Abstract
Despite the risk of transmitting HIV-1, mothers in resource-poor areas are encouraged to breastfeed their infants because of beneficial immunologic and nutritional factors in milk. Interestingly, in the absence of antiretroviral prophylaxis, the overwhelming majority of HIV-1-exposed, breastfeeding infants are naturally protected from infection. To understand the role of HIV-1 envelope (Env)-specific antibodies in breast milk in natural protection against infant virus transmission, we produced 19 HIV-1 Env-specific monoclonal antibodies (mAbs) isolated from colostrum B cells of HIV-1-infected mothers and investigated their specificity, evolution, and anti-HIV-1 functions. Despite the previously reported genetic compartmentalization and gp120-specific bias of colostrum HIV Env-specific B cells, the colostrum Env-specific mAbs described here demonstrated a broad range of gp120 epitope specificities and functions, including inhibition of epithelial cell binding and dendritic cell-mediated virus transfer, neutralization, and antibody-dependent cellular cytotoxicity. We also identified divergent patterns of colostrum Env-specific B-cell lineage evolution with respect to crossreactivity to gastrointestinal commensal bacteria, indicating that commensal bacterial antigens play a role in shaping the local breast milk immunoglobulin G (IgG) repertoire. Maternal vaccine strategies to specifically target this breast milk B-cell population may be necessary to achieve safe breastfeeding for all HIV-1-exposed infants.
Similar content being viewed by others
INTRODUCTION
Mother-to-child transmission accounts for ∼260,000 new HIV-1 infections annually,1 with a majority occurring in sub-Saharan Africa. One third to one half of infant infections occur postpartum.2, 3, 4 With up to 1 liter of virus-containing breast milk ingested daily for up to 2 years of life by breastfeeding infants born to HIV-1-infected mothers, it is remarkable that the HIV-1 transmission rate via breastfeeding is <10%, even in the absence of maternal antiretroviral treatment.5 This relatively low risk of infection in the face of chronic mucosal HIV-1 exposure of the infant warrants the study of naturally protective immune factors in breast milk.
Breast milk is known to provide protection against a multitude of neonatal infections, and also provides homeostasis between the infant and its gastrointestinal microbiota.6, 7, 8 Breast milk immunoglobulins A and G (IgA and IgG), as well as a number of innate antimicrobial factors, likely all contribute to infant antimicrobial protection against neonatal pathogens.9 HIV-1 exposure in the infant mucosa occurs in the presence of maternal breast milk HIV-1 envelope (Env)-specific antibodies that have been shown to have neutralizing and antibody-dependent cellular cytotoxicity (ADCC) activity.10, 11 It is possible that these antibodies play a role in protection against infant HIV-1 acquisition via effector functions in the breast milk compartment or at the infant mucosal barrier.
We previously reported that HIV-1 Env-specific antibodies isolated from colostrum of HIV-1-infected, lactating women are exclusively IgG1 isotype, have distinct variable immunoglobulin gene usage from the HIV-1 Env-specific B cells in peripheral blood, and are predominantly specific for the gp120 portion of the HIV-1 Env.12 As the anti-HIV-1 functions of these compartmentalized, potentially protective mucosal gp120-specific antibodies have not been well defined, we sought to provide insight into the evolution and antiviral functions of the HIV-1 Env-specific antibody repertoire of breast milk B cells. Defining the antiviral functions and protective role of milk antibodies directed against HIV-1 would guide the development of immunologic interventions to make breastfeeding safe for all infants in areas of high HIV-1 prevalence.
RESULTS
Selection of colostrum Env-specific IgG1 mAbs
In this study, we aimed to characterize the epitope specificity, function, and evolution of colostrum-derived Env-specific IgG1 antibodies. Using our initial panel of 39 HIV-1 Env-specific IgG monoclonal antibodies (mAbs) previously isolated from colostrum B cells of 17 HIV-1-infected Malawian women,12 we selected a panel of Env-specific colostrum mAbs for functional characterization using the Env-binding properties of the mAbs determined after small-scale mAb production by transient transfection. Four criteria were applied to select the panel of mAbs for large-scale production and in-depth study: (i) robust binding to the clade C 1086gp140 (half-maximal effective concentration (EC50) <0.05 μg ml−1; Table 1); (ii) cross-clade Env gp120 binding (bound ≥3 of 4 Env proteins: Consensus (ConS), Clade A (A244), B (MN), and C (1086) gp120 Envs13); (iii) part of an isolated clonal B-cell lineage,4 included to study lineage evolution and affinity maturation; and (iv) gp41-specific mAbs with strong binding to C.1086gp140 (EC50 <0.03), included for functional comparison with the predominant gp120-specific colostrum mAbs. We also selected four blood Env-specific mAbs isolated from two of the subjects from whom the majority of the colostrum mAbs were isolated (CH9105 and CH0404) for functional comparisons.
This selected panel of 19 colostrum Env-specific mAbs was isolated from a total of 6 HIV-1-infected, lactating Malawian women from the CHAVI009 cohort12 (Table 2), one of whom transmitted the virus postnatally to her infant (CH0404). The breast milk viral load from the nontransmitting colostrum B-cell donors was below the limit of quantification, <240 copies per ml,14 and the median peripheral CD4+ T-cell count was 324, whereas the single transmitting colostrum B-cell donor (CH0404) had high viral load (29,425 copies per ml) and low peripheral CD4+ T-cell count (80 cells per ml). Of the selected 15 gp120-specific and 4 gp41-specific colostrum mAbs, a high percentage (52.6%) used the immunoglobulin heavy-chain variable (VH) gene 1-69 that is common among HIV-1 Env-specific mAbs15, 16 and is used at significantly higher rates in colostrum Env-specific mAbs compared with those in blood12 (Table 1). As broadly HIV-1-neutralizing antibodies (bnAbs) frequently have a high VH mutation rate and long complementarity determining region 3 (CDR3) regions,17 we examined these characteristics in our panel of Env-specific colostrum mAbs. The median VH and light-chain variable gene (VK/VL) amino-acid mutation rate of the colostrum Env-specific mAb panel was 9% (range: 3–13%) and 5% (range: 1–8%), respectively. The median VH CDR3 length of the colostrum Env-specific mAb panel was 15 amino acids (range: 12–29 amino acids), with three colostrum gp120-specific mAbs having a CDR3 length >20 amino acids (DH276=22; DH284=23; and DH388=29 amino acids). The median VK/VL CDR3 length of the colostrum Env-specific mAbs was 9 amino acids (range: 8–11 amino acids). Although the selected colostrum mAbs bound robustly and broadly to HIV-1 Env antigens, they did not demonstrate unusually high mutation frequency and only a minority of colostrum Env-specific mAbs had long CDR3 lengths, and thus did not commonly display characteristics of bnAbs.17
Diverse epitope specificities of HIV-1 gp120-specific colostrum mAbs
Next, we sought to determine the epitope specificities of the gp120-specific colostrum mAbs. Using a combination of binding and blocking competition enzyme-linked immunosorbent assays (ELISAs), we defined the specificities of 12/15 of the colostrum gp120-specific mAbs (Figure 1a). Of these, 27% (4/15) were specific for constant regions 1 and 2 (C1/C2),18 defined by blocking the Env binding of mAb A32.19 However, these C1/C2-specific mAbs did not block galactosyl ceramide (Galcer) liposome binding as previously described for some A32-like C1/C2-specific mAbs20 (Supplementary Figure S1). Of the colostrum gp120-specific mAbs, 20% (3/15) were specific to the third variable loop (V3), binding to both linear peptides and scaffolded V3 proteins21 (Supplementary Table S1). Another 20% (3/15) of the colostrum gp120-specific mAbs blocked the binding of soluble CD4 to Env, and are therefore presumably specific for the CD4-binding site. Interestingly, DH388 mAb binding was dependent on an N-linked Asparagine at amino-acid position 334 in the C3 region of the gp120 Env (Figure 1b), a glycosylation site that is often targeted by antibodies in infected subjects who later develop neutralization breadth.22 DH388 glycan-binding mAb also had the longest VH CDR3 length (29 amino acids) of the panel of gp120-specific colostrum mAbs, yet had a rather typical VH mutation rate (8%). Another antibody, DH285, bound specifically to the V1/V2 loop of the HIV-1 Env, a specificity of antibody responses that predicted reduced risk of infection in the RV144 Thai vaccine trial.23 Finally, 20% (3/15) of colostrum mAbs had gp120-specificity unable to be defined by ELISA.
Fine specificity of isolated colostrum anti-gp120 immunoglobulin G (IgG) monoclonal antibodies (mAbs). (a) V3 and V1/V2 specificity was determined by direct V3 peptide and scaffolded V1/V2 protein-binding enzyme-linked immunosorbent assay (ELISA). CD4-blocking mAbs were defined by envelope (Env) soluble CD4 (sCD4)-blocking ELISA. C1/C2 specificity was determined by mAb A32-blocking ELISA. (b) The mAb DH388 Env binding was dependent on N334 glycan for binding by ELISA using CH505gp120 and a mutant CH505gp120 that has an alanine substitution at position 334. DH388 binding was not dependent on the glycan located at amino acid position 332. OD, optical density.
Colostrum gp120-specific mAb heterologous and autologous HIV-1 neutralization potency
The ability of locally produced colostrum IgG1 mAbs to neutralize heterologous tier 1 and tier 2 HIV-1 and simian/human immunodeficiency virus (SHIV) variants was investigated in the TZM-bl assay. Whole breast milk neutralization potency against the tier 1 clade C variant MW965 was determined for each subject14 (Table 2). This response did not appear to correlate with the isolation of neutralizing colostrum mAbs. Of the colostrum gp120-specific mAbs, 53% (8/15) neutralized the tier 1 clade C virus, MW965 (Supplementary Figure S2B) at a concentration of <0.01 μg ml−1, well below the range of concentrations of Env-specific IgG (∼1–10 μg ml−1) found in the milk of HIV-infected women.14 Of these, seven neutralized other tier 1 clade B and C SHIVs (Figure 2). Of note, DH378, a CD4-blocking mAb, demonstrated weak neutralization at a concentration of 41 μg ml−1, against one tier 2 SHIV variant (SHIV1157ipd3N4) (Supplementary Figure S2A). Despite its glycan dependence and long CDR3 length, mAb DH388 did not neutralize any of the tested HIV-1 variants. None of the four selected gp41-specific colostrum mAbs neutralized tier 1 or tier 2 variants. In addition to heterologous virus neutralization, we also assessed whether the neutralizing colostrum mAbs isolated from subject CH9105 had autologous neutralization potency against two HIV-1 variants isolated from plasma of that subject at 4 weeks after delivery. Four of five of the neutralizing colostrum antibodies isolated from CH9105 (DH374, DH376, DH377, and DH378) demonstrated low-level neutralization of these two autologous viruses (Table 3 and Supplementary Figure S2 C,D).
The gp120 envelope (Env)-specific colostrum immunoglobulin G (IgG) monoclonal antibodies (mAbs) isolated from nontransmitting subjects showed neutralization activity against tier 1 HIV-1/SHIV variants in TZM-bl cells. The inhibitory concentration 50% (IC50) was determined against a panel of tier 1 and 2 HIV-1 and simian/human immunodeficiency virus (SHIV) variants in TZM-bl cells using an initial concentration of 25 μg ml−1. When assayed at an initial concentration of 50 μg ml−1, DH378, a CD4-blocking mAb, neutralized the tier 2 SHIV1157ipd3N4 variant (IC50=41 μg ml−1). All mAbs were screened against tier 1 C.MW965 and tier 2 C.1086. The mAbs with neutralizing activity or broadly HIV-1-neutralizing antibody (BnAb) properties (DH388) were then screened against a wider panel of tier 1 and tier 2 heterologous viruses.
Env-specific colostrum mAbs can inhibit binding of HIV-1 virions to GI epithelial cells
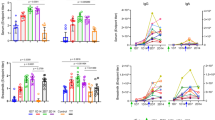
Preventing HIV-1 infectious virions from binding to and crossing the gastrointestinal (GI) tract epithelial cell layer in infants is a potential target for blocking establishment of HIV-1 infection via breastfeeding. Therefore, we sought to determine whether the colostrum Env-specific mAbs can impede this initial step of mucosal infection. Colostrum Env-specific mAbs were tested for their ability to inhibit binding of a Malawian tier 1 clade C HIV-1 variant, C.MW965, to colonic epithelial cells.24 Of the 19 Env-specific mAbs, 11 (57.8%) inhibited viral attachment to colonic epithelial cells (Figure 3a). Additionally, all eight mAbs that neutralized MW965 also inhibited epithelial cell binding at a concentration of 30 μg ml−1, with mean percent inhibition ranging from 68.7 to 98.3%. In contrast, DH390, a nonneutralizing gp120-specific mAb that mediated this activity, only had marginal inhibition (12.8% mean percent inhibition). Furthermore, two gp41-specific mAbs, DH288 and DH389, from the postnatally transmitting subject CH0404 and nontransmitting subject CH9606 weakly inhibited the binding of HIV-1 MW965 to epithelial cells (23.9 and 10.79% mean percent inhibition, respectively). These findings may indicate that weak inhibition of epithelial cell binding by local, nonneutralizing mAbs was not enough to confer protection from breast milk transmission in the setting of high milk virus load in subject CH0404.
Inhibition of HIV-1 epithelial cell binding and dendritic cell (DC)-mediated trans-infection by envelope (Env)-specific colostrum monoclonal antibodies (mAbs). (a) Of the 15 colostrum Env-specific mAbs, 11 inhibit epithelial cell binding of the clade C HIV-1 variant MW965. Epithelial cell binding inhibition was mediated at a high level by the HIV-1 MW965-neutralizing mAbs (red bars), and at a low level by one nonneutralizing gp120-specific mAb (DH390) and gp41 mAbs (DH288 and DH389). (b) Of the 15 colostrum gp120-specific antibodies, 6 inhibit DC transfer of clade C HIV-1 variant MW965 from mature DCs to CD4+/CCR5+ target cells (TZM-bl). The cutoff for significant response in both assays was determined by the mean response of the negative control (anti-influenza mAb CH65) plus 2 s.d. (dotted line). (c) Maximum antibody-dependent cellular cytotoxicity (ADCC) activity is reported, indicating that 8 of 15 colostrum gp120-specific mAbs mediate ADCC of 1086Cgp120-coated natural killer (NK) cells. All antibodies reported to mediate ADCC of 1086C-coated NK cells were compared with uncoated NK cells as controls (Ctrls) and all ADCC activity against uncoated NK cells was found to be below the cutoff of positivity. For the three assays above, mAbs were considered positive if their mean was above the cutoff. GzB, granzyme B.
Neutralizing gp120-specific colostrum mAbs inhibit HIV-1 transfer from DCs to CD4+ target cells
Another important step for establishment of mucosal infection in the infant GI tract is the transfer of virions from dendritic cells (DCs) to CD4+ T lymphocytes. We therefore sought to determine whether colostrum Env-specific mAbs can inhibit the transfer of HIV-1 virions from DCs to CD4+ target cells. Six of eight (75%) mAbs that neutralized HIV-1 MW965 also inhibited DC transfer to CD4+/CCR5+ target cells (TZM-bl) at a concentration of 30 μg ml−1, with mean percent inhibition ranging from 91.4 to 100% (Figure 3b). Although the majority of the neutralizing antibodies efficiently inhibited DC viral transfer, none of the nonneutralizing gp120 or gp41-specific mAbs inhibited DC-C.MW965 virus transfer above background levels. Thus, the ability of colostrum Env-specific mAbs to inhibit infectious virus transfer by DCs may be restricted to neutralizing mAbs.
C1/C2-specific colostrum mAbs mediate ADCC
Once HIV-1 virions or infected cells cross the mucosal epithelium, there are few mechanisms by which the adaptive immune system may protect from further dissemination of HIV-1. ADCC may be an effective way to locally contain HIV-1 infection once HIV-1 crosses the mucosal barrier,25 as it has been associated with protection against postnatal HIV-1 transmission.11 Vaccine-induced antibodies specific for the C1/C2 region of the gp120 Env have been previously described to mediate ADCC.26 Thus, we tested our colostrum mAbs for ADCC activity against natural killer (NK) cells coated with 1086Cgp120 Env and found that 53% mediated ADCC. In concordance with previous findings, all four of our C1/C2-specific colostrum mAbs mediated ADCC, with three at a high titer: DH280, DH382, and DH383 (16–27% granzyme B (GzB) activity, Figure 3c). Two of three V3-specific colostrum mAbs also mediated ADCC. In addition, two conformationally gp120-specific mAbs mediated ADCC at a high level, one neutralizing mAb (DH276) and one nonneutralizing mAb (DH375). The ADCC mediating mAbs demonstrated peak activity at either 10 or 40 μg ml−1. However, end point concentration for the mAbs with high activity was <0.1 μg ml−1 (Supplementary Table S2). Interestingly, 71% of the nonneutralizing gp120-specific colostrum mAbs mediated ADCC, and thus, the ability to mediate ADCC may be an important role of nonneutralizing Env-specific colostrum mAbs.
A subset of colostrum Env gp120-specific mAbs crossreact with intestinal microbiota antigens
Although it has been established that mammary gland IgA-producing B cells originate in the gut-associated lymphoid tissue,27, 28 the origin of IgG-producing B cells in milk is still uncertain. It has been previously reported that gp41-specific mAbs derived from terminal ileum B cells of acute and chronically HIV-1-infected individuals are frequently crossreactive to intestinal microbiota whole-cell lysate (WCL)29 isolated from human stool samples of HIV-1-uninfected individuals. Therefore, we aimed to determine whether colostrum Env-specific mAbs are similarly crossreactive to commensal GI microbiota. We tested our panel of 19 Env-specific colostrum mAbs and two previously described colostrum Env-specific mAbs10 for reactivity against aerobic and anaerobic intestinal microbiota WCL by western blot analysis (Supplementary Figure S3). None of the gp41-specific colostrum mAbs crossreacted with the bacterial WCL antigens (Supplementary Table S3). However, 76% (13/17) of gp120-specific colostrum mAbs crossreacted with either aerobic or anaerobic intestinal microbiota WCL antigens in western blot. As a subset of gp120-specific mAbs induced by Env vaccination crossreact with intestinal microbiota WCLs,30 we sought to determine whether this crossreactivity was specific to colostrum B cell-derived gp120-specific mAbs. Thus, we selected 18 well-characterized, gp120-specific antibodies isolated from blood of vaccinated or chronically HIV-1-infected individuals with a range of gp120 epitope specificities similar to that of the colostrum mAbs from our panel as well as one blood gp120-specific mAb from the lactating HIV-1-infected subject CH9105 to assess crossreactivity to bacterial WCL. Of the 19 blood gp120-specific mAbs, 9 (47%) were reactive to either aerobic or anaerobic commensal bacteria WCL antigens in western blot analysis, and this was not different than the proportion of colostrum gp120-specific mAbs (13 of 17, 76%) that crossreacted with stool bacteria WCL (P=0.096, Fisher’s exact test; Table 4).
Antibody reactivity to the aerobic and anaerobic commensal bacteria in native nonreduced conformation was then confirmed by surface plasmon resonance (SPR). Of the 13 colostrum gp120-specific mAbs that bound to bacterial WCL in Western blot, 10 were also reactive to bacterial WCL by SPR and 2 others that were negative by western blot were reactive by SPR. In addition, four of the blood gp120-specific antibodies that bound to WCL in western blot were confirmed to bind bacterial antigens by SPR and six additional blood gp120s were also reactive by SPR. The difference in mAbs found to be intestinal microbiota reactive in sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) western blot and SPR may be explained by the reduced vs. native conditions of the WCL proteins in each assay, respectively. Overall, colostrum-derived gp120-specific mAbs (15 of 17) did not have significantly higher rate of intestinal microbiota reactivity than blood-derived gp120-specific mAbs (15 of 18; P=0.662, Fisher’s exact test). Colostrum antibody reactivity to intestinal microbiota did not appear to depend on gp120-epitope specificity or particular mAb genetic characteristics, as mAbs with varying gp120 specificities (V3, CD4 blocking, C1/C2, N334 glycan dependency, and V1/V2) and with long and short CDR3 lengths demonstrated reactivity with bacterial antigens (Table 4). We also explored whether any of the measured antiviral functions were associated with intestinal microbiota crossreactivity and found that there were significantly less crossreactive gp120 colostrum mAbs with the ability to inhibit DC-mediated virus transfer compared with non-crossreactive mAbs (P=0.02, Fisher’s exact test). For a full summary of antibody characteristics and function, refer to Supplementary Table S3.
To determine whether intestinal microbiota WCL crossreactivity could be a product of polyreactive/autoreactive properties of the mAbs, we tested both the colostrum and blood gp120-specific antibodies for reactivity against autoantigens in Luminex AtheNA ANA II and HEp-2 immunofluorescence ANA assays. Only 2 of 15 colostrum-derived and 1 of 15 blood-derived intestinal microbiota crossreactive, gp120-specific mAbs showed reactivity to non-HIV-1 antigens by these assays (Table 4). Thus, there is pervasive crossreactivity of blood and breast milk gp120-specific mAbs with commensal bacteria antigens that does not appear to be due primarily to antibody polyreactivity.31
Crossreactivity of a colostrum-isolated gp120-specific mAb with Escherichia coli Chaperonin 60
One gp120 V3-specific colostrum mAb that strongly crossreacted with commensal bacteria WCL (DH374) by western blot and SPR specifically bound to a single 60 kDa bacterial antigen band on a reduced and nonreduced SDS-PAGE Western blot (Figure 4a). This 60 kDa band was isolated by gel extraction and the antigen was identified by liquid chromatography–tandem mass spectrometry as Chaperonin 60 from multiple bacterial species (Escherichia coli, Bacteriodes thetaiotaomicron, and Eubacterium eligens). The binding of DH374 to E. coli Chaperonin 60 was confirmed by western blot and ELISA (Figure 4b,c) but did not bind by SPR or native gel (Supplementary Figure S4). This could be due to the native conformation of the antigen in solution when measured by SPR vs. the nonnative/reduced conformation of the protein in the SDS-PAGE western blot and ELISA. As there is limited amino-acid homology between the linear Chaperonin 6032 and the linear V3 sequence (Supplementary Figure S5), our findings suggest that the colostrum gp120 V3-specific mAb DH374 may crossreact with a conformational epitope on the monomer of the heptameric protein.
HIV-1 gp120-specific colostrum monoclonal antibodies (mAbs) are crossreactive with commensal bacteria whole-cell lysate (WCL) by western blot, including specific reactivity against Escherichia coli Chaperonin 60. (a) The gp120-specific mAbs were tested for reactivity to bacterial WCL by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) reduced and nonreduced western blot. The mAb DH374 was strongly reactive (++) against a single protein band in the WCL, whereas DH280 was weakly reactive (+) against a number of protein bands, and DH386 was nonreactive (−) to gut microbiota WCL. (b) The specificity of DH374 for E. coli Chaperonin 60 was confirmed by both reduced and nonreduced SDS-PAGE western blot. (c) DH374 binding to Chaperonin 60 was also confirmed by binding enzyme-linked immunosorbent assay (ELISA) to the nonreduced E. coli Chaperonin 60 protein (threefold dilutions, ranging from 100 to 0.01 μg ml−1). DH375, a noncommensal bacteria-reactive colostrum mAb, was used as a negative control. OD, optical density.
Affinity maturation of commensal bacteria crossreactive colostrum Env-specific mAbs
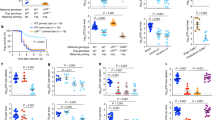
To define the role that commensal bacterial antigens may play in the development of HIV-1 gp120-specific colostrum mAbs, we inferred the heavy- and light-chain unmutated common ancestors12 of two gp120 and commensal bacteria-reactive colostrum mAbs, DH284 and DH285, and recombinantly produced their unmutated common ancestors and intermediates (Figure 5a,b). In both clonal lineages, there was a progressive increase in the affinity for the gp120 (DH284) or V1V2 (DH285) Env antigen with antibody maturation (Figure 5c,d). However, differences emerged in the binding affinity for bacterial WCL within each lineage. For DH284, the binding affinity to commensal bacteria WCL antigens increased over the maturation of this mAb (75 peak response units (RUs) to 350 peak RUs) (Figure 5e) during affinity maturation for gp120. In contrast, the binding strength of the V1V2-specific mAb DH285 to commensal bacteria WCL decreased with maturation (binding affinity peak from 100 peak RUs to 25 peak RUs) (Figure 5f), suggesting that this mAb evolved away from its specificity for bacterial antigens as it increased in affinity for HIV-1 Env. Notably, the DH285 clonal lineage progressively developed tier 1 neutralization potency (C.MW965) during affinity maturation (Figure 5b). These two examples of colostrum gp120-specific mAb evolution indicate that crossreactivity for commensal bacteria can be gained (Figure 5g) and lost (Figure 5h) during HIV-1 gp120-specific affinity maturation, demonstrating that GI bacterial antigens may contribute to shaping the milk B-cell repertoire.
Affinity maturation of colostrum gp120-specific and commensal bacteria crossreactive monoclonal antibodies (mAbs). (a, b) Phylogenetic trees are rooted on the inferred unmutated common ancestor (UCA). Color boxes surrounding lineage antibodies (UCA, green; intermediate, red; mature, blue) indicate the antibody HIV-1 C.1086 envelope (Env) binding strength (EC50), MW965 neutralization potency (IC50), and aerobic commensal bacteria whole-cell lysate (WCL) reactivity (avidity score); color corresponds to graphs below. (c) The mAb DH284 clonal lineage C.1086gp120 antigen reactivity by surface plasmon resonance (SPR). (d) The mAb DH285 clonal lineage C.1086V1/V2 antigen reactivity by SPR. (e,f) The mAb DH284 and mAb DH285 clonal lineage commensal bacteria WCL reactivity by SPR, respectively. (g,h) Schematic of the reactivity of clonal lineages DH284 and DH285 to anaerobic commensal bacteria and HIV-1 Env through lineage maturation.
DISCUSSION
Despite widespread ARV access, breast milk transmission of HIV-1 persists throughout areas of high HIV-1 prevalence. Immunologic strategies that are less dependent on daily adherence to antiretroviral drugs are needed to reduce postnatal HIV-1 transmission while maintaining the benefits of breastfeeding. Natural, innate antiviral factors in breast milk, such as Tenascin-C,33 may contribute to the inherently low rate of HIV-1 transmission via breastfeeding. Furthermore, the potential for immunologic interventions to enhance potentially protective maternal antibody responses in breast milk, the characteristics of the natural milk antibody repertoire needs further exploration. IgG represents <10% of immunoglobulin secreted in breast milk, whereas IgA represents the majority of remaining total milk immunoglobulin.34 Yet, we and others have reported that the concentration of HIV-1 Env-specific IgG in breast milk, which makes up <10% of the total milk IgG,14, 35 exceeds that of the HIV-1 Env-specific IgA levels in milk by 1–2 logs.2, 14, 36 It is important to define the qualities and functions of the locally produced milk HIV-1 Env-specific IgG relative to the concentration in milk to develop strategies for enhancing these potentially protective antibodies that can achieve levels that may be functional in vivo. Our study also sought to provide insight into the ontogeny of mAbs isolated from colostrum B cells that may contribute to blocking mucosal infection of the breastfeeding infant.
We previously reported that the Env-specific B cells in breast milk produced mAbs that had restricted isotype and gene usage compared with those in blood.12 Moreover, compared with the systemic Env-specific B-cell repertoire, the breast milk Env-specific B-cell repertoire was biased toward gp120 specificity. This finding was surprising because of the observed gp41-specific mucosal mAb predominance in acute infection37 and in the gut-associated lymphoid tissue of chronically infected individuals.29 Although most of the milk antibodies characterized in this work were isolated from HIV-1-infected lactating women with a predominance of gp120-specific B cells isolated from milk, the single transmitting mother, CH0404, displayed a predominantly gp41-specific colostrum Env-specific B-cell repertoire (Table 2). However, this subject also had considerable risk factors for postnatal transmission, including a high milk viral load and low peripheral CD4+ T-cell count,38 raising the question of whether the gp41 specificity predominance of the colostrum B-cell repertoire in this subject is related to the high viral load and maternal immunosuppression, yet unrelated to the transmission outcome. Additional studies are necessary to investigate whether a gp41-specific B-cell repertoire in breast milk provides less protection of the infant against HIV-1 infection compared with a gp120-biased B-cell repertoire.
Despite colostrum Env-specific B cells being compartmentalized from those in blood on the basis of gp120 vs. gp41 specificity in this population,12 our panel of colostrum gp120-specific antibodies demonstrate specificity for a wide range of epitopes on the gp120 spike.39 Twenty percent of the gp120-specific colostrum mAbs blocked the CD4-binding site on the Env, and this could be an important function of mucosal antibodies in blocking HIV-1 target cell infection at mucosal surfaces.40 The isolated colostrum mAbs were also specific for the V3 loop that could work in concert with those directed against the CD4-binding site.41 As has been previously reported for weakly neutralizing anti-V3 mAbs,42 our colostrum mAbs also showed gene preference for VH5-51 usage. We also showed that colostrum-derived gp120-specific mAbs can target the V1/V2 loop, a response that has been implicated as protective when elicited systemically by HIV-1 vaccination.13 Interestingly, we additionally identified a colostrum mAb, DH388, specific for a glycan-binding site at position 334. This glycosylation site in the Env constant region 3 (C3) is part of a larger area known as the “high-mannose patch”, thought to be a promising epitope of antiviral activity for bnAbs.43 It has been postulated that a high mannose glycan located at the asparagine of amino acid 332 is involved in viral escape from bnAbs and escaped viruses switch the glycosylation to position 334 at ∼2 years following infection.22 However, this mAb did not show neutralizing activity against any of the HIV-1 variants tested. Finally, despite the high rate of VH 1-69 usage among colostrum Env-specific B cells,12 we found no correlation between usage of this heavy-chain gene and distinct gp120 epitope specificity or function.
This study also characterized the neutralizing potency of colostrum mAbs and their ability to inhibit virion transmission across a mucosal barrier. We found that a high rate of colostrum gp120-specific IgG mAbs could neutralize tier 1 HIV-1 viruses. However, only one colostrum antibody, DH378, weakly neutralized a tier 2 virus variant (SHIV1157ipd3N4). Of potentially greater importance is the observation that four tier 1 virus-neutralizing mAbs isolated from colostrum of the nontransmitting subject CH9105 weakly neutralized autologous virus variants. Although this function was only demonstrated in one individual, it may exemplify a route to block vertical virus transmission, as the infant is solely exposed to maternal virus variants that coevolved with the autologous breast milk antibody responses. However, the function of our colostrum mAbs was not limited to neutralization, as we found that the neutralizing colostrum Env-specific mAbs also had other functions that may mediate protection against mucosal virus acquisition; including the capacity to inhibit HIV-1 virus transfer from DCs to CD4+ target cells44, 45 and to inhibit viral particle binding to an epithelial cell layer.46 Milk antibody blocking of virion interaction with mucosal cells before infection of CD4+ target cells may be key to protection of infants against HIV-1 infection in oral cavity or GI tract.
Whether or not bnAbs can be induced at mucosal sites remains unclear, however, we isolated two colostrum derived gp120 mAbs that may be precursor HIV-1 bnAbs. DH388 was sensitive to changes in the outer domain glycan at amino-acid position 334 in the C3 region of Env, the same high-mannose glycan patch targeted by bnAbs, PGT128, 130, and 131.47 DH388 additionally had a long CDR3 (29 amino acids) that is characteristic of bnAbs, yet it did not demonstrate any neutralizing function. Another potential bnAb precursor, the CD4 binding site-directed colostrum mAb DH378, weakly neutralized autologous viruses and a tier 2 heterologous SHIV virus, as well as inhibited epithelial and DC virus transfer. BnAbs have been shown to be effective at blocking oral virus transmission in neonatal rhesus macaques,48 but have yet to be elicited in mucosal or systemic compartments through vaccination. Thus, it will be important to determine whether these nonbroadly neutralizing colostrum IgG mAbs offer any protection against infant oral acquisition.
It is well described that IgA is trafficked from the gut-associated lymphoid tissue to the mammary gland through the gut–mammary axis16, 17 and it is theorized that IgG-producing B cells in milk follow this same trafficking pattern.49 It was recently reported that a subset of gp41-specific mAbs isolated from blood and intestine B cells have reactivity to intestinal microbiota WCLs29, 50 and may develop as a result of preinfection B-cell repertoire. Williams30 reported that Env-vaccine induced gp120 mAbs also crossreact with intestinal microbiota and we found that a large proportion of the gp120-specific colostrum mAbs isolated from infected individuals had crossreactivity to intestinal microbiota antigens. Of note, the panel of gp120-specific mAbs isolated from blood of chronically infected or vaccinated individuals with matched epitope specificity to our colostrum gp120-specific mAb panel had a similar proportion of commensal bacteria crossreactive mAbs. This suggests that the preexisting B-cell repertoire stimulated by commensal bacterial antigens may contribute to shaping both the systemic and breast milk Env-specific B-cell repertoire. We identified a specific bacterial antigen, Chaperonin 60, that crossreacted with the gp120-specific colostrum mAb, DH374. This crossreactive gp120-specific mAb bound to the linear and scaffolded V3 loop, yet there was limited homology between the linear V3 and Chaperonin sequence. As this mAb only bound to the nonnative Chaperonin 60 protein, it may target an epitope only displayed on the monomer of this heptameric protein.
In our analysis of two clonal lineages of gp120-specific colostrum mAbs isolated from the same subject, we observed two distinct patterns of evolution for gp120-specific and commensal microbiota reactive mAbs. In one clonal lineage, DH285, the germline naive B-cell receptor, or unmutated common ancestor, showed affinity maturation for the V1/V2 loop and away from commensal bacteria reactivity. In this case, the IgG-producing B cells may have been initially stimulated in the gut-associated lymphoid tissue and developed a strong affinity for the V1/V2 loop of gp120 through somatic hypermutation. A second pattern, demonstrated by the lineage of DH284, also involved affinity maturation toward gp120 Env specificity but demonstrated increasing cross-reactivity with commensal bacteria. As is theorized with HIV-1 gp41-specific antibodies,29, 51 this pattern may represent molecular mimicry of gp120 and commensal bacterial antigens. Alternatively, this commensal bacteria antigen crossreactivity may be a common feature of milk B cell-produced antibodies, as the maternal milk antibodies have also been described as regulators of the infant GI microbiome.52
There are several limitations to our study. First, we only characterized antibodies from a small number of HIV-1-infected, lactating subjects, focusing on subjects in whom we were able to isolate a large number of antibodies.12 Moreover, we characterized the function of only a few subject-matched systemic antibodies in parallel with the colostrum Env-specific antibody panel. Yet, the systemic Env-specific B-cell repertoire has been studied extensively in previous reports,29, 39, 53, 54 and therefore we focused our investigations on the colostrum antibodies. Beyond our association of milk and blood HIV-1 Env-specific and microbiota crossreactive antibodies, it would be relevant to probe the hypothesis that the preexisting commensal microbiota influences the HIV-1-specific IgG-producing B-cell repertoire. Furthermore, it would be interesting to isolate milk Env-specific B cells at various time points postpartum to track development of clonal lineages over time and further characterize mammary gland B-cell trafficking and evolution.
In summary, these data provide evidence that IgG antibodies produced by B cells resident in breast milk mediate important anti-HIV-1 functions that may contribute to protection of the infant against postnatal virus acquisition. This work supports the development of strategies to specifically target the induction of functional, HIV-1 Env-specific IgG-producing B cells in breast milk. In fact, we have previously reported that combined systemic and mucosal maternal HIV-1 Env vaccination can elicit functional antibody responses in milk of lactating nonhuman primates.9, 55 Maternal vaccination during breastfeeding to specifically target the B-cell population that traffics to and resides in breast milk may be a feasible and effective strategy to make breastfeeding safe for all infants born to HIV-1-infected women.
METHODS
Study subjects The subjects whose blood and colostrum samples were selected for analysis of antibody function were from the previously described CHAVI009 Malawian HIV-1-infected maternal cohort enrolled between 2008 and 2009.12, 14, 56 The majority of subjects were untreated or antiretrovirals were initiated during the third trimester. Single-dose Nevirapine was provided to all mothers and infants at the time of delivery. Maternal blood and milk samples were collected postpartum through the period of breastfeeding. HIV-1 transmission was determined by infant whole blood HIV-1 DNA PCR.24 The subjects’ baseline plasma viral load and peripheral CD4+ T-cell count was measured at the third trimester of pregnancy. Milk virus load was quantified by reverse transcriptase–PCR, with a detection limit of 240 RNA copies per ml, yet if virus RNA was detected in the sample but below the minimum for quantification, 120 RNA copies per ml was assigned.14 Blood and breast milk samples used in this study to isolate B cells were collected within 1 week postpartum, and therefore are referred to as colostrum.
Isolation of Env-specific B cells from colostrum and blood, antibody variable region gene amplification, sequencing, and screening for HIV-1 Env reactivity As described, breast milk cells and peripheral blood mononuclear cells isolated from EDTA-anticoagulated blood were thawed, stained, isolated, and sorted according to our previously reported data.12 B cells were sorted using fluorescently labeled ConS gp140 staining, with the exception of B cells from PTID CH8802 that used ConC gp120 antigen.
B-cell immunoglobulin gene PCR, sequencing, and HIV-1 Env reactivity screening were performed as previously described.12 Briefly, B cell expressed variable heavy- and light-chain genes were amplified from reverse transcriptase and nested PCR products, then purified and sequenced. Using the first round, functional immunoglobulin, PCR products were transiently transfected in 293T cells and supernatants of the transfected cells were screened for reactivity against HIV-1 Env proteins (C.1086gp12057 or MNgp41, obtained from Immunodiagnostics, Woburn, MA) by ELISA.
Selection of colostrum Env-specific mAbs for characterization The panel of colostrum Env-specific mAbs selected was originally from six HIV-1-infected mothers, five postpartum nontransmitting mothers, and one transmitting mother whose B-cell repertoires were previously analyzed.12 We selected colostrum IgG mAbs from three mothers (040-4, 910-5, and 920-8) who had a high number of mAbs isolated (between 7 and 14) and from three other mothers who had few antibodies isolated. We selected 19 colostrum mAbs for further characterization using the criteria of strong Env binding strength and breadth, and membership in an identified B-cell lineage. Antibody gene sequences were inserted into competent pcDNA 3.1 expression plasmids using standard molecular technology, and then these mAbs heavy- and light-chain plasmids were transfected into 293F and resulting IgG was protein A purified before binding and functional assays.
ELISA binding and blocking assays Recombinant mAbs expressed in small- and large-scale transfections were assayed for antibody reactivity to HIV-1 antigens by ELISA as previously described.50, 58 Purified antibodies were incubated with HIV-1 Env antigen-coated wells at concentrations ranging from 100 μg ml−1 to 5.6E−4 μg ml−1 at threefold dilutions with positivity cutoffs for reactivity set at threefold above background and an optical density of 0.180 at 100 μg ml−1. Fine specificity of gp120 colostrum mAbs was determined by binding to BconV3, CconV3,21 gp70MNV3,59 C.1086 V1/V2tags,60 CH505 gp120, and CH505 gp120 N334 mutant. The mAb blocking of A32 and soluble CD4 was performed as previously described.61 Briefly, 0.2 μg of protein (C.97ZMgp140, A244gp120, or ConSgp140)57 was added to each well and blocked with assay diluent (phosphate-buffered saline containing 4% (wt/vol) whey protein, 15% normal goat serum, 0.5% Tween-20, and 0.05% sodium azide). Each assay step was conducted in assay diluent and incubated for 1 h at room temperature and was followed by washing with phosphate-buffered saline/0.1% Tween-20, except the substrate and subsequent reaction stop step. The mAbs were titrated at concentrations ranging from 100 to 0.001 μg ml−1 at threefold dilutions. In addition, CH3162 and CH10663 were included as positive controls. For the CD4 blocking assay, soluble CD4 was added to each well at a concentration of 0.64 μg ml−1. Biotinylated A32 or OKT-4 (eBioscience, San Diego, CA) was added at the 50% effective concentration determined by a direct binding curve (biotinylated mAb vs. JRFL protein). Inhibition of biotin-mAb binding was detected with streptavidin–alkaline phosphatase at 1:1,000 (Promega, Madison, WI, V5591), followed by incubation with alkaline phosphatase substrate (2 mM MgCl2, 1 mg ml−1 p-NPP, and CBC, pH 9.6). Plates were read with a plate reader at 405 nm. For duplicate wells, background values were subtracted, and the results were averaged. Percent inhibition was calculated as follows: [1−(mean of triplicate wells/mean of no-inhibition control)] × 100.
HIV-1 pseudovirus neutralization in TZM-bl cells HIV-1 Env heterologous pseudovirus neutralization assays were performed as previously described.64 Briefly, colostrum mAbs were incubated with virus for 45–90 min, after which TZM-bl cells were added at a concentration of 1.0 × 105. Luminometry was used to determine whether antibodies were able to inhibit viral entry; then inhibitor concentration was calculated as a 50% reduction of relative light units, with respect to virus-only wells. All antibodies were tested for neutralization capacity against C.MW965 and 1086C in TZM-bl cells and, if positive, were screened against additional viruses. Virus variants BF 1677.613a, BF942.218d, and BF1266.431a are postnatal T/F HIV viruses,24 whereas 0724 and 0711 are 910-5 autologous virus variants. To isolate autologous virus from available plasma with detectable viral load, single genome amplification and sequencing was performed as previously described65 on plasma aliquots from PTID 910-5, resulting in only two viral variants, likely because of low virus load. Once functional viruses were amplified and sequenced, cloning and viral production was implemented as previously described.24
Epithelial cell inhibition assay To determine the ability of colostrum IgG mAbs to impede infectious virus binding to colonic epithelial cells (HT29), a modified previously reported protocol was used.24 Percent inhibition was calculated by dividing the relative light units of each well by the median relative light units of epithelial-bound virus that was not preincubated with isolated colostrum mAbs. The anti-influenza mAb CH65 IgG was used as a negative control, whereas the broadly neutralizing anti-HIV-1 CD4-binding site mAb VRC01 was used as a positive control. The cutoff value was determined by the mean inhibition plus 2 s.d. of CH65 IgG relative to no antibody.
DC HIV-1 trans-infection assay To determine whether our mAbs blocked virus transfer to monocyte-derived DCs, we used a method reported previously.24 The ability of various Abs to prevent the DC-mediated transfer of HIV MW965 was measured by comparing the TCID50 of the DC-bound virus and the total virus added to the DCs. The anti-influenza mAb CH65 IgG was used as a negative control, whereas the broadly neutralizing anti-HIV-1 CD4-binding site mAb VRC01 was used as a positive control. The cutoff value was determined by the mean plus 2 s.d. of CH65 relative to no antibody.
Antibody-dependent cellular cytotoxicity ADCC activity was determined by the GranToxiLux assay as previously described.66 Briefly, CEM.NKRCCR5 target cells (NIH AIDS Reagent Program, Division of AIDS, NIAID, NIH, from Alexandra Trkola, Germantown, MD) were coated with recombinant HIV-1 1086.C gp120. NK cells isolated by negative selection with magnetic beads (Miltenyi Biotec, Bergisch Gladbach, Germany) from cryopreserved peripheral blood mononuclear cells collected from a healthy HIV-1 seronegative donor with the F/F low-affinity Fc-gamma receptor (FcRγ) IIIa phenotype were used as effector cells. The NK cells were added to the gp120-coated target cells in wells of a 96-well plate at a ratio of 10:1. The colostrum mAbs were tested at a final concentration range of 40 to 0.01 μg ml−1. The RSV-specific mAb Palivizumab and the HIV-1-specific anti-C1 mAb A32 (ref. 67) were included as negative and positive controls, respectively. The assay plates were incubated for 1 h at 37 °C in 5% CO2. A minimum of 2,500 events representing viable target cells was acquired for each well using a LSRII flow cytometer (BD Biosciences, San Jose, CA). Data analysis was performed using FlowJo 9.8.1 software (Tree Star, Ashland, OR). The % GzB activity was defined as the percentage of cells positive for proteolytically active GzB out of the total viable target cell population. The final results are expressed after subtracting the background % GzB activity observed in wells containing no mAb samples. ADCC Ab titers were determined by interpolating the concentration of mAb that intersect the positive cutoff (>average+3 s.d. % GzB activity of mAbs tested against uncoated target cells) using GraphPad Prism 5 software (GraphPad, La Jolla, CA).
Western blot analysis of intestinal microbiota WCLs Colostrum mAbs were tested for gastrointestinal microbiota antigen reactivity as previously described.29 Previously characterized10 CH07 and CH08 were added to our panel of colostrum mAbs to round out the panel of 17 colostrum mAbs. A panel of gp120-specific mAbs isolated from the blood of both HIV-1-infected and vaccinated individuals matched by epitope specificity to the panel of colostrum-isolated gp120-specific mAbs including V2-specific mAbs CH58,60 CH59,60 HG107;60 V3-specific mAbs F39F,68 19b,69 CH22,70 CH16 (provided by B Haynes and the Center for HIV/AIDS Vaccine Immunology, Durham, NC); V1/V2 mAb PG9,71 V2/V3 mAb PG16;71 CD4i mAbs 17b,72 A32,73 CH38;74 CD4-binding site mAbs CH31,75 VRC01,76 CH13, CH17, and CH18 (provided by B Haynes and the Center for HIV/AIDS Vaccine Immunology); conformationally dependent mAbs CH21(ref. 74) and DH379 were included for comparison. Briefly, western blot analysis of intestinal microbiota WCL reactivity, 100 μg of human stool aerobic, and anaerobic bacterial WCLs were run on 4–12% Tris-Bis SDS-PAGE (Life Technologies, Grand Island, NY) for 1.5 h at 150 V in both reduced and nonreduced conditions. NuPAGE sample-reducing agent at 1 × was used for reducing conditions (Life Technologies). Antigens were transferred to nitrocellulose using Life Technologies iBlot Gel Transfer system. Antibody binding was tested at 20 μg ml−1 for all antibodies, and the anti-human IgG (whole molecule)-alkaline phosphatase antibody produced in goat (Sigma, St. Louis, MO) was used in a 1:5,000 dilution. Detection occurred directly on the nitrocellulose using Western Blue (Promega).
Autoantibody assays Assays to test autoreactivity of isolated colostrum mAbs were performed as previously described.31 Briefly, antibodies were assayed for reactivity to HEp-2 cells at 50 and 25 μg ml−1 (Inverness Medical Professional Diagnostics, Waltham, MA) by indirect immunofluorescence staining. Autoreactivity was also determined by antibody multiplex AtheNA Multi-Lyte ANA II test (Wampole Laboratories, Princeton, NJ) with a dose dilution starting at 50 μg ml−1 and determined positive when assay scores were ≥225 mean fluorescence intensity.
Surface plasmon resonance and Galcer blocking assays To confirm the reactivity of colostrum mAbs to commensal bacteria WCL and C.1086 gp120 Env, SPR binding assays were performed on a Biacore 4000 (GE Healthcare, Pittsburgh, PA) maintained at 25 °C as previously described.30 Briefly, the colostrum mAbs were immobilized on a Series S CM5 sensor chip (GE Healthcare) to 5,000 to 6,000 RUs using standard amine coupling chemistry. Uninfected human stool bacterial WCL, 1086Cgp120, or C.1086V1/V2 protein was injected over the immobilized colostrum mAbs. Injection time was 150 s and the dissociation activity was monitored for an additional 100 s. The maximal RU of binding at 150 s was reported. The dissociation constant (kd) was calculated using a 1:1 Langmuir model from 160 to 250 s. For bacterial WCL, avidity scores were calculated using the formula, RU/kd. All data analyses were performed using the BIAevaluation 4.1 analysis software (GE Healthcare).
The lipids, lyophilized powder of D-galactosyl-β-1,1' N-octanoyl-D-erythro-sphingosine (Galcer) and chloroform stock of 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), were purchased from Avanti Polar Lipids (Alabaster, AL).The Galcer liposomes were prepared in a 1:1 Galcer/POPC molar ratio and used in the Galcer blocking assay as previously detailed.20
References
Global Report: UNAIDS Report on the Global AIDS Epidemic: 2013. [cited]Available from http://www.unaids.org/sites/default/files/en/media/unaids/contentassets/documents/epidemiology/2013/gr2013/UNAIDS_Global_Report_2013_en.pdf.
Van de Perre, P. et al. Infective and anti-infective properties of breastmilk from HIV-1-infected women. Lancet 341, 914–918 (1993).
Datta, P. et al. Mother-to-child transmission of human immunodeficiency virus type 1: report from the Nairobi Study. J. Infect. Dis. 170, 1134–1140 (1994).
Fowler, M.G. & Newell, M.L. Breast-feeding and HIV-1 transmission in resource-limited settings. J. Acquir. Immune Defic. Syndr. 30, 230–239 (2002).
Coovadia, H.M. et al. Mother-to-child transmission of HIV-1 infection during exclusive breastfeeding in the first 6 months of life: an intervention cohort study. Lancet 369, 1107–1116 (2007).
Rogier, E.W. et al. Secretory antibodies in breast milk promote long-term intestinal homeostasis by regulating the gut microbiota and host gene expression. Proc. Natl. Acad. Sci. USA 111, 3074–3079 (2014).
Sadeharju, K. et al. Maternal antibodies in breast milk protect the child from enterovirus infections. Pediatrics 119, 941–946 (2007).
Newman, J. How breast milk protects newborns. Sci. Am. 273, 76–79 (1995).
Fouda, G.G. et al. Mucosal immunization of lactating female rhesus monkeys with a transmitted/founder HIV-1 envelope induces strong Env-specific IgA antibody responses in breast milk. J. Virol. 87, 6986–6999 (2013).
Friedman, J. et al. Isolation of HIV-1-neutralizing mucosal monoclonal antibodies from human colostrum. PLoS One 7, e37648 (2012).
Mabuka, J., Nduati, R., Odem-Davis, K., Peterson, D. & Overbaugh, J. HIV-specific antibodies capable of ADCC are common in breastmilk and are associated with reduced risk of transmission in women with high viral loads. PLoS Pathog. 8, e1002739 (2012).
Sacha, C.R. et al. Restricted isotype, distinct variable gene usage, and high rate of gp120 specificity of HIV-1 envelope-specific B cells in colostrum compared with those in blood of HIV-1-infected, lactating African women. Mucosal Immunol. 8, 316–326 (2014).
Haynes, B.F. et al. Immune-correlates analysis of an HIV-1 vaccine efficacy trial. N. Engl. J. Med. 366, 1275–1286 (2012).
Fouda, G.G. et al. HIV-specific functional antibody responses in breast milk mirror those in plasma and are primarily mediated by IgG antibodies. J. Virol. 85, 9555–9567 (2011).
Morris, L. et al. Isolation of a human anti-HIV gp41 membrane proximal region neutralizing antibody by antigen-specific single B cell sorting. PLoS One 6, e23532 (2011).
Gorny, M.K. et al. Functional and immunochemical cross-reactivity of V2-specific monoclonal antibodies from HIV-1-infected individuals. Virology 427, 198–207 (2012).
Mascola, J.R. & Haynes, B.F. HIV-1 neutralizing antibodies: understanding nature's pathways. Immunol. Rev. 254, 225–244 (2013).
Acharya, P. et al. Structural definition of an antibody-dependent cellular cytotoxicity response implicated in reduced risk for HIV-1 infection. J. Virol. 88, 12895–12906 (2014).
Moore, J.P. et al. Immunochemical analysis of the gp120 surface glycoprotein of human immunodeficiency virus type 1: probing the structure of the C4 and V4 domains and the interaction of the C4 domain with the V3 loop. J. Virol. 67, 4785–4796 (1993).
Dennison, S.M. et al. Vaccine-induced HIV-1 envelope gp120 constant region 1-specific antibodies expose a CD4-inducible epitope and block the interaction of HIV-1 gp140 with galactosylceramide. J. Virol. 88, 9406–9417 (2014).
Gao, F. et al. Cross-reactive monoclonal antibodies to multiple HIV-1 subtype and SIVcpz envelope glycoproteins. Virology 394, 91–98 (2009).
Moore, P.L. et al. Evolution of an HIV glycan-dependent broadly neutralizing antibody epitope through immune escape. Nat. Med. 18, 1688–1692 (2012).
Zolla-Pazner, S. et al. Vaccine-induced IgG antibodies to V1V2 regions of multiple HIV-1 subtypes correlate with decreased risk of HIV-1 infection. PLoS One 9, e87572 (2014).
Fouda, G.G. et al. Postnatally-transmitted HIV-1 Envelope variants have similar neutralization-sensitivity and function to that of nontransmitted breast milk variants. Retrovirology 10, 3 (2013).
Brenner, B.G., Gryllis, C. & Wainberg, M.A. Role of antibody-dependent cellular cytotoxicity and lymphokine-activated killer cells in AIDS and related diseases. J. Leukoc. Biol. 50, 628–640 (1991).
Pollara, J. et al. HIV-1 vaccine-induced C1 and V2 Env-specific antibodies synergize for increased antiviral activities. J. Virol. 88, 7715–7726 (2014).
Roux, M.E., McWilliams, M., Phillips-Quagliata, J.M., Weisz-Carrington, P. & Lamm, M.E. Origin of IgA-secreting plasma cells in the mammary gland. J. Exp. Med. 146, 1311–1322 (1977).
Barone, F. et al. IgA-producing plasma cells originate from germinal centers that are induced by B-cell receptor engagement in humans. Gastroenterology 140, 947–956 (2011).
Trama, A.M. et al. HIV-1 envelope gp41 antibodies can originate from terminal ileum B cells that share cross-reactivity with commensal bacteria. Cell Host Microbe 16, 215–226 (2014).
Williams, W.B. et al. Antibody Repertoire Induced by the Multiclade (Env A, B, C) HIV-1 DNA Prime, rAd5 Boost VRC Vaccine. Paper presented at: AIDS Vaccine Meeting; October 2013, Barcelona, Spain.
Haynes, B.F. et al. Cardiolipin polyspecific autoreactivity in two broadly neutralizing HIV-1 antibodies. Science 308, 1906–1908 (2005).
Ohtaka, C., Nakamura, H. & Ishikawa, H. Structures of chaperonins from an intracellular symbiont and their functional expression in Escherichia coli groE mutants. J. Bacteriol. 174, 1869–1874 (1992).
Fouda, G.G. et al. Tenascin-C is an innate broad-spectrum, HIV-1-neutralizing protein in breast milk. Proc. Natl. Acad. Sci. USA 110, 18220–18225 (2013).
Goldman, A.S. The immune system of human milk: antimicrobial, antiinflammatory and immunomodulating properties. Pediatr. Infect. Dis. J. 12, 664–671 (1993).
Telemo, E. & Hanson, L.A. Antibodies in milk. J. Mammary Gland Biol. Neoplasia 1, 243–249 (1996).
Rychert, J. & Amedee, A.M. The antibody response to SIV in lactating rhesus macaques. J. Acquir. Immune Defic. Syndr. 38, 135–141 (2005).
Yates, N.L. et al. Vaccine-induced Env V1-V2 IgG3 correlates with lower HIV-1 infection risk and declines soon after vaccination. Sci. Transl. Med. 6, 228ra239 (2014).
Mmiro, F.A. et al. Predictors of early and late mother-to-child transmission of HIV in a breastfeeding population: HIV Network for Prevention Trials 012 experience, Kampala, Uganda. J. Acquir. Immune Defic. Syndr. 52, 32–39 (2009).
Scheid, J.F. et al. Broad diversity of neutralizing antibodies isolated from memory B cells in HIV-infected individuals. Nature 458, 636–640 (2009).
Carias, A.M. et al. Defining the interaction of HIV-1 with the mucosal barriers of the female reproductive tract. J. Virol. 87, 11388–11400 (2013).
Montefiori, D.C. et al. V3-specific neutralizing antibodies in sera from HIV-1 gp160-immunized volunteers block virus fusion and act synergistically with human monoclonal antibody to the conformation-dependent CD4 binding site of gp120. NIH-NIAID AIDS Vaccine Clinical Trials Network. J. Clin. Invest. 92, 840–847 (1993).
Gorny, M.K. et al. Human anti-V3 HIV-1 monoclonal antibodies encoded by the VH5-51/VL lambda genes define a conserved antigenic structure. PLoS One 6, e27780 (2011).
Sok, D. et al. Promiscuous glycan site recognition by antibodies to the high-mannose patch of gp120 broadens neutralization of HIV. Sci. Transl. Med. 6, 236ra263 (2014).
Su, B. et al. Neutralizing antibodies inhibit HIV-1 transfer from primary dendritic cells to autologous CD4 T lymphocytes. Blood 120, 3708–3717 (2012).
Coleman, C.M., Gelais, C.S. & Wu, L. Cellular and viral mechanisms of HIV-1 transmission mediated by dendritic cells. Adv. Exp. Med. Biol. 762, 109–130 (2013).
Bobardt, M.D. et al. Cell-free human immunodeficiency virus type 1 transcytosis through primary genital epithelial cells. J. Virol. 81, 395–405 (2007).
Walker, L.M. et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477, 466–470 (2011).
Pegu, A. et al. Neutralizing antibodies to HIV-1 envelope protect more effectively in vivo than those to the CD4 receptor. Sci. Transl. Med. 6, 243ra288 (2014).
Tuaillon, E. et al. Human milk-derived B cells: a highly activated switched memory cell population primed to secrete antibodies. J. Immunol. 182, 7155–7162 (2009).
Liao, H.X. et al. Initial antibodies binding to HIV-1 gp41 in acutely infected subjects are polyreactive and highly mutated. J. Exp. Med. 208, 2237–2249 (2011).
Yang, G. et al. Identification of autoantigens recognized by the 2F5 and 4E10 broadly neutralizing HIV-1 antibodies. J. Exp. Med. 210, 241–256 (2013).
Latuga, M.S., Stuebe, A. & Seed, P.C. A review of the source and function of microbiota in breast milk. Semin. Reprod. Med. 32, 68–73 (2014).
Bonsignori, M. et al. Analysis of a clonal lineage of HIV-1 envelope V2/V3 conformational epitope-specific broadly neutralizing antibodies and their inferred unmutated common ancestors. J. Virol. 85, 9998–10009 (2011).
Huang, J. et al. Broad and potent neutralization of HIV-1 by a gp41-specific human antibody. Nature 491, 406–412 (2012).
Wilks, A.B. et al. Robust vaccine-elicited cellular immune responses in breast milk following systemic simian immunodeficiency virus DNA prime and live virus vector boost vaccination of lactating rhesus monkeys. J. Immunol. 185, 7097–7106 (2010).
Salazar-Gonzalez, J.F. et al. Origin and evolution of HIV-1 in breast milk determined by single-genome amplification and sequencing. J. Virol. 85, 2751–2763 (2011).
Liao, H.X. et al. Antigenicity and immunogenicity of transmitted/founder, consensus, and chronic envelope glycoproteins of human immunodeficiency virus type 1. J. Virol. 87, 4185–4201 (2013).
Liao, H.X. et al. High-throughput isolation of immunoglobulin genes from single human B cells and expression as monoclonal antibodies. J. Virol. Methods 158, 171–179 (2009).
Kayman, S.C., Wu, Z., Revesz, K., Chen, H., Kopelman, R. & Pinter, A. Presentation of native epitopes in the V1/V2 and V3 regions of human immunodeficiency virus type 1 gp120 by fusion glycoproteins containing isolated gp120 domains. J. Virol. 68, 400–410 (1994).
Liao, H.X. et al. Vaccine induction of antibodies against a structurally heterogeneous site of immune pressure within HIV-1 envelope protein variable regions 1 and 2. Immunity 38, 176–186 (2013).
Alam, S.M. et al. Human immunodeficiency virus type 1 gp41 antibodies that mask membrane proximal region epitopes: antibody binding kinetics, induction, and potential for regulation in acute infection. J. Virol. 82, 115–125 (2008).
Kwong, P.D. & Mascola, J.R. Human antibodies that neutralize HIV-1: identification, structures, and B cell ontogenies. Immunity 37, 412–425 (2012).
Liao, H.X. et al. Co-evolution of a broadly neutralizing HIV-1 antibody and founder virus. Nature 496, 469–476 (2013).
Li, M. et al. Human immunodeficiency virus type 1 env clones from acute and early subtype B infections for standardized assessments of vaccine-elicited neutralizing antibodies. J. Virol. 79, 10108–10125 (2005).
Salazar-Gonzalez, J.F. et al. Deciphering human immunodeficiency virus type 1 transmission and early envelope diversification by single-genome amplification and sequencing. J. Virol. 82, 3952–3970 (2008).
Pollara, J. et al. High-throughput quantitative analysis of HIV-1 and SIV-specific ADCC-mediating antibody responses. Cytometry A 79, 603–612 (2011).
Ferrari, G. et al. An HIV-1 gp120 envelope human monoclonal antibody that recognizes a C1 conformational epitope mediates potent antibody-dependent cellular cytotoxicity (ADCC) activity and defines a common ADCC epitope in human HIV-1 serum. J. Virol. 85, 7029–7036 (2011).
Gao, F. et al. Antigenicity and immunogenicity of a synthetic human immunodeficiency virus type 1 group m consensus envelope glycoprotein. J. Virol. 79, 1154–1163 (2005).
Robinson, J.E., Holton, D., Pacheco-Morell, S., Liu, J. & McMurdo, H. Identification of conserved and variant epitopes of human immunodeficiency virus type 1 (HIV-1) gp120 by human monoclonal antibodies produced by EBV-transformed cell lines. AIDS Res. Hum. Retroviruses 6, 567–579 (1990).
Montefiori, D.C. et al. Magnitude and breadth of the neutralizing antibody response in the RV144 and Vax003 HIV-1 vaccine efficacy trials. J. Infect. Dis. 206, 431–441 (2012).
Walker, L.M. et al. Broad and potent neutralizing antibodies from an African donor reveal a new HIV-1 vaccine target. Science 326, 285–289 (2009).
Aasa-Chapman, M.M. et al. In vivo emergence of HIV-1 highly sensitive to neutralizing antibodies. PLoS One 6, e23961 (2011).
Banerjee, K. et al. Enzymatic removal of mannose moieties can increase the immune response to HIV-1 gp120 in vivo. Virology 389, 108–121 (2009).
Bonsignori, M. et al. Antibody-dependent cellular cytotoxicity-mediating antibodies from an HIV-1 vaccine efficacy trial target multiple epitopes and preferentially use the VH1 gene family. J. Virol. 86, 11521–11532 (2012).
Wu, X. et al. Focused evolution of HIV-1 neutralizing antibodies revealed by structures and deep sequencing. Science 333, 1593–1602 (2011).
Wu, X. et al. Rational design of envelope identifies broadly neutralizing human monoclonal antibodies to HIV-1. Science 329, 856–861 (2010).
Acknowledgements
We acknowledge the following individuals for their technical contributions and support: Shaunna Shen, Ryan Duffy, Georgia Tomaras, David Martinez, Whitney Edwards, Rob Parks, Sabrina Arora, Krissey Lloyd, Jamie Pritchett, David Easterhoff, and Wilton Williams. Research reported in this publication was supported by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health R01 (AI106380), by the Center for HIV/AIDS Vaccine Immunology and Immunogen Discovery (UM1-AI100645-01), and by the Duke Center for AIDS Research (CFAR), an NIH-funded program (5P30 A1064518). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Supplementary information
Rights and permissions
About this article
Cite this article
Jeffries, T., Sacha, C., Pollara, J. et al. The function and affinity maturation of HIV-1 gp120-specific monoclonal antibodies derived from colostral B cells. Mucosal Immunol 9, 414–427 (2016). https://doi.org/10.1038/mi.2015.70
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2015.70