Abstract
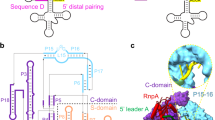
RNase P is the only endonuclease responsible for processing the 5′ end of transfer RNA by cleaving a precursor and leading to tRNA maturation1,2. It contains an RNA component and a protein component and has been identified in all organisms. It was one of the first catalytic RNAs identified3 and the first that acts as a multiple-turnover enzyme in vivo. RNase P and the ribosome are so far the only two ribozymes known to be conserved in all kingdoms of life. The RNA component of bacterial RNase P can catalyse pre-tRNA cleavage in the absence of the RNase P protein in vitro and consists of two domains: a specificity domain and a catalytic domain4,5. Here we report a 3.15-Å resolution crystal structure of the 154-nucleotide specificity domain of Bacillus subtilis RNase P. The structure reveals the architecture of this domain, the interactions that maintain the overall fold of the molecule, a large non-helical but well-structured module that is conserved in all RNase P RNA, and the regions that are involved in interactions with the substrate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Altman, S. & Kirsebom, L. A. in The RNA World (eds Gesteland, R. F., Cech, T. R. & Atkins, J. F.) 351–380 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1999)
Frank, D. N. & Pace, N. R. Ribonuclease P: unity and diversity in a tRNA processing ribozyme. Annu. Rev. Biochem. 67, 153–180 (1998)
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983)
Loria, A. & Pan, T. Domain structure of the ribozyme from eubacterial ribonuclease P. RNA 2, 551–563 (1996)
Pan, T. Higher order folding and domain analysis of the ribozyme from Bacillus subtilis ribonuclease P. Biochemistry 34, 902–909 (1995)
Qin, H., Sosnick, T. R. & Pan, T. Modular construction of a tertiary RNA structure: the specificity domain of the Bacillus subtilis RNase P RNA. Biochemistry 40, 11202–11210 (2001)
Brown, J. W. et al. Comparative analysis of ribonuclease P RNA using gene sequences from natural microbial populations reveals tertiary structural elements. Proc. Natl Acad. Sci. USA 93, 3001–3006 (1996)
Massire, C., Jaeger, L. & Westhof, E. Derivation of the three-dimensional architecture of bacterial ribonuclease P RNAs from comparative sequence analysis. J. Mol. Biol. 279, 773–793 (1998)
Chen, J. L., Nolan, J. M., Harris, M. E. & Pace, N. R. Comparative photocross-linking analysis of the tertiary structures of Escherichia coli and Bacillus subtilis RNase P RNAs. EMBO J. 17, 1515–1525 (1998)
Pan, T. Novel RNA substrates for the ribozyme from Bacillus subtilis ribonuclease P identified by in vitro selection. Biochemistry 34, 8458–8464 (1995)
Barrera, A. et al. Dimeric and monomeric Bacillus subtilis RNase P holoenzyme in the absence and presence of pre-tRNA substrates. Biochemistry 41, 12986–12994 (2002)
Nissen, P., Ippolito, J. A., Ban, N., Moore, P. B. & Steitz, T. A. RNA tertiary interactions in the large ribosomal subunit: the A-minor motif. Proc. Natl Acad. Sci. USA 98, 4899–4903 (2001)
Cate, J. H. et al. Crystal structure of a group I ribozyme domain: principles of RNA packing. Science 273, 1678–1685 (1996)
Odell, L., Huang, V., Jakacka, M. & Pan, T. Interaction of structural modules in substrate binding by the ribozyme from Bacillus subtilis RNase P. Nucleic Acids Res. 26, 3717–3723 (1998)
LaGrandeur, T. E., Huttenhofer, A., Noller, H. F. & Pace, N. R. Phylogenetic comparative chemical footprint analysis of the interaction between ribonuclease P RNA and tRNA. EMBO J. 13, 3945–3952 (1994)
Cate, J. H. et al. RNA tertiary structure mediation by adenosine platforms. Science 273, 1696–1699 (1996)
Moore, P. B. Structural motifs in RNA. Annu. Rev. Biochem. 68, 287–300 (1999)
Tanner, M. A. & Cech, T. R. An important RNA tertiary interaction of group I and group II introns is implicated in gram-positive RNase P RNAs. RNA 1, 349–350 (1995)
Costa, M. & Michel, F. Frequent use of the same tertiary motif by self-folding RNAs. EMBO J. 14, 1276–1285 (1995)
Costa, M. & Michel, F. Rules for RNA recognition of GNRA tetraloops deduced by in vitro selection: comparison with in vivo evolution. EMBO J. 16, 3289–3302 (1997)
Chen, J.-L. & Pace, N. R. Identification of the universally conserved core of ribonuclease P RNA. RNA 3, 557–560 (1997)
Brown, J. W. The Ribonuclease P Database. Nucleic Acids Res. 27, 314 (1999)
Frank, D. N., Adamidi, C., Ehringer, M. A., Pitulle, C. & Pace, N. R. Phylogenetic-comparative analysis of the eukaryal ribonuclease P RNA. RNA 6, 1895–1904 (2000)
Pan, T., Loria, A. & Zhong, K. Probing of tertiary interactions in RNA: 2′-hydroxyl-base contacts between the RNase P RNA and pre-tRNA. Proc. Natl Acad. Sci. USA 92, 12510–12514 (1995)
Loria, A. & Pan, T. Recognition of the T stem-loop of a pre-tRNA substrate by the ribozyme from Bacillus subtilis ribonuclease P. Biochemistry 36, 6317–6325 (1997)
Nolan, J. M., Burke, D. H. & Pace, N. R. Circularly permuted tRNAs as specific photoaffinity probes of ribonuclease P RNA structure. Science 261, 762–765 (1993)
Hendrickson, W. A. Determination of macromolecular structures from anomalous diffraction of synchrotron radiation. Science 254, 51–58 (1991)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Brunger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Haas, E. S., Banta, A. B., Harris, J. K., Pace, N. R. & Brown, J. W. Structure and evolution of ribonuclease P RNA in Gram-positive bacteria. Nucleic Acids Res. 24, 4775–4782 (1996)
Acknowledgements
We thank X. Liu and Y. Xiao for technical assistance, A. Changela, H. Feinberg, V. Grum and members of DND–CAT for help with data collection, and A. Changela, C. Correll, V. Grum, E. Sontheimer, B. Taneja and J. Wedekind for comments and suggestions. Research was supported by the NIH (to A.M.) and an NIH NRSA Fellowship to A.K. Support from the R.H. Lurie Cancer Center of Northwestern University to the Structural Biology Center is acknowledged. Portions of this work were performed at the DuPont–Northwestern–Dow Collaborative Access Team (DND–CAT) Synchrotron Research Center at the Advanced Photon Source (APS) and at the Stanford Synchrotron Radiation Laboratory (SSRL). DND–CAT is supported by DuPont, Dow and the NSF, and use of the APS is supported by the DOE. SSRL is operated by the DOE, Office of Basic Energy Sciences. The SSRL Biotechnology Program is supported by the NIH and the DOE.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Krasilnikov, A., Yang, X., Pan, T. et al. Crystal structure of the specificity domain of ribonuclease P. Nature 421, 760–764 (2003). https://doi.org/10.1038/nature01386
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01386
This article is cited by
-
Fluorogenic RNA Mango aptamers for imaging small non-coding RNAs in mammalian cells
Nature Communications (2018)
-
NMR resonance assignments of RNase P protein from Thermotoga maritima
Biomolecular NMR Assignments (2018)
-
Essential is Not Irreplaceable: Fitness Dynamics of Experimental E. coli RNase P RNA Heterologous Replacement
Journal of Molecular Evolution (2014)
-
Co-crystal structure of a T-box riboswitch stem I domain in complex with its cognate tRNA
Nature (2013)
-
Three-dimensional RNA structure refinement by hydroxyl radical probing
Nature Methods (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.