Abstract
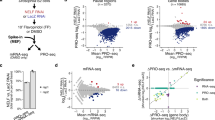
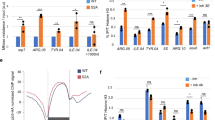
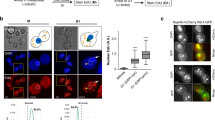
Chromosome condensation and the global repression of gene transcription1 are features of mitosis in most eukaryotes. The logic behind this phenomenon is that chromosome condensation prevents the activity of RNA polymerases. In budding yeast, however, transcription was proposed to be continuous during mitosis2. Here we show that Cdc14, a protein phosphatase required for nucleolar segregation3 and mitotic exit4, inhibits transcription of yeast ribosomal genes (rDNA) during anaphase. The phosphatase activity of Cdc14 is required for RNA polymerase I (Pol I) inhibition in vitro and in vivo. Moreover Cdc14-dependent inhibition involves nucleolar exclusion of Pol I subunits. We demonstrate that transcription inhibition is necessary for complete chromosome disjunction, because ribosomal RNA (rRNA) transcripts block condensin binding to rDNA, and show that bypassing the role of Cdc14 in nucleolar segregation requires in vivo degradation of nascent transcripts. Our results show that transcription interferes with chromosome condensation, not the reverse. We conclude that budding yeast, like most eukaryotes, inhibit Pol I transcription before segregation as a prerequisite for chromosome condensation and faithful genome separation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Taylor, J. Nucleic acid synthesis in relation to the cell division cycle. Ann. NY Acad. Sci. 90, 409–421 (1960)
Elliott, S. G. & McLaughlin, C. S. Regulation of RNA synthesis in yeast. III. Synthesis during the cell cycle. Mol. Gen. Genet. 169, 237–243 (1979)
Granot, D. & Snyder, M. Segregation of the nucleolus during mitosis in budding and fission yeast. Cell Motil. Cytoskeleton 20, 47–54 (1991)
Stegmeier, F. & Amon, A. Closing mitosis: the functions of the Cdc14 phosphatase and its regulation. Annu. Rev. Genet. 38, 203–232 (2004)
Lavoie, B. D., Hogan, E. & Koshland, D. In vivo requirements for rDNA chromosome condensation reveal two cell-cycle-regulated pathways for mitotic chromosome folding. Genes Dev. 18, 76–87 (2004)
Machin, F., Torres-Rosell, J., Jarmuz, A. & Aragon, L. Spindle-independent condensation-mediated segregation of yeast ribosomal DNA in late anaphase. J. Cell Biol. 168, 209–219 (2005)
Stegmeier, F., Visintin, R. & Amon, A. Separase, polo kinase, the kinetochore protein Slk19, and Spo12 function in a network that controls Cdc14 localization during early anaphase. Cell 108, 207–220 (2002)
Machin, F. et al. Transcription of ribosomal genes can cause nondisjunction. J. Cell Biol. 173, 893–903 (2006)
Tomson, B. N., D’Amours, D., Adamson, B. S., Aragon, L. & Amon, A. Ribosomal DNA transcription-dependent processes interfere with chromosome segregation. Mol. Cell. Biol. 26, 6239–6247 (2006)
Shou, W. et al. Exit from mitosis is triggered by Tem1-dependent release of the protein phosphatase Cdc14 from nucleolar RENT complex. Cell 97, 233–244 (1999)
Visintin, R., Hwang, E. S. & Amon, A. Cfi1 prevents premature exit from mitosis by anchoring Cdc14 phosphatase in the nucleolus. Nature 398, 818–823 (1999)
Azzam, R. et al. Phosphorylation by cyclin B-Cdk underlies release of mitotic exit activator Cdc14 from the nucleolus. Science 305, 516–519 (2004)
Shou, W. et al. Net1 stimulates RNA polymerase I transcription and regulates nucleolar structure independently of controlling mitotic exit. Mol. Cell 8, 45–55 (2001)
Hendrickson, C., Meyn, M. A., Morabito, L. & Holloway, S. L. The KEN box regulates Clb2 proteolysis in G1 and at the metaphase-to-anaphase transition. Curr. Biol. 11, 1781–1787 (2001)
Fath, S. et al. Differential roles of phosphorylation in the formation of transcriptional active RNA polymerase I. Proc. Natl Acad. Sci. USA 98, 14334–14339 (2001)
Gerber, J. et al. Site specific phosphorylation of yeast RNA polymerase I. Nucleic Acids Res. 36, 793–802 (2008)
D’Amours, D., Stegmeier, F. & Amon, A. Cdc14 and condensin control the dissolution of cohesin-independent chromosome linkages at repeated DNA. Cell 117, 455–469 (2004)
Sullivan, M., Higuchi, T., Katis, V. L. & Uhlmann, F. Cdc14 phosphatase induces rDNA condensation and resolves cohesin-independent cohesion during budding yeast anaphase. Cell 117, 471–482 (2004)
Torres-Rosell, J., Machin, F., Jarmuz, A. & Aragon, L. Nucleolar segregation lags behind the rest of the genome and requires Cdc14p activation by the FEAR network. Cell Cycle 3, 496–502 (2004)
Wang, B. D., Yong-Gonzalez, V. & Strunnikov, A. V. Cdc14p/FEAR pathway controls segregation of nucleolus in S. cerevisiae by facilitating condensin targeting to rDNA chromatin in anaphase. Cell Cycle 3, 960–967 (2004)
Wang, B. D., Butylin, P. & Strunnikov, A. Condensin function in mitotic nucleolar segregation is regulated by rDNA transcription. Cell Cycle 5, 2260–2267 (2006)
Fujii, T., Yamaoka, H., Gomi, K., Kitamoto, K. & Kumagai, C. Cloning and nucleotide sequence of the ribonuclease T1 gene (rntA) from Aspergillus oryzae and its expression in Saccharomyces cerevisiae and Aspergillus oryzae. Biosci. Biotechnol. Biochem. 59, 1869–1874 (1995)
D’Ambrosio, C., Kelly, G., Shirahige, K. & Uhlmann, F. Condensin-dependent rDNA decatenation introduces a temporal pattern to chromosome segregation. Curr. Biol. 18, 1084–1089 (2008)
Freeman, L., Aragon-Alcaide, L. & Strunnikov, A. The condensin complex governs chromosome condensation and mitotic transmission of rDNA. J. Cell Biol. 149, 811–824 (2000)
Baxter, J. & Diffley, J. F. Topoisomerase II inactivation prevents the completion of DNA replication in budding yeast. Mol. Cell 30, 790–802 (2008)
Bhalla, N., Biggins, S. & Murray, A. W. Mutation of YCS4, a budding yeast condensin subunit, affects mitotic and nonmitotic chromosome behavior. Mol. Biol. Cell 13, 632–645 (2002)
Nelson, J. D., Denisenko, O. & Bomsztyk, K. Protocol for the fast chromatin immunoprecipitation (ChIP) method. Nature Protocols 1, 179–185 (2006)
Keener, J., Josaitis, C. A., Dodd, J. A. & Nomura, M. Reconstitution of yeast RNA polymerase I transcription in vitro from purified components. TATA-binding protein is not required for basal transcription. J. Biol. Chem. 273, 33795–33802 (1998)
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004)
Powers, T. & Walter, P. Regulation of ribosome biogenesis by the rapamycin-sensitive TOR-signaling pathway in Saccharomyces cerevisiae. Mol. Biol. Cell 10, 987–1000 (1999)
Acknowledgements
We are grateful to A. Amon, D. Morgan, K. Kitamoto and J. Van Etten for providing reagents, and M. Nomura for advice. We thank members of the Aragon laboratory for discussions. We acknowledge grant support from UAB HSF-GEF to D.A.S. and from ‘La Junta de Extremadura’ to A.C.-B. The Aragon laboratory is supported by the Medical Research Council of the UK.
Author contributions F.M. made the initial observation that transcription impairs rDNA segregation in the absence of Cdc14 function. All experiments shown were performed by A.C.-B., except ChIPs which were performed by M.M.-S. and in vitro phosphatase assays which were performed by D.A.S. A.J. cloned the RNAse constructs. H.T. provided a battery of Rpa43 mutants. All experiments shown were conceived by L.A. and the person performing the specific experiment. L.A. wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-10 with Legends and Supplementary Table 1 (PDF 5109 kb)
Rights and permissions
About this article
Cite this article
Clemente-Blanco, A., Mayán-Santos, M., Schneider, D. et al. Cdc14 inhibits transcription by RNA polymerase I during anaphase. Nature 458, 219–222 (2009). https://doi.org/10.1038/nature07652
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07652
This article is cited by
-
Cdc14 phosphatase counteracts Cdk-dependent Dna2 phosphorylation to inhibit resection during recombinational DNA repair
Nature Communications (2023)
-
Budding yeast complete DNA synthesis after chromosome segregation begins
Nature Communications (2020)
-
Histone H2A insufficiency causes chromosomal segregation defects due to anaphase chromosome bridge formation at rDNA repeats in fission yeast
Scientific Reports (2019)
-
Condensin action and compaction
Current Genetics (2019)
-
SMC complexes differentially compact mitotic chromosomes according to genomic context
Nature Cell Biology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.