Abstract
Deep-sea hydrothermal vents are important in global biogeochemical cycles, providing biological oases at the sea floor that are supported by the thermal and chemical flux from the Earth's interior. As hot, acidic and reduced hydrothermal fluids mix with cold, alkaline and oxygenated sea water, minerals precipitate to form porous sulphide–sulphate deposits. These structures provide microhabitats for a diversity of prokaryotes that exploit the geochemical and physical gradients in this dynamic ecosystem1. It has been proposed that fluid pH in the actively venting sulphide structures is generally low (pH < 4.5)2, yet no extreme thermoacidophile has been isolated from vent deposits. Culture-independent surveys based on ribosomal RNA genes from deep-sea hydrothermal deposits have identified a widespread euryarchaeotal lineage, DHVE2 (deep-sea hydrothermal vent euryarchaeotic 2)3,4,5,6. Despite the ubiquity and apparent deep-sea endemism of DHVE2, cultivation of this group has been unsuccessful and thus its metabolism remains a mystery. Here we report the isolation and cultivation of a member of the DHVE2 group, which is an obligate thermoacidophilic sulphur- or iron-reducing heterotroph capable of growing from pH 3.3 to 5.8 and between 55 and 75 °C. In addition, we demonstrate that this isolate constitutes up to 15% of the archaeal population, providing evidence that thermoacidophiles may be key players in the sulphur and iron cycling at deep-sea vents.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Reysenbach, A. L. & Shock, E. L. Merging genomes with geochemistry in hydrothermal ecosystems. Science 296, 1077–1082 (2002)
Tivey, M. K. in Subseafloor Biosphere at Mid-Ocean Ridges (eds Wilcock, W., Cary, C., DeLong, E., Kelley, D. & Baross, J.) 137–152 (Geophysical Monograph Series, American Geophysical Union, Washington DC, 2004)
Takai, K. & Horikoshi, K. Genetic diversity of archaea in deep-sea hydrothermal vent environments. Genetics 152, 1285–1297 (1999)
Reysenbach, A.-L., Longnecker, K. & Kirshtein, J. Novel bacterial and archaeal lineages from an in situ growth chamber deployed at a Mid-Atlantic Ridge hydrothermal vent. Appl. Environ. Microbiol. 66, 3798–3806 (2000)
Nercessian, O., Reysenbach, A.-L., Prieur, D. & Jeanthon, C. Compositional changes in archaeal communities associated with deep-sea hydrothermal vent samples from the East Pacific Rise (13°N). Environ. Microbiol. 5, 492–502 (2003)
Hoek, J., Banta, A. B., Hubler, R. F. & Reysenbach, A.-L. Microbial diversity of a deep-sea hydrothermal vent biofilm from Edmond vent field on the Central Indian Ridge. Geobiology 1, 119–127 (2003)
McCollom, T. M. & Shock, E. L. Geochemical constraints on chemolithoautotrophic metabolism by microorganisms in seafloor hydrothermal systems. Geochim. Cosmochim. Acta 61, 4375–4391 (1997)
Von Damm, K. L. in Physical, Chemical, Biological, and Geological Interactions within Seafloor Hydrothermal Systems (eds Humphris, S., Lupton, J., Mullineaux, L. & Zierenberg, R.) 222–247 (American Geophysical Union, Washington DC, 1995)
Le Bris, N., Zbinden, M. & Gaill, F. Processes controlling the physicochemical micro-environments associated with Pompeii worms. Deep Sea Res. 52, 1071–1083 (2005)
Takai, K. et al. Lebetimonas acidiphila gen. nov., sp. nov., a novel thermophilic, acidophilic, hydrogen-oxidizing chemolithoautotroph within the ‘Epsilonproteobacteria’, isolated from a deep-sea hydrothermal fumarole in the Mariana Arc. Int. J. Syst. Evol. Microbiol. 55, 183–189 (2005)
Prokofeva, M. et al. Cultivated anaerobic acidophilic/acidotolerant thermophiles from terrestrial and deep-sea hydrothermal habitats. Extremophiles 9, 437–448 (2005)
Kobayashi, T. in Bergeys Manual of Systematic Bacteriology (eds Boone, D. R., Castenholz, R. W. & Garrity, G. M.) 342–346 (Springer, New York, 2001)
Schrenk, M. O., Kelley, D. S., Delaney, J. R. & Baross, J. A. Incidence and diversity of microorganisms within the walls of an active deep-sea sulfide chimney. Appl. Environ. Microbiol. 69, 3580–3592 (2003)
Nakagawa, T. et al. Geomicrobiological exploration and characterization of a novel deep-sea hydrothermal system at the TOTO caldera in the Mariana volcanic arc. Environ. Microbiol. 8, 37–49 (2006)
Nakagawa, S. et al. Distribution, phylogenetic diversity and physiological characteristics of epsilon-Proteobacteria in a deep-sea hydrothermal field. Environ. Microbiol. 7, 1619–1632 (2005)
Takai, K., Komatsu, T., Inagari, F. & Horikoshi, K. Distribution of Archaea in a black smoker chimney structure. Appl. Environ. Microbiol. 67, 3618–3629 (2001)
Moussard, H. et al. Thermophilic lifestyle for an uncultured archaeon from hydrothermal vents: evidence from environmental genomics. Appl. Environ. Microbiol. 72, 2268–2271 (2006)
Ishibashi, J. et al. Expedition reveals changes in Lau Basin hydrothermal system. Eos 87, 13–17 (2006)
Inagaki, F. et al. Biogeographical distribution and diversity of microbes in methane hydrate-bearing deep marine sediments on the Pacific Ocean Margin. Proc. Natl Acad. Sci. USA 103, 2815–2820 (2006)
Segerer, A., Langworthy, T. A. & Stetter, K. O. Thermoplasma acidophilum and Thermoplasma volcanium sp. nov. from solfatara fields. Syst. Appl. Microbiol. 10, 161–171 (1988)
Langmuir, C. H. et al. Hydrothermal prospecting and petrological sampling in the Lau Basin: Background data for the integrated study site. Eos 85, B13A–0189 (2004)
Macalady, J. L. et al. Membrane monolayers in Ferroplasma spp.: a key to survival in acid. Extremophiles 8, 411–419 (2004)
Schleper, C. et al. Picrophilus gen. nov., fam. nov.: a novel aerobic, heterotrophic, thermoacidophilic genus and family comprising Archaea capable of growth around pH 0. J. Bacteriol. 177, 7050–7059 (1995)
Beveridge, T. J. Structures of Gram-negative cell walls and their derived membrane vesicles. J. Bacteriol. 7, 253–260 (1999)
Sleytr, U. B. & Beveridge, T. J. Bacterial S-layers. Trends Microbiol. 7, 253–260 (1999)
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004)
Olsen, G., Matsuda, H., Hagstrom, R. & Overbeek, R. FastDNAmL: a tool for construction of phylogenetic trees of DNA sequences using maximum likelihood. Comput. Appl. Biosci. 10, 41–48 (1994)
Hopmans, E. C. et al. Analysis of intact tetraether lipids in archaeal cell material and sediments by high performance liquid chromatography/atmospheric pressure chemical ionization mass spectrometry. Rapid Comm. Mass. Spectrom. 14, 585–589 (2000)
Segerer, A., Langworthy, T. A. & Stetter, K. O. Thermoplasma acidophilum and Thermoplasma volcanium sp. nov. from solfatara fields. Syst. Appl. Microbiol. 10, 161–171 (1988)
Acknowledgements
We thank the crew of the RV Melville, RV Atlantis and the DSROV Jason II and DSV Alvin teams for their assistance. This work is funded by the US National Science Foundation (A.-L.R., M.K.T, K.L.V.D.), a PSU Faculty Enhancement Award (A.-L. R.), the Natural Science and Engineering Research Council of Canada (NSERC), the US Department of Energy (T.J.B.), the US National Research Program, Water Resources Division, USGS, and NASA Exobiology (M.A.V.). We thank D. Moyles and R. Harris for technical help with TEM; P. Craddock for XRD; M. Baas and E. Hopmans for analytical assistance with the lipid analyses; I. Cozzarelli, M. Doughten and J. Jaeschke for the organic acid analysis; G. Sincerny, A. O'Neill and D. Decker for their assistance; and I. Ferrera for sharing qPCR results. J. Euzeby and M. Bartlett are acknowledged for assistance with naming the new archaeon. Author Contributions A.-L. R. and Y.L. isolated and characterized the acidophile; A.B.B. did all molecular phylogenetic and environmental DNA analyses; M.A.V. and J.D.K. did the qPCR analysis; T.J.B. did the ultrastructure analysis and interpretation; S.S. performed membrane lipid analysis; M.K.T. provided XRD data and geochemical interpretation; and K.L.V.D. provided samples, data on the sample locations and geochemical context. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The new sequences described in this manuscript have been deposited in GenBank, accession numbers DQ451875 (T469) and DQ451876 (T449). The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains the Supplementary Methods and Supplementary Tables 1–3, Supplementary Figures 1–3 and additional references. (PDF 302 kb)
Supplementary Figure 2
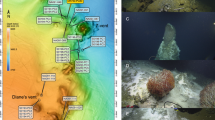
This is a high resolution version of Supplementary Figure 2. Example of a vent deposit from Mariner deep-sea vents (22°10.82'S, 176°36.09'W) and the outer accumulations of iron-oxide and sulphur biofilms typically where DHVE2 are detected. (PDF 304 kb)
Rights and permissions
About this article
Cite this article
Reysenbach, AL., Liu, Y., Banta, A. et al. A ubiquitous thermoacidophilic archaeon from deep-sea hydrothermal vents. Nature 442, 444–447 (2006). https://doi.org/10.1038/nature04921
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04921
This article is cited by
-
Identification of a deep-branching thermophilic clade sheds light on early bacterial evolution
Nature Communications (2023)
-
Global patterns of diversity and metabolism of microbial communities in deep-sea hydrothermal vent deposits
Microbiome (2022)
-
Moisture effects on the active prokaryotic communities in a saline soil unraveled by 18O-informed metagenomics
Journal of Soils and Sediments (2021)
-
New approaches for archaeal genome-guided cultivation
Science China Earth Sciences (2021)
-
Microorganisms from deep-sea hydrothermal vents
Marine Life Science & Technology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.