nat Genet 2004; 36: 1084–1089.
Soekarman D, von Lindern M, Daenen S, de Jong B, Fonatsch C, Heinze B et al. The translocation (6;9) (p23;q34) shows consistent rearrangement of two genes and defines a myeloproliferative disorder with specific clinical features. Blood 1992; 79: 2990–2997.
von Lindern M, Breems D, van Baal S, Adriaansen H, Grosveld G . Characterization of the translocation breakpoint sequences of two DEK-CAN fusion genes present in t(6;9) acute myeloid leukemia and a SET-CAN fusion gene found in a case of acute undifferentiated leukemia. Genes Chromosomes Cancer 1992; 5: 227–234.
Cobaleda C, Schebesta A, Delogu A, Busslinger M . Pax5: the guardian of B cell identity and function. Nat Immunol 2007; 8: 463–470.
Fazio G, Palmi C, Rolink A, Biondi A, Cazzaniga G . PAX5/TEL acts as a transcriptional repressor causing down-modulation of CD19, enhances migration to CXCL12, and confers survival advantage in pre-BI cells. Cancer Res 2008; 68: 181–189.
Acknowledgements
This study was supported by a grant of the Austrian Ministry of Science and Research (GEN-AU II, GZ 200.136/1-VI/1/2005) (to SS) and the St Anna Kinderkrebsforschung. We thank Meinrad Busslinger (IMP, Vienna, Austria) for kindly providing the PAX5 cosmid probes and Tilman Johannes (MetaSystems, Altlussheim, Germany) for assistance with the Metafer4-Metacyte system. We thank all those people, who perform the routine diagnostic work-up and consistently provide the basis for our research. Further, we are indebted to Dasa Janousek for the efficient clinical data management and analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu)
Supplementary information
Rights and permissions
About this article
Cite this article
Nebral, K., Denk, D., Attarbaschi, A. et al. Incidence and diversity of PAX5 fusion genes in childhood acute lymphoblastic leukemia. Leukemia 23, 134–143 (2009). https://doi.org/10.1038/leu.2008.306
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2008.306
Keywords
This article is cited by
-
International Consensus Classification of acute lymphoblastic leukemia/lymphoma
Virchows Archiv (2023)
-
Advances in germline predisposition to acute leukaemias and myeloid neoplasms
Nature Reviews Cancer (2021)
-
Functional analysis of a novel fusion protein PAX5-KIDINS220 identified in a pediatric Ph-like ALL patient
International Journal of Hematology (2020)
-
Inhibition of inflammatory signaling in Pax5 mutant cells mitigates B-cell leukemogenesis
Scientific Reports (2020)
-
Implementation of RNA sequencing and array CGH in the diagnostic workflow of the AIEOP-BFM ALL 2017 trial on acute lymphoblastic leukemia
Annals of Hematology (2020)