Abstract
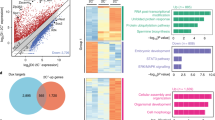
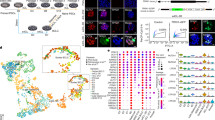
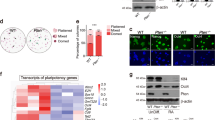
Induced pluripotent stem cells (iPSCs) have been generated by enforced expression of defined sets of transcription factors in somatic cells. It remains controversial whether iPSCs are molecularly and functionally equivalent to blastocyst-derived embryonic stem (ES) cells. By comparing genetically identical mouse ES cells and iPSCs, we show here that their overall messenger RNA and microRNA expression patterns are indistinguishable with the exception of a few transcripts encoded within the imprinted Dlk1–Dio3 gene cluster on chromosome 12qF1, which were aberrantly silenced in most of the iPSC clones. Consistent with a developmental role of the Dlk1–Dio3 gene cluster, these iPSC clones contributed poorly to chimaeras and failed to support the development of entirely iPSC-derived animals (‘all-iPSC mice’). In contrast, iPSC clones with normal expression of the Dlk1–Dio3 cluster contributed to high-grade chimaeras and generated viable all-iPSC mice. Notably, treatment of an iPSC clone that had silenced Dlk1–Dio3 with a histone deacetylase inhibitor reactivated the locus and rescued its ability to support full-term development of all-iPSC mice. Thus, the expression state of a single imprinted gene cluster seems to distinguish most murine iPSCs from ES cells and allows for the prospective identification of iPSC clones that have the full development potential of ES cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Hochedlinger, K. & Plath, K. Epigenetic reprogramming and induced pluripotency. Development 136, 509–523 (2009)
Maherali, N. et al. A high-efficiency system for the generation and study of human induced pluripotent stem cells. Cell Stem Cell 3, 340–345 (2008)
Mikkelsen, T. S. et al. Dissecting direct reprogramming through integrative genomic analysis. Nature 454, 49–55 (2008)
Wernig, M. et al. In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 448, 318–324 (2007)
Okita, K., Ichisaka, T. & Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nature 448, 313–317 (2007)
Boland, M. J. et al. Adult mice generated from induced pluripotent stem cells. Nature 461, 91–94 (2009)
Zhao, X. Y. et al. iPS cells produce viable mice through tetraploid complementation. Nature 461, 86–90 (2009)
Kang, L., Wang, J., Zhang, Y., Kou, Z. & Gao, S. iPS cells can support full-term development of tetraploid blastocyst-complemented embryos. Cell Stem Cell 5, 135–138 (2009)
Nagy, A. et al. Embryonic stem cells alone are able to support fetal development in the mouse. Development 110, 815–821 (1990)
Eggan, K. et al. Hybrid vigor, fetal overgrowth, and viability of mice derived by nuclear cloning and tetraploid embryo complementation. Proc. Natl Acad. Sci. USA 98, 6209–6214 (2001)
Kim, J. B. et al. Pluripotent stem cells induced from adult neural stem cells by reprogramming with two factors. Nature 454, 646–650 (2008)
Hanna, J. et al. Direct reprogramming of terminally differentiated mature B lymphocytes to pluripotency. Cell 133, 250–264 (2008)
Meissner, A., Wernig, M. & Jaenisch, R. Direct reprogramming of genetically unmodified fibroblasts into pluripotent stem cells. Nature Biotechnol. 25, 1177–1181 (2007)
Chin, M. H. et al. Induced pluripotent stem cells and embryonic stem cells are distinguished by gene expression signatures. Cell Stem Cell 5, 111–123 (2009)
Marchetto, M. C. et al. Transcriptional signature and memory retention of human-induced pluripotent stem cells. PLoS One 4, e7076 (2009)
Wilson, K. D. et al. MicroRNA profiling of human-induced pluripotent stem cells. Stem Cells Dev. 18, 749–758 (2009)
Pick, M. et al. Clone- and gene-specific aberrations of parental imprinting in human induced pluripotent stem cells. Stem Cells 27, 2686–2690 (2009)
Doi, A. et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nature Genet. 41, 1350–1353 (2009)
Deng, J. et al. Targeted bisulfite sequencing reveals changes in DNA methylation associated with nuclear reprogramming. Nature Biotechnol. 27, 353–360 (2009)
Brambrink, T., Hochedlinger, K., Bell, G. & Jaenisch, R. ES cells derived from cloned and fertilized blastocysts are transcriptionally and functionally indistinguishable. Proc. Natl Acad. Sci. USA 103, 933–938 (2006)
Wakayama, S. et al. Equivalency of nuclear transfer-derived embryonic stem cells to those derived from fertilized mouse blastocysts. Stem Cells 24, 2023–2033 (2006)
Soldner, F. et al. Parkinson’s disease patient-derived induced pluripotent stem cells free of viral reprogramming factors. Cell 136, 964–977 (2009)
Stadtfeld, M., Maherali, N., Borkent, M. & Hochedlinger, K. A reprogrammable mouse strain from gene-targeted embryonic stem cells. Nature Methods 7, 53–55 (2009)
Sommer, C. A. et al. Induced pluripotent stem cell generation using a single lentiviral stem cell cassette. Stem Cells 27, 543–549 (2009)
Beard, C., Hochedlinger, K., Plath, K., Wutz, A. & Jaenisch, R. Efficient method to generate single-copy transgenic mice by site-specific integration in embryonic stem cells. Genesis 44, 23–28 (2006)
da Rocha, S. T., Edwards, C. A., Ito, M., Ogata, T. & Ferguson-Smith, A. C. Genomic imprinting at the mammalian Dlk1-Dio3 domain. Trends Genet. 24, 306–316 (2008)
Humpherys, D. et al. Epigenetic instability in ES cells and cloned mice. Science 293, 95–97 (2001)
Seitz, H. et al. Imprinted microRNA genes transcribed antisense to a reciprocally imprinted retrotransposon-like gene. Nature Genet. 34, 261–262 (2003)
Seitz, H. et al. A large imprinted microRNA gene cluster at the mouse Dlk1-Gtl2 domain. Genome Res. 14, 1741–1748 (2004)
Takahashi, N. et al. Deletion of Gtl2, imprinted non-coding RNA, with its differentially methylated region induces lethal parent-origin-dependent defects in mice. Hum. Mol. Genet. 18, 1879–1888 (2009)
Lin, S. P. et al. Differential regulation of imprinting in the murine embryo and placenta by the Dlk1-Dio3 imprinting control region. Development 134, 417–426 (2007)
Steshina, E. Y. et al. Loss of imprinting at the Dlk1-Gtl2 locus caused by insertional mutagenesis in the Gtl2 5′ region. BMC Genet. 7, 44 (2006)
da Rocha, S. T. et al. Gene dosage effects of the imprinted delta-like homologue 1 (dlk1/pref1) in development: implications for the evolution of imprinting. PLoS Genet. 5, e1000392 (2009)
Lin, S. P. et al. Asymmetric regulation of imprinting on the maternal and paternal chromosomes at the Dlk1-Gtl2 imprinted cluster on mouse chromosome 12. Nature Genet. 35, 97–102 (2003)
Carr, M. S., Yevtodiyenko, A., Schmidt, C. L. & Schmidt, J. V. Allele-specific histone modifications regulate expression of the Dlk1-Gtl2 imprinted domain. Genomics 89, 280–290 (2007)
Dean, W. et al. Altered imprinted gene methylation and expression in completely ES cell-derived mouse fetuses: association with aberrant phenotypes. Development 125, 2273–2282 (1998)
Tevendale, M., Watkins, M., Rasberry, C., Cattanach, B. & Ferguson-Smith, A. C. Analysis of mouse conceptuses with uniparental duplication/deficiency for distal chromosome 12: comparison with chromosome 12 uniparental disomy and implications for genomic imprinting. Cytogenet. Genome Res. 113, 215–222 (2006)
Schmidt, J. V., Matteson, P. G., Jones, B. K., Guan, X. J. & Tilghman, S. M. The Dlk1 and Gtl2 genes are linked and reciprocally imprinted. Genes Dev. 14, 1997–2002 (2000)
Eggan, K. et al. Mice cloned from olfactory sensory neurons. Nature 428, 44–49 (2004)
Li, J., Ishii, T., Feinstein, P. & Mombaerts, P. Odorant receptor gene choice is reset by nuclear transfer from mouse olfactory sensory neurons. Nature 428, 393–399 (2004)
Ono, Y. & Kono, T. Irreversible barrier to the reprogramming of donor cells in cloning with mouse embryos and embryonic stem cells. Biol. Reprod. 75, 210–216 (2006)
Stadtfeld, M., Nagaya, M., Utikal, J., Weir, G. & Hochedlinger, K. Induced pluripotent stem cells generated without viral integration. Science 322, 945–949 (2008)
Yu, J. et al. Human induced pluripotent stem cells free of vector and transgene sequences. Science 324, 797–801 (2009)
Navarro, P. et al. Molecular coupling of Xist regulation and pluripotency. Science 321, 1693–1695 (2008)
Donohoe, M. E., Silva, S. S., Pinter, S. F., Xu, N. & Lee, J. T. The pluripotency factor Oct4 interacts with Ctcf and also controls X-chromosome pairing and counting. Nature 460, 128–132 (2009)
Eminli, S. et al. Differentiation stage determines potential of hematopoietic cells for reprogramming into induced pluripotent stem cells. Nature Genet. 41, 968–976 (2009)
Hochedlinger, K., Yamada, Y., Beard, C. & Jaenisch, R. Ectopic expression of Oct-4 blocks progenitor-cell differentiation and causes dysplasia in epithelial tissues. Cell 121, 465–477 (2005)
Coser, K. R. et al. Global analysis of ligand sensitivity of estrogen inducible and suppressible genes in MCF7/BUS breast cancer cells by DNA microarray. Proc. Natl Acad. Sci. USA 100, 13994–13999 (2003)
Eisen, M. B., Spellman, P. T., Brown, P. O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863–14868 (1998)
Acknowledgements
We are grateful to H. Arnold for assistance with GeneSifter analysis; K. Coser, K. Claycomb and P. August for technical support on Affymetrix expression profiling; S. Sato and M. Machida for technical assistance; V. Greco for advice on keratinocyte isolation; and S. Schubert for advice on miRNA isolation. We thank members of the Hochedlinger laboratory for helpful suggestions and A. Umezawa for discussions and support. M.S. was supported by a postdoctoral fellowship from the Schering Foundation, E.A. was supported by a Jane Coffin Childs postdoctoral fellowship and K.H. was supported by a NIH Director’s Innovator Award and by funds provided by the Harvard Stem Cell Institute, MGH and HHMI.
Author information
Authors and Affiliations
Contributions
Author Contributions M.S., E.A. and K.H. conceived the ideas for this study, designed and analysed experiments and wrote the manuscript. M.S. derived iPSC lines, conducted in vitro differentiation assays and performed expression array analysis. E.A. conducted qPCR analyses, in situ hybridizations and chromatin immunoprecipitations. H.A. and A.F. performed nuclear transfer experiments. P.F. did blastocyst injections. T.S. performed microarray experiments and analyses. S.N. and T.K. provided important study materials.
Corresponding author
Ethics declarations
Competing interests
K.H. is on the advisory board of iPierian.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-10 with legends and Supplementary Tables 1, 4, 6 and 7. (PDF 5027 kb)
Supplementary Table 2
This table shows expression levels of imprinted genes in ES cells and iPSCs. (XLS 41 kb)
Supplementary Table 3
This table shows the global miRNA expression in ES cells and iPSCs. (XLS 230 kb)
Supplementary Table 5
This table shows the global miRNA expression of 4n complementation-competent and non-competent iPSCs. (XLS 189 kb)
Pease note that the descriptions for Tables 2, 3 and 5 were updated on 2 May 2010
Rights and permissions
About this article
Cite this article
Stadtfeld, M., Apostolou, E., Akutsu, H. et al. Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature 465, 175–181 (2010). https://doi.org/10.1038/nature09017
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09017
This article is cited by
-
Modelling in vitro gametogenesis using induced pluripotent stem cells: a review
Cell Regeneration (2023)
-
Functional evolutionary convergence of long noncoding RNAs involved in embryonic development
Communications Biology (2023)
-
Time-resolved Small-RNA Sequencing Identifies MicroRNAs Critical for Formation of Embryonic Stem Cells from the Inner Cell Mass of Mouse Embryos
Stem Cell Reviews and Reports (2023)
-
Maintenance of methylation profile in imprinting control regions in human induced pluripotent stem cells
Clinical Epigenetics (2022)
-
SETD5 modulates homeostasis of hematopoietic stem cells by mediating RNA Polymerase II pausing in cooperation with HCF-1
Leukemia (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.