Abstract
MicroRNAs (miRNAs) are a growing family of small non-protein-coding regulatory genes that regulate the expression of homologous target-gene transcripts. They have been implicated in the control of cell death and proliferation in flies1,2, haematopoietic lineage differentiation in mammals3, neuronal patterning in nematodes4 and leaf and flower development in plants5,6,7,8. miRNAs are processed by the RNA-mediated interference machinery. Drosha is an RNase III enzyme that was recently implicated in miRNA processing. Here we show that human Drosha is a component of two multi-protein complexes. The larger complex contains multiple classes of RNA-associated proteins including RNA helicases, proteins that bind double-stranded RNA, novel heterogeneous nuclear ribonucleoproteins and the Ewing's sarcoma family of proteins. The smaller complex is composed of Drosha and the double-stranded-RNA-binding protein, DGCR8, the product of a gene deleted in DiGeorge syndrome. In vivo knock-down and in vitro reconstitution studies revealed that both components of this smaller complex, termed Microprocessor, are necessary and sufficient in mediating the genesis of miRNAs from the primary miRNA transcript.
Similar content being viewed by others
Main
It was recently suggested that most miRNA genes originate from independent transcription units9,10,11. However, about a quarter of human miRNA genes are located in introns of pre-mRNAs. Because these miRNAs have the same orientation as pre-mRNAs, it is likely that they are not transcribed from their own promoters but are processed from the introns12,13,14. The remaining miRNAs are clustered in the genome, predicting a multi-cistronic transcript9,10. Regardless of how different miRNAs originate, the primary miRNA transcript (pri-miRNA)15 must be processed to yield a mature 22-nucleotide (nt) miRNA. The present model for the pathway by which mammalian miRNAs undergo maturation begins with cleavage of the pri-miRNA in the nucleus to release a ∼60–70-nt stem loop intermediate, known as the pre-miRNA15,16. This processing event is mediated by an RNase III endonuclease, Drosha, which cleaves both strands of the stem at sites near the base of the primary stem loop17. Whether Drosha in itself is sufficient for this processing or whether other components associated with Drosha contribute to the cleavage of pri-miRNAs has not previously been addressed.
To gain insight into the components of the miRNA-processing machinery, we isolated a Drosha-containing complex from human cells. This was accomplished by developing HEK-293-derived stable cell lines expressing Flag-tagged Drosha. Flag-Drosha was isolated by immunoaffinity chromatography. As Fig. 1a demonstrates, the Flag affinity eluate contains a rich harvest of polypeptides. Drosha was represented in two forms differing in molecular mass by ∼10–15 kDa (Drosha a, ∼160 kDa, and Drosha b, ∼145 kDa; see Fig. 1a). It is currently not clear whether these two forms reflect post-translational modification of Drosha or result from proteolytic cleavage sustained during preparation of the nuclear extract.
a, Fractions of the immunoaffinity eluate from M2 anti-Flag beads resolved by SDS–PAGE (4–12%). Flag-Drosha was revealed by silver staining and western blotting with anti-Flag antibodies. Molecular masses of marker proteins (left) and the polypeptide masses of associated subunits (right) are indicated. b, Silver staining of Superose 6 gel-filtration fractions. Top, fraction number; bottom, molecular mass markers. DAP, Drosha-associated proteins having different molecular masses; asterisks, contaminating polypeptides; IP, immunoprecipitation.
To characterize the elution profiles of the two forms of Drosha and to demonstrate that the Drosha-associated polypeptides (DAPs) constitute a multi-protein complex, the Flag-affinity eluate was fractionated on a gel-filtration column (Fig. 1b). This analysis revealed the presence of the larger form of Drosha (Drosha a) in two distinct complexes: a high-molecular-mass complex containing most Drosha-associated polypeptides (fractions 16–18) and a lower-molecular-mass complex (fractions 22–26, ∼600 kDa). The smaller form of Drosha (Drosha b) displayed an elution profile consistent with a smaller complex with a molecular mass of ∼400 kDa (fractions 30–32).
We next determined the identity of the Drosha–associated polypeptides. The Flag-affinity eluate was separated on a polyacrylamide gel, bands were stained with colloidal blue, and individual polypeptides were excised from the gel and subjected to mass spectrometric sequencing. Nineteen specific Drosha-associated polypeptides were identified in two independent sequencing analyses (Supplementary Fig. S1). The Drosha-associated proteins comprised specific classes of RNA-associated proteins displaying common structural domains. These included the DEAD-box and DEAH-box family of RNA helicases, proteins with domains that bind double-stranded RNA, heterogeneous nuclear ribonucleoproteins (hnRNPs) and the Ewing's sarcoma family of proteins containing an RNA recognition motif (RRM) and a zinc-finger domain.
To address the association of the aforementioned different classes of polypeptides with Drosha, we analysed alternate fractions of Superose 6 gel-filtration chromatography by western blot analysis. This analysis revealed that Drosha coelutes with the protein product of Ewing's sarcoma gene (EWS), the RNA helicase, DDX17/P72 and hnRNPM4 in a large complex peaking in fraction 17 (Fig. 2a). In contrast with these polypeptides, a single polypeptide of 120 kDa corresponding to DGCR8 coelutes with the smaller complex peaking in fractions 25–27 (Figs 1b and 2a).
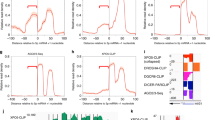
a, Western blotting with the antibodies shown at the left. IP, immunoprecipitation. b, Flag-affinity eluates and an enriched Dicer fraction were used for miRNA processing. ‘1 × ’ corresponds to 10 ng of Drosha and 5 µl of enriched Dicer fraction. c, miRNA processing with a pri-miRNA miR-(17,18,19a,20,19b-1) fragment. Flag-Drosha eluate (Input) corresponded to 5, 10, 20 and 40 ng of Drosha, determined as described in Methods.
We next analysed the Flag-affinity eluate for pri-miRNA processing. A 790-base-pair fragment corresponding to a cluster of pri-miRNAs miR-(17,18,19a,20,19b-1) was used as a substrate for analysis of pri-miRNA processing. It was shown recently that pri-miRNA processing catalysed by Drosha resulted in the formation of a ∼60–70-base-pair pre-miRNA precursor17. We first examined the activity of the Flag-Drosha affinity eluate in processing the pri-miRNA. Addition of increasing concentrations of Flag-Drosha affinity eluate to the pri-miRNA processing reaction resulted in the appearance of a distinct 63-nt pre-miRNA fragment (Fig. 2b). Further addition of a fraction enriched for Dicer, which catalyses further processing of the pre-miRNA, converted this fragment to a mature ∼22-nt miRNA, confirming that this pre-miRNA is the correct processing product (Fig. 2b).
We next analysed the gel-filtration fractions for pri-miRNA processing activity and found it was present in two distinct peaks corresponding to the two Drosha-containing complexes (Fig. 2c). Although the larger complex (fractions 16–18) displayed some pri-miRNA processing activity, the bulk of pri-miRNA processing activity co-eluted with the smaller Drosha complex containing DGCR8. Moreover, closer examination of the reaction revealed that the fractions containing the larger Drosha complex contained a non-specific RNase activity, resulting in reduced probe concentrations.
To characterize the polypeptide composition of the smaller Drosha-containing complex, fractions 24–27 were concentrated by trichloroacetic acid, treated as described above and subjected to mass spectrometric sequencing. The three largest polypeptides corresponded to Drosha a and b forms and the DGCR8 protein (Supplementary Fig. S2). (The smaller polypeptide corresponded to SKB1, a common contaminant of Flag-affinity purification.) We then analysed the specific pri-miRNA processing activity of the two Drosha complexes by normalizing the amounts of Drosha in each complex (Supplementary Fig. S2). These studies revealed that the Drosha–DGCR8 complex displays a nearly eightfold greater pri-miRNA processing activity than the large Drosha complex (Supplementary Fig. S2).
We next developed HEK-293-derived stable cell lines expressing Flag-DGCR8. Flag-DGCR8 was isolated by affinity chromatography. As shown in Fig. 3a, the DGCR8 affinity eluate contained a single polypeptide corresponding to Drosha in addition to DGCR8 itself. The polypeptide below DGCR8 corresponds to a carboxy-terminal truncation of the protein, because only the amino-terminal DGCR8 antibodies recognize this truncated protein (Fig. 3a). The Flag-DGCR8–Drosha complex was then used to assess its activity for pri-miRNA processing. Consistent with the results obtained with the smaller Drosha complex (Supplementary Fig. S2), Flag-DGCR8–Drosha exhibited robust pri-miRNA processing activity (Fig. 3b). These results demonstrate the stable association of Drosha and DGCR8 in an active pri-miRNA processing complex. For the sake of consistency we have called this complex Microprocessor18. A parallel study shows that a Drosophila homologue of DGCR8 associates with Drosha and pri-miRNA processing activity in Drosophila, Caenorhabditis elegans and mammals for a role of this protein in miRNA biogenesis18.
a, Isolation of Flag-DGCR8 complex. SDS–PAGE followed by silver staining and western blot analysis with antibodies shown on the figure are displayed. b, Analysis of pri-miRNA processing activity of Flag-DGCR8–Drosha complex. c, Analysis of miRNA processing activity of Microprocessor purified through Flag-Drosha (fraction 25; Fxn 25) or Flag-DGCR8 with the use of the miRNAs shown.
We next analysed the activity of Microprocessor purified by either Flag-Drosha or Flag-DGCR8 affinity chromatography for processing of two other pri-miRNAs. miR-(15,16) and miR-(23,27,24-2) were processed to yield expected miRNA precursors as was shown previously15 (Fig. 3c). Moreover, analysis of the processed fragments by northern blot analysis and further processing by Dicer confirmed the specific processing by Microprocessor (Supplementary Fig. S3).
To demonstrate rigorously that pri-miRNA processing required DGCR8 in addition to the RNase III Drosha, we reconstituted pri-miRNA processing activity by using recombinant Drosha produced in insect cells and recombinant DGCR8 generated in bacteria (Fig. 4a). Both recombinant proteins were purified to near homogeneity and were used in near-stoichiometric amounts in pri-miRNA processing assays (Fig. 4b, c). Although native Microprocessor displayed robust pri-miRNA processing activity, neither recombinant Drosha nor recombinant DGCR8 showed any significant pri-miRNA processing activity (Fig. 4c, lanes 4–7). However, addition of the two recombinant proteins reconstituted the pri-miRNA processing activity to levels similar to those seen with native complex (Fig. 4c, lanes 8 and 9). Whereas the addition of increasing concentrations of DGCR8 inhibited the processing reaction perhaps through a squelching mechanism, a further increase in Drosha stimulated the processing of pri-miRNA (Fig. 4c, compare lanes 10, 11 and 12, 13). Interestingly, whereas Drosha alone displayed non-specific RNase activity on the substrate, the addition of DGCR8 inhibited these non-specific effects and promoted Drosha's pri-miRNA processing activity (Fig. 4c, compare lanes 5 and 9). These results show the requirement for DGCR8 in directing the specific processing of pri-miRNAs by Drosha.
a, Analysis of recombinant Drosha and DGCR8 with colloidal blue staining; 100 ng of each protein was analysed. b, As in a except that ‘1 × ’ corresponds to 50 ng of BSA, used to determine Drosha and DGCR8 concentrations. c, Reconstitution of miRNA processing by using recombinant Drosha and DGCR8. ‘1 × ’ corresponds to 10 ng of Drosha in fraction 26 (Fxn 26) of Superose 6 and 20 ng of each of the recombinant (r) proteins assayed for pri-miRNA processing activity as described in Methods.
To assess the role of Microprocessor in the initiation of miRNA processing in vivo, we used RNA interference to deplete DGCR8 and Drosha. For these experiments short interfering RNA (siRNA) against Drosha was used as a positive control, whereas siRNA against transcription factor TFII-I was used as negative control (Fig. 5a). Knock-down of DGCR8 caused a similar effect to Drosha depletion for all miRNAs tested, resulting in a pronounced decrease in mature miRNA levels (Fig. 5b). Depletion of both Drosha and DGCR8, however, resulted in a substantial accumulation of pri-miRNA, showing the requirement for Microprocessor in miRNA processing in vivo (Fig. 5c). These results show the obligatory role for Drosha and DGCR8 in microRNA processing in vivo and in vitro.
a, Analysis of transcript levels by using RT–PCR for Drosha and DGCR8 after treatment of HeLa cells with siRNA against each protein. siRNA against TFII-I was used as control. b, Northern blot analysis of miRNA-21, miRNA-16, miRNA-23, let-7a-1 and miRNA-20 after treatment of HeLa cells with siRNA against Drosha, DGCR8 or control siRNA. c, Analysis of pri-miRNA processing after depletion of Drosha and DGCR8. -RT, absence of reverse transcriptase.
We have isolated two multi-protein complexes that contain Drosha as their catalytic engine. We show that the smaller Microprocessor complex containing Drosha and DGCR8 is necessary and sufficient for the processing of pri-miRNA to pre-miRNA. This contention is based on the following observations. First, Microprocessor purified by Flag-Drosha affinity purification displays specific and robust activity in pri-miRNA processing. Second, isolation of Microprocessor through Flag-DGCR8 using stable cell lines revealed its close association with Drosha and miRNA processing activity. Third, miRNA processing activity could be reconstituted with recombinant Drosha and DGCR8. Last, knock-down of Drosha and DGCR8 resulted in diminished mature miRNA and accumulation of pri-miRNA in vivo.
DGCR8 is an evolutionarily conserved protein that contains two double-stranded RNA-binding domains and a WW domain known to interact with proline-rich peptides. The WW domain of DGCR8 is most probably the interacting surface with the proline-rich N-terminal domain of Drosha. DGCR8 is one of an estimated 30 genes in the chromosomal region 22q11.2 whose heterozygous deletion results in the most common human genetic deletion syndrome, known as DiGeorge syndrome19,20. The clinical manifestations of the disease are highly variable, with 75% of patients displaying congenital heart defects. Other common features include, among many others, characteristic facial appearance, immunodeficiency resulting from thymic hypoplasia, and developmental and behavioural problems. It will be important to explain the role of DGCR8 in the genesis and development of DiGeorge syndrome. Future experiments with knockout mouse models of DGCR8 in homozygous and heterozygous animals will shed light on the likely function of DGCR8 in developmental control and the expression of DiGeorge syndrome-like phenotypes.
In the past we have used the 2-MDa chromatin remodelling complex, SWI–SNF, as a size maker for gel-filtration chromatography. Comparing the elution profiles of the large Drosha complex with that of SWI–SNF (which peaks in fraction 20 on Superose 6) we conclude that the large Drosha complex is larger than SWI–SNF. We have identified nearly 20 polypeptides that are specifically associated with Drosha in this complex. These included RNA helicases containing a DEAD-box or DEAH-box, hnRNPs and proteins with either double-stranded RNA-binding or RRM domains. We have also identified EWS as components of this larger Drosha complex. Ewing's family of tumours result from tumour-associated chromosomal translocations between EWS genes and one of five different ETS transcription factors21.
We cannot exclude a role for the large Drosha complex in miRNA processing because it displayed a weak pre-miRNA processing activity in vitro. Moreover, our analysis in vivo with siRNA knock-down of three components, p62/DDX5, p72/DDX17 and hnRNPU1-like, revealed a small decrease in mature miRNAs, although abrogation of these subunits never approached the effects observed after DGCR8 and Drosha depletions. It is therefore more likely that the large Drosha-containing complex has a function in other RNA processing pathways. Because Drosha has also been previously shown to participate in preribosomal RNA processing22, the large Drosha complex might mediate such preribosomal RNA processing activities.
Methods
Affinity purification of Flag-Drosha and Flag-DGCR8
Flag-Drosha and a selectable marker for puromycin resistance were cotransfected into HEK-293 human embryonic kidney cells. Transfected cells were grown in the presence of 2.5 µg ml-1 puromycin, and individual colonies were isolated and analysed for Flag-Drosha expression. To purify the Drosha complex, nuclear extract generated from 200 15-cm plates (4 × 109 cells or ∼150 mg of nuclear extract) was incubated with anti-Flag M2 affinity resin (Sigma). After two washes with buffer A (20 mM Tris-HCl pH 7.9, 0.5 M KCl, 10% glycerol, 1 mM EDTA, 5 mM dithiothreitol, 0.5% Nonidet P40, 0.2 mM phenylmethylsulphonyl fluoride), the affinity column was eluted with buffer A containing Flag peptide (400 µg ml-1) in accordance with the manufacturer's instructions (Sigma). Analysis of Drosha by Superose 6 gel filtration was similar to that previously described23,24. Fractions from the gel-filtration chromatography were concentrated by precipitation with trichloroacetic acid and analysed by SDS–PAGE followed by silver staining. Flag-DGCR8 was isolated by using a similar protocol. Protein identification by liquid chromatography–tandem mass spectroscopy was performed as detailed23. Multiple sequencing analyses have determined SKB1, MEP50 and α–tubulin as common contaminants of Flag-affinity purification. Anti-hnRNP M4 antibody was obtained from Santa Cruz Biotechnology. Anti-Drosha and anti-DGCR8 were generated against the last 20 amino acids in the amino and carboxy termini of each protein (Open Biosystems). Recombinant Drosha and DGCR8 were purified by methods previously described for the purification of recombinant proteins from insect cells and bacteria23,24. In brief, each protein was expressed as a Flag-tagged protein and purified with anti-Flag M2 affinity resin similar in manner to the protocol described for Flag-Drosha and Flag-DGCR8.
miRNA processing
In vitro transcription was performed with the Promega Riboprobe system, using linearized pGEM-7Z vector containing miR-(17,18,19a,20,19b-1), miR-(15,16) and miR-(23,27,24-2) as described15. In brief, the miRNA probes were amplified from HEK-293 RNA by reverse transcriptase-mediated polymerase chain reaction (RT–PCR) with 5′-tgctgaatttgtatggtttatagttgtta-3′ as 5′ primer and 5′-tacttttctacagacttttcactaccaca-3′ as 3′ primer for miR-(17,18,19a,20,19b-1), 5′-CGCCCGGTGCCCCCCTCACCCCTGTGCCAC-3′ as 5′ primer and 5′-CCCTGTTCCTGCTGAACTGAGCCAGTGTAC-3′ as 3′ primer for miR-(23,27,24-2) and 5′-CCTTGGAGTAAAGTAGCAGCAACTAATG-3′ as 5′ primer and 5′-CTTACTCTGAGTTAAATCTTGAATAC-3′ as 3′ primer for miR-(15,16). The processing reaction, containing the indicated amounts of Drosha complex, 3 µl of solution containing 32 mM MgCl2, 10 mM ATP, 200 mM creatine phosphate, 1 U µl-1 HPRI (Takara) and the labelled pri-miRNA (2 × 105 c.p.m.) and buffer (20 mM Tris-HCl pH 7.9, 0.1 M KCl, 10% glycerol, 5 mM dithiothreitol, 0.2 mM phenylmethylsulphonyl fluoride), was added to a final volume of 30 µl. The reaction mixture was incubated at 37 °C for 90 min and extracted with phenol:chloroform mixture, then with chloroform and precipitated with 300 mM sodium acetate and ethanol. The precipitated RNA was loaded on 15% denaturing polyacrylamide gels. For the reconstitution experiments, the two recombinant proteins were incubated for 1 h on ice in buffer A containing 100 mM KCl before the addition of the reaction mix. All Drosha quantitative analyses were performed by comparing the Flag-affinity eluate and recombinant Drosha by quantitative western blot analysis. The amounts of recombinant Drosha were then deduced by using colloidal blue staining in comparison with known amounts of BSA.
siRNAs and transfection
The siRNAs were synthesized by Dharmacon. The sequence of Drosha siRNA was 5′-AAGGACCAAGUAUUCAGCAAG-3′, the DGCR8 siRNA was 5′-AUCCGUUGAUCUCGAGGAATT-3′, and the control siRNA against TFIII was 5′-UGUGGGGAAGCUCUUGGCCTT-3′. siRNA transfection in HeLa cells was performed with Lipofectamine 2000 (Invitrogen). In brief, cells were plated in 10-cm dishes to 40% confluence. For each dish, 1.6 nmol of siRNA was mixed with 24 µl of Lipofectamine 2000 in 3 ml of Opti-MEM medium. The mixture was added to cells and incubated for 6 h. After 24 h a second transfection was performed in the same way. Total RNA was prepared 3 days after the second transfection and was used for northern blot analysis or RT–PCR.
RNA isolation, RT–PCR and northern blot analysis
Total RNA from HeLa cells was prepared in TRIzol reagents (Invitrogen) in accordance with the manufacturer's instructions. To examine the effect of siRNA, RNA (2 µg) was subjected to complementary DNA synthesis with oligo(dT), using the SuperScript first-strand synthesis system for RT–PCR (Invitrogen). To examine pri-miRNAs, RNA (2 µg) was subjected to cDNA synthesis with random primers. As a control, RT–PCRs were performed in the absence of reverse transcriptase. Different PCR cycles were examined to determine linear amplification.
Primer sequences for RT–PCRs in Fig. 5a were Drosha (5′-CATGCACCAGATTCTCCT GTA-3′ and 5′-GTCTCCTGCATAACTCAACTG-3′) and DGCR8 (5′-TATCAGATCCTCCACGAGTG-3′ and 5′-TCTTGGAGCTTGCTGAGGAT-3′). Primer sequences for RT–PCRs in Fig. 5c were pri-let-7a-1 (5′-GATTCCTTTTCACCATTCACCCTGGATGTT-3′ and 5′-TTTCTATCAGACCGCCTGGATGCAGACTTT-3′), pri-miR30a (5′-ATTGCTGTTTGAATGAGGCTTCAGTACTTT-3′ and 5′-TTCAGCTTTGTAAAAATGTATCAAAGAGAT-3′) and pri-miR17 (5′-TGCTGAATTTGTATGGTTTATAGTTGTTAG-3′ and 5′-CACTACCACAGTCAGTTTTGCATGGATTTG-3′). β-Actin was used for internal control in Fig. 5a, c, and the primer sequences were 5′-AAAGACCTGTACGCCAACAC-3′ and 5′-GTCATACTCCTGCTTGCTGAT-3′.
For northern blot analysis, total RNA (10 µg) was resolved on a 15% denaturing polyacrylamide gel and electrotransferred to Hybond N+ nylon membrane (Amersham). The membranes were crosslinked under ultraviolet and prewashed for 1 h at 65 °C in 0.1 × SSC/0.1% SDS. Prehybridization and hybridization were performed at 42 °C in 10 × Denharts solution, 6 × SSC, 0.1% SDS. Oligonucleotides complementary to miRNAs were end-labelled with [γ-32P]ATP and used as probes for northern analysis. The sequences of the oligonucleotides were 5′-gaaaatccctggcaatgtgat-3′ (miR-23), 5′-actatacaacctactacctca-3′ (let-7a-1), 5′-gccaatatttacgtgctgcta-3′ (miR-16), 5′-tacctgcactataagcacttta-3′ (miR-20), 5′-tcaacatcagtctgataagcta-3′ (miR-21) and 5′-caggcccgaccctgcttagcttccgagatcagacgagat-3′ (5S rRNA). All of the probes were washed twice for 10 min at 25 °C in 6 × SSC/0.1% SDS.
References
Brennecke, J. et al. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113, 25–36 (2003)
Xu, P. et al. The Drosophila microRNA mir-14 suppresses cell death and is required for normal fat metabolism. Curr. Biol. 13, 790–795 (2003)
Chen, C. Z. et al. MicroRNAs modulate hematopoietic lineage differentiation. Science 303, 83–86 (2004)
Johnston, R. J. & Hobert, O. A microRNA controlling left/right neuronal asymmetry in Caenorhabditis elegans. Nature 426, 845–849 (2003)
Aukerman, M. J. & Sakai, H. Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15, 2730–2741 (2003)
Chen, X. et al. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303, 2022–2025 (2004)
Emery, J. F. et al. Radial patterning of Arabidopsis shoots by class III HD-ZIP and KANADI genes. Curr. Biol. 13, 1768–1774 (2003)
Palatnik, J. F. et al. Control of leaf morphogenesis by microRNAs. Nature 425, 257–263 (2003)
Lagos-Quintana, M. et al. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001)
Lau, N. C. et al. An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294, 858–862 (2001)
Lee, R. C. & Ambros, V. An extensive class of small RNAs in Caenorhabditis elegans. Science 294, 862–864 (2001)
Aravin, A. A. et al. The small RNA profile during Drosophila melanogaster development. Dev. Cell 5, 337–350 (2003)
Lagos-Quintana, M. et al. New microRNAs from mouse and human. RNA 9, 175–179 (2003)
Lim, L. P. et al. The microRNAs of Caenorhabditis elegans. Genes Dev. 17, 991–1008 (2003)
Lee, Y., Jeon, K., Lee, J. T., Kim, S. & Kim, V. N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21, 4663–4670 (2002)
Zeng, Y. & Cullen, B. R. MicroRNAs and small interfering RNAs can inhibit mRNA expression by similar mechanisms. Proc. Natl Acad. Sci. USA 100, 9779–9784 (2003)
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003)
Denali, A. M., Tops, B. B. J., Plasterk, R. H. A., Ketting, R. F. & Hannon, G. J. Processing of primary microRNAs by the Microprocessor complex. Nature doi:10.1038/nature03049 (this issue)
Yamagishi, H. & Srivastava, D. Unraveling the genetic and developmental mysteries of 22q11 deletion syndrome. Trends Mol. Med. 9, 383–389 (2003)
Shiohama, A., Sasaki, T., Noda, S., Minoshima, S. & Shimizu, N. Molecular cloning and expression analysis of a novel gene DGCR8 located in DiGeorge syndrome chromosomal region. Biochem. Biophys. Res. Commun. 304, 184–190 (2003)
Arvand, A. & Denny, C. T. Biology of EWS/ETS fusions in Ewing's family tumors. Oncogene 20, 5747–5754 (2001)
Wu, H., Xu, H., Miraglia, L. J. & Crooke, S. T. Human RNase III is a 160-kDa protein involved in preribosomal RNA processing. J. Biol. Chem. 275, 36957–36965 (2000)
Bochar, D. A. et al. BRCA1 is associated with a human SWI/SNF-related complex: linking chromatin remodeling to breast cancer. Cell 102, 257–265 (2000)
Dong, Y. et al. Regulation of BRCC, a holoenzyme complex containing BRCA1 and BRCA2, by a signalosome-like subunit and its role in DNA repair. Mol. Cell 12, 1087–1099 (2003)
Acknowledgements
We thank V. N. Kim for the Drosha cDNA; O. Delattre and A. I. Lamond for the gift of EWS and DDX17 antibodies, respectively; K. Nishikura for providing recombinant Drosha; and T. Beer (Wistar Proteomocs Facility) for expertise in the microcapillary HPLC/mass spectrometry. R.S. was supported by grants from the NIH and the American Cancer Institute. R.G. is a fellow of the Jane Coffin Child Memorial Fund for Medical Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure 1
Diagrammatic depiction of Drosha-associated proteins and their domain structures. (PDF 47 kb)
Supplementary Figure 2
The bulk of miRNA processing activity resides in Drosha/DGCR8 complex. (PDF 281 kb)
Supplementary Figure 3
Analysis of miRNA precursors. (PDF 425 kb)
Rights and permissions
About this article
Cite this article
Gregory, R., Yan, Kp., Amuthan, G. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004). https://doi.org/10.1038/nature03120
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature03120
This article is cited by
-
N4-acetylcytidine modifies primary microRNAs for processing in cancer cells
Cellular and Molecular Life Sciences (2024)
-
An Overview of Targeted Genome Editing Strategies for Reducing the Biosynthesis of Phytic Acid: an Anti-nutrient in Crop Plants
Molecular Biotechnology (2024)
-
Nuclear receptor RXRα binds the precursor of miR-103 to inhibit its maturation
BMC Biology (2023)
-
The various role of microRNAs in breast cancer angiogenesis, with a special focus on novel miRNA-based delivery strategies
Cancer Cell International (2023)
-
Non-coding RNAs identification and regulatory networks in pathogen-host interaction in the microsporidia congenital infection
BMC Genomics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.