Abstract
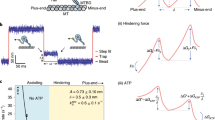
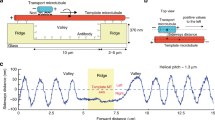
Microtubules (MTs) are important components of the eukaryotic cytoskeleton: they contribute to cell shape and movement, as well as to the motions of organelles including mitotic chromosomes. MTs bind motor enzymes that drive many such movements, but MT dynamics can also contribute to organelle motility1,2,3,4,5,6,7,8. Each MT polymer is a store of chemical energy that can be used to do mechanical work, but how this energy is converted to motility remains unknown. Here we show, by conjugating glass microbeads to tubulin polymers through strong inert linkages, such as biotin–avidin, that depolymerizing MTs exert a brief tug on the beads, as measured with laser tweezers. Analysis of these interactions with a molecular-mechanical model of MT structure and force production9,10 shows that a single depolymerizing MT can generate about ten times the force that is developed by a motor enzyme; thus, this mechanism might be the primary driving force for chromosome motion. Because even the simple coupler used here slows MT disassembly, physiological couplers may modulate MT dynamics in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Inoue, S. & Salmon, E. D. Force generation by microtubule assembly/disassembly in mitosis and related movements. Mol. Biol. Cell 6, 1619–1640 (1995)
Dogterom, M., Kerssemakers, J. W., Romet-Lemonne, G. & Janson, M. E. Force generation by dynamic microtubules. Curr. Opin. Cell Biol. 17, 67–74 (2005)
Waterman-Storer, C. M. & Salmon, E. D. Endoplasmic reticulum membrane tubules are distributed by microtubules in living cells using three distinct mechanisms. Curr. Biol. 8, 798–806 (1998)
Waterman-Storer, C. M., Worthylake, R. A., Liu, B. P., Burridge, K. & Salmon, E. D. Microtubule growth activates Rac1 to promote lamellipodial protrusion in fibroblasts. Nature Cell Biol. 1, 45–50 (1999)
Koshland, D. E., Mitchison, T. J. & Kirschner, M. W. Polewards chromosome movement driven by microtubule depolymerization in vitro. Nature 331, 499–504 (1988)
Coue, M., Lombillo, V. A. & McIntosh, J. R. Microtubule depolymerization promotes particle and chromosome movement in vitro. J. Cell Biol. 112, 1165–1175 (1991)
Lombillo, V. A., Nislow, C., Yen, T. J., Gelfand, V. I. & McIntosh, J. R. Antibodies to the kinesin motor domain and CENP-E inhibit microtubule depolymerization-dependent motion of chromosomes in vitro. J. Cell Biol. 128, 107–115 (1995)
Lombillo, V. A., Stewart, R. J. & McIntosh, J. R. Minus-end-directed motion of kinesin-coated microspheres driven by microtubule depolymerization. Nature 373, 161–164 (1995)
Molodtsov, M. I. et al. A molecular-mechanical model of the microtubule. Biophys. J. 88, 3167–3179 (2005)
Molodtsov, M. I., Grishchuk, E. L., Efremov, A. K., McIntosh, J. R. & Ataullakhanov, F. I. Force production by depolymerizing microtubules: a theoretical study. Proc. Natl Acad. Sci. USA 102, 4353–4358 (2005)
Muller-Reichert, T., Chretien, D., Severin, F. & Hyman, A. A. Structural changes at microtubule ends accompanying GTP hydrolysis: information from a slowly hydrolyzable analogue of GTP, guanylyl (α,β)methylenediphosphonate. Proc. Natl Acad. Sci. USA 95, 3661–3666 (1998)
Gigant, B. et al. The 4Å X-ray structure of a tubulin:stathmin-like domain complex. Cell 102, 809–816 (2000)
Caplow, M., Ruhlen, R. L. & Shanks, J. The free energy for hydrolysis of a microtubule-bound nucleotide triphosphate is near zero: all of the free energy for hydrolysis is stored in the microtubule lattice. J. Cell Biol. 127, 779–788 (1994)
Howard, J. & Hyman, A. A. Dynamics and mechanics of the microtubule plus end. Nature 422, 753–758 (2003)
Mandelkow, E. M., Mandelkow, E. & Milligan, R. A. Microtubule dynamics and microtubule caps: a time-resolved cryo-electron microscopy study. J. Cell Biol. 114, 977–991 (1991)
Hyman, A. A., Salser, S., Drechsel, D. N., Unwin, N. & Mitchison, T. J. Role of GTP hydrolysis in microtubule dynamics: information from a slowly hydrolyzable analogue, GMPCPP. Mol. Biol. Cell 3, 1155–1167 (1992)
Vigers, G. P., Coue, M. & McIntosh, J. R. Fluorescent microtubules break up under illumination. J. Cell Biol. 107, 1011–1024 (1988)
Allersma, M. W., Gittes, F., deCastro, M. J., Stewart, R. J. & Schmidt, C. F. Two-dimensional tracking of ncd motility by back focal plane interferometry. Biophys. J. 74, 1074–1085 (1998)
Visscher, K. & Block, S. M. Versatile optical traps with feedback control. Methods Enzymol. 298, 460–489 (1998)
Svoboda, K. & Block, S. M. Force and velocity measured for single kinesin molecules. Cell 77, 773–784 (1994)
Westermann, S. et al. Formation of a dynamic kinetochore-microtubule interface through assembly of the Dam1 ring complex. Mol. Cell 17, 277–290 (2005)
Gildersleeve, R. F., Cross, A. R., Cullen, K. E., Fagen, A. P. & Williams, R. C. Jr Microtubules grow and shorten at intrinsically variable rates. J. Biol. Chem. 267, 7995–8006 (1992)
O'Brien, E. T., Salmon, E. D., Walker, R. A. & Erickson, H. P. Effects of magnesium on the dynamic instability of individual microtubules. Biochemistry 29, 6648–6656 (1990)
Walker, R. A. et al. Dynamic instability of individual microtubules analyzed by video light microscopy: rate constants and transition frequencies. J. Cell Biol. 107, 1437–1448 (1988)
Fygenson, D. K., Braun, E. & Libchaber, A. Phase diagram of microtubules. Phys. Rev. E 50, 1579–1588 (1994)
Hunt, A. J. & McIntosh, J. R. The dynamic behaviour of individual microtubules associated with chromosomes in vitro. Mol. Biol. Cell 9, 249–261 (1998)
Miranda, J. L., De Wulf, P., Sorger, P. & Harrison, S. C. The yeast DASH complex forms closed rings on microtubules. Nature Struct. Mol. Biol. 12, 138–143 (2005)
Nicklas, R. B. How cells get the right chromosomes. Science 275, 632–637 (1997)
McIntosh, J. R., Grishchuk, E. L. & West, R. R. Chromosome–microtubule interactions during mitosis. Annu. Rev. Cell. Dev. Biol. 18, 193–219 (2002)
Brouhard, G. J., Schek, H. T. III & Hunt, A. J. Advanced optical tweezers for the study of cellular and molecular biomechanics. IEEE Trans. Biomed. Eng. 50, 121–125 (2003)
Acknowledgements
We thank A. Hunt and G. J. Bouchard for sharing the plans for their laser tweezers; T. Perkins for advice, the quadrant photo detector design and programs to help with instrument calibration; H. Higuchi for tips on buffers; and members of McIntosh laboratory, and A. I. Vorobjev and G. P. Georgiev for help and support. V. Sarbash, T. Buxkemper and C. Bowen helped with building the trap. This work was supported in part by grants from the NIH to J.R.M., who is a Research Professor of the American Cancer Society.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains a description of our mathematical model of MT depolymerization, of MT-bead association, and of the laser trap. We calculate the position of the bead and shape of the MT when depolymerization is triggered via two scenarios described in the article. These calculations are compared with experimental results. The last section presents equations that were used to fit the experimental curves (DOC 43 kb)
Supplementary Figures
This file contains Supplementary Figures 1–3 and their legends. The figures display calculated profiles of the bead-MT system and the corresponding graphs. (PDF 499 kb)
Supplementary Video
Each frame shows the result of a calculation of the MT-bead configuration for “depolymerization” of one dimer layer via scenario 1. The bead moves when bending occurs in the bead–associated dimers and the dimers immediately downstream from it. All parameters are as in Supplementary Figure 2, except the attached dimers are # 10–14. The initial configuration is shown in gray. (MOV 132 kb)
Rights and permissions
About this article
Cite this article
Grishchuk, E., Molodtsov, M., Ataullakhanov, F. et al. Force production by disassembling microtubules. Nature 438, 384–388 (2005). https://doi.org/10.1038/nature04132
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04132
This article is cited by
-
Measuring and modeling forces generated by microtubules
Biophysical Reviews (2023)
-
Regulation of microtubule dynamics, mechanics and function through the growing tip
Nature Reviews Molecular Cell Biology (2021)
-
Mechanisms of microtubule dynamics and force generation examined with computational modeling and electron cryotomography
Nature Communications (2020)
-
Microtubule dynamics: an interplay of biochemistry and mechanics
Nature Reviews Molecular Cell Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.