Abstract
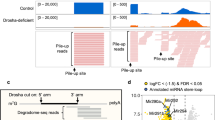
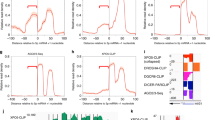
The nucleolytic activity of animal Argonaute proteins is deeply conserved, despite its having no obvious role in microRNA-directed gene regulation. In mice, Ago2 (also known as Eif2c2) is uniquely required for viability, and only this family member retains catalytic competence. To investigate the evolutionary pressure to conserve Argonaute enzymatic activity, we engineered a mouse with catalytically inactive Ago2 alleles. Homozygous mutants died shortly after birth with an obvious anaemia. Examination of microRNAs and their potential targets revealed a loss of miR-451, a small RNA important for erythropoiesis. Though this microRNA is processed by Drosha (also known as Rnasen), its maturation does not require Dicer. Instead, the pre-miRNA becomes loaded into Ago and is cleaved by the Ago catalytic centre to generate an intermediate 3′ end, which is then further trimmed. Our findings link the conservation of Argonaute catalysis to a conserved mechanism of microRNA biogenesis that is important for vertebrate development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hutvagner, G. & Simard, M. J. Argonaute proteins: key players in RNA silencing. Nature Rev. Mol. Cell Biol. 9, 22–32 (2008)
Joshua-Tor, L. The Argonautes. Cold Spring Harb. Symp. Quant. Biol. 71, 67–72 (2006)
Elbashir, S. M., Lendeckel, W. & Tuschl, T. RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev. 15, 188–200 (2001)
Elbashir, S. M., Martinez, J., Patkaniowska, A., Lendeckel, W. & Tuschl, T. Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J. 20, 6877–6888 (2001)
Yuan, Y. R. et al. Crystal structure of A. aeolicus argonaute, a site-specific DNA-guided endoribonuclease, provides insights into RISC-mediated mRNA cleavage. Mol. Cell 19, 405–419 (2005)
Martinez, J. & Tuschl, T. RISC is a 5′ phosphomonoester-producing RNA endonuclease. Genes Dev. 18, 975–980 (2004)
Schwarz, D. S., Tomari, Y. & Zamore, P. D. The RNA-induced silencing complex is a Mg2+-dependent endonuclease. Curr. Biol. 14, 787–791 (2004)
Malone, C. D. & Hannon, G. J. Small RNAs as guardians of the genome. Cell 136, 656–668 (2009)
Yigit, E. et al. Analysis of the C. elegans Argonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127, 747–757 (2006)
Bohmert, K. et al. AGO1 defines a novel locus of Arabidopsis controlling leaf development. EMBO J. 17, 170–180 (1998)
Baumberger, N. & Baulcombe, D. C. Arabidopsis ARGONAUTE1 is an RNA Slicer that selectively recruits microRNAs and short interfering RNAs. Proc. Natl Acad. Sci. USA 102, 11928–11933 (2005)
Qi, Y., Denli, A. M. & Hannon, G. J. Biochemical specialization within Arabidopsis RNA silencing pathways. Mol. Cell 19, 421–428 (2005)
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009)
Yekta, S., Shih, I. H. & Bartel, D. P. MicroRNA-directed cleavage of HOXB8 mRNA. Science 304, 594–596 (2004)
Davis, E. et al. RNAi-mediated allelic trans-interaction at the imprinted Rtl1/Peg11 locus. Curr. Biol. 15, 743–749 (2005)
Harfe, B. D., McManus, M. T., Mansfield, J. H., Hornstein, E. & Tabin, C. J. The RNaseIII enzyme Dicer is required for morphogenesis but not patterning of the vertebrate limb. Proc. Natl Acad. Sci. USA 102, 10898–10903 (2005)
Sekita, Y. et al. Role of retrotransposon-derived imprinted gene, Rtl1, in the feto-maternal interface of mouse placenta. Nature Genet. 40, 243–248 (2008)
Hornstein, E. et al. The microRNA miR-196 acts upstream of Hoxb8 and Shh in limb development. Nature 438, 671–674 (2005)
Tolia, N. H. & Joshua-Tor, L. Slicer and the argonautes. Nature Chem. Biol. 3, 36–43 (2007)
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004)
Rivas, F. V. et al. Purified Argonaute2 and an siRNA form recombinant human RISC. Nature Struct. Mol. Biol. 12, 340–349 (2005)
Song, J. J., Smith, S. K., Hannon, G. J. & Joshua-Tor, L. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305, 1434–1437 (2004)
Azuma-Mukai, A. et al. Characterization of endogenous human Argonautes and their miRNA partners in RNA silencing. Proc. Natl Acad. Sci. USA 105, 7964–7969 (2008)
Ender, C. et al. A human snoRNA with microRNA-like functions. Mol. Cell 32, 519–528 (2008)
Babiarz, J. E., Ruby, J. G., Wang, Y., Bartel, D. P. & Blelloch, R. Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes Dev. 22, 2773–2785 (2008)
Tam, O. H. et al. Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature 453, 534–538 (2008)
Watanabe, T. et al. Endogenous siRNAs from naturally formed dsRNAs regulate transcripts in mouse oocytes. Nature 453, 539–543 (2008)
Kaneda, M., Tang, F., O’Carroll, D., Lao, K. & Surani, M. A. Essential role for Argonaute2 protein in mouse oogenesis. Epigenetics Chromatin 2, 9 (2009)
Ma, J. et al. MicroRNA activity is suppressed in mouse oocytes. Curr. Biol. 20, 265–270 (2010)
Murchison, E. P. et al. Critical roles for Dicer in the female germline. Genes Dev. 21, 682–693 (2007)
Suh, N. et al. MicroRNA function is globally suppressed in mouse oocytes and early embryos. Curr. Biol. 20, 271–277 (2010)
Alisch, R. S., Jin, P., Epstein, M., Caspary, T. & Warren, S. T. Argonaute2 is essential for mammalian gastrulation and proper mesoderm formation. PLoS Genet. 3, e227 (2007)
Morita, S. et al. One Argonaute family member, Eif2c2 (Ago2), is essential for development and appears not to be involved in DNA methylation. Genomics 89, 687–696 (2007)
Sasaki, T., Shiohama, A., Minoshima, S. & Shimizu, N. Identification of eight members of the Argonaute family in the human genome. Genomics 82, 323–330 (2003)
Rossant, J. & Cross, J. C. in Mouse development: Pattering, Morphogenesis and Organogenesis (eds Rossant, J. & Tam, P. P. L.) 155–180 (Academic Press, 2002)
Rossant, J. & Cross, J. C. Placental development: lessons from mouse mutants. Nature Rev. Genet. 2, 538–548 (2001)
Papapetrou, E. P., Korkola, J. E. & Sadelain, M. A genetic strategy for single and combinatorial analysis of miRNA function in mammalian hematopoietic stem cells. Stem Cells 28, 287–296 (2009)
Dore, L. C. et al. A GATA-1-regulated microRNA locus essential for erythropoiesis. Proc. Natl Acad. Sci. USA 105, 3333–3338 (2008)
Kim, V. N., Han, J. & Siomi, M. C. Biogenesis of small RNAs in animals. Nature Rev. Mol. Cell Biol. 10, 126–139 (2009)
Siolas, D. et al. Synthetic shRNAs as potent RNAi triggers. Nature Biotechnol. 23, 227–231 (2004)
Chendrimada, T. P. et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436, 740–744 (2005)
Wang, H. W. et al. Structural insights into RNA processing by the human RISC-loading complex. Nature Struct. Mol. Biol. 16, 1148–1153 (2009)
Song, J. J. et al. The crystal structure of the Argonaute2 PAZ domain reveals an RNA binding motif in RNAi effector complexes. Nature Struct. Biol. 10, 1026–1032 (2003)
Wang, Y., Sheng, G., Juranek, S., Tuschl, T. & Patel, D. J. Structure of the guide-strand-containing argonaute silencing complex. Nature 456, 209–213 (2008)
Diederichs, S. & Haber, D. A. Dual role for argonautes in microRNA processing and posttranscriptional regulation of microRNA expression. Cell 131, 1097–1108 (2007)
O’Carroll, D. et al. A Slicer-independent role for Argonaute 2 in hematopoiesis and the microRNA pathway. Genes Dev. 21, 1999–2004 (2007)
Bandres, E. et al. microRNA-451 regulates macrophage migration inhibitory factor production and proliferation of gastrointestinal cancer cells. Clin. Cancer Res. 15, 2281–2290 (2009)
Pfeffer, S. et al. Identification of microRNAs of the herpesvirus family. Nature Methods 2, 269–276 (2005)
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003)
Nagy, A., Gertsenstein, M., Vintersten, K. & Behringer, R. Manipulating the Mouse Embryo: A Laboratory Manual 3rd edn (Cold Spring Harbor Press, 2003)
Nagy, A. & Rossant, J. in Gene Targeting: A Practical Approach 2nd edn (ed. Joyner, A. L.) 189–192 (Oxford Univ. Press, 2000)
Socolovsky, M. et al. Ineffective erythropoiesis in Stat5a-/-5b-/- mice due to decreased survival of early erythroblasts. Blood 98, 3261–3273 (2001)
Liu, Y. et al. Suppression of Fas-FasL coexpression by erythropoietin mediates erythroblast expansion during the erythropoietic stress response in vivo . Blood 108, 123–133 (2006)
Aravin, A. & Tuschl, T. Identification and characterization of small RNAs involved in RNA silencing. FEBS Lett. 579, 5830–5840 (2005)
Denli, A. M., Tops, B. B., Plasterk, R. H., Ketting, R. F. & Hannon, G. J. Processing of primary microRNAs by the Microprocessor complex. Nature 432, 231–235 (2004)
Acknowledgements
We thank M. Mosquera, S. Y. Kim, D. Frendewey and A. Economides for help with generating mutants and caring for animals. A. Nagy, M. Shen, J. Murn, V. Vagin and A. Aravin provided helpful discussion. N. Kim, D. Littman, I. Ibarra and E. Wagenblast provided critical reagents, and O. Tam, R. Sachidanandam, Z. Xuan, D. McCombie and M. Rooks provided support for generation and analysis of deep sequencing data. This work was supported by grants from the NIH and by a gift from K. W. Davis.
Author Contributions S.C. and G.J.H. planned experiments, interpreted data and wrote the paper. S.C. and C.O.D.S. performed studies, and M.M.W.C. provided critical unpublished reagents and discussion.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1 and Supplementary Figures S1-S8 with legends. (PDF 6132 kb)
Rights and permissions
About this article
Cite this article
Cheloufi, S., Dos Santos, C., Chong, M. et al. A dicer-independent miRNA biogenesis pathway that requires Ago catalysis. Nature 465, 584–589 (2010). https://doi.org/10.1038/nature09092
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09092
This article is cited by
-
A catalytic function for mammalian Argonautes
Nature Reviews Molecular Cell Biology (2024)
-
Unveiling the world of bee microRNAs: computational identification and characterization of pathway genes, conserved microRNAs, and their targets
International Journal of Tropical Insect Science (2024)
-
CD95/Fas ligand mRNA is toxic to cells through more than one mechanism
Molecular Biomedicine (2023)
-
p53 and HuR combinatorially control the biphasic dynamics of microRNA-125b in response to genotoxic stress
Communications Biology (2023)
-
microRNAs in action: biogenesis, function and regulation
Nature Reviews Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.