Abstract
Members of the Wiskott–Aldrich syndrome protein (WASP) family control cytoskeletal dynamics by promoting actin filament nucleation with the Arp2/3 complex. The WASP relative WAVE regulates lamellipodia formation within a 400-kilodalton, hetero-pentameric WAVE regulatory complex (WRC). The WRC is inactive towards the Arp2/3 complex, but can be stimulated by the Rac GTPase, kinases and phosphatidylinositols. Here we report the 2.3-ångstrom crystal structure of the WRC and complementary mechanistic analyses. The structure shows that the activity-bearing VCA motif of WAVE is sequestered by a combination of intramolecular and intermolecular contacts within the WRC. Rac and kinases appear to destabilize a WRC element that is necessary for VCA sequestration, suggesting the way in which these signals stimulate WRC activity towards the Arp2/3 complex. The spatial proximity of the Rac binding site and the large basic surface of the WRC suggests how the GTPase and phospholipids could cooperatively recruit the complex to membranes.
Similar content being viewed by others
Main
Members of the WASP family are central to the control of cellular actin dynamics1,2,3. These proteins receive information from multiple signalling pathways and respond by promoting the actin nucleating activity of the ubiquitous Arp2/3 complex. In this way, WASP proteins control actin assembly spatially and temporally in processes including cell migration, polarization, adhesion and vesicle trafficking.
The WASP family is defined by a conserved C-terminal VCA motif (for the verprolin-homology, central and acidic regions), which binds and activates the Arp2/3 complex1,3. This element must be tightly regulated to ensure proper spatial and temporal control over actin assembly. In the best-understood family members, WASP and N-WASP, the VCA is autoinhibited by intramolecular interactions with a regulatory element termed the GTPase binding domain (GBD)4. Various ligands can bind to WASP/N-WASP simultaneously, and destabilize GBD–VCA contacts, leading to activation1,3. Activation of all family members appears to be restricted to membranes. Superimposed on allosteric control and coupled with membrane recruitment, the activity of WASP proteins can be substantially increased by dimerization, or more generally oligomerization/clustering at membranes5.
Although WASP and N-WASP can exist independently in cells, WAVE proteins are constitutively associated with four additional proteins inside cells: Sra1/Cyfip1, Nap1/Hem-2, Abi and HSPC300 (refs 6 and 7). The components of this ∼400-kDa pentamer, termed the WRC, have all been implicated in control of Arp2/3-complex-mediated actin assembly in a wide range of systems1,8. Sra1/Cyfip1 also has a distinct role in translational control9,10. WAVE proteins lack an inhibitory GBD, and the mechanism of VCA regulation within the WRC is not known. The WRC can be activated by a wide range of stimuli, including the Rac GTPase and acidic phospholipids6,11,12,13,14, which appear to act cooperatively at the plasma membrane12,14. Furthermore, components of the WRC can be phosphorylated at numerous positions (http://www.phosphosite.org/proteinAction.do?id=7256&showAllSites=true), with some modifications enhancing signalling activity14,15,16,17,18,19. The mechanisms by which ligands act individually and cooperatively to recruit and activate the WRC are not known.
Here we report the 2.3-Å crystal structure of the WRC and complementary biochemical and cell biological analyses. The combined data reveal how the WAVE VCA is inhibited within the complex and provide plausible mechanisms for WRC activation by Rac and phosphorylation, and for cooperative membrane recruitment by Rac and phospholipids. Our analyses provide an integrated picture of how the WRC orchestrates multiple signalling pathways to control actin polymerization at the plasma membrane.
Overall structure of the WRC
To facilitate crystallization of the WRC we genetically deleted the C-terminal proline-rich region and SH3 domain of Abi2, and replaced the proline-rich region of WAVE1 with an 18-residue linker. Sra1, Nap1 and HSPC300 were full-length. The resulting miniWRC is inactive towards the Arp2/3 complex but can be stimulated by Rac1-GMPPNP13.
Crystals of miniWRC contained one complex in the asymmetric unit and diffracted to 2.3 Å at a synchrotron light source. Phases were obtained by multiple isomorphous replacement with anomalous scattering (MIRAS) using preparations containing selenomethionine-labelled Sra1 and Nap1 (Supplementary Table 1). The final structure was refined to Rwork/Rfree = 18.8%/23.7%. MiniWRC has an elongated shape with approximate dimensions of 200 Å × 110 Å × 80 Å (Fig. 1). Two-dimensional class averages from electron micrographs of negatively stained miniWRC and full-length WRC are indistinguishable, and have dimensions similar to the crystal structure (Supplementary Fig. 1). The structure of miniWRC is thus probably a faithful representation of the structured elements of the WRC.
a, Stereo view of miniWRC. Sra1, Nap1, WAVE1, Abi2 and HSPC300 are green, blue, magenta, orange and yellow, respectively. The A-region (residues 545–559), α6–V region linker (residues 185–485) and the sequence connecting V- and C-helices (residues 519–528) are not observed in the electron density. The latter two are shown as dashed lines. b, 180° rotation about a horizontal axis from a. The polybasic region and the proposed Rac1 and eIF4E binding sites are indicated.
MiniWRC can be delineated into two subcomplexes: an Sra1:Nap1 dimer and a WAVE1:Abi2:HSPC300 trimer (Fig. 1). Sra1 and Nap1 have homologous structures (see below) and interact extensively to create an elongated pseudo-symmetric dimer, which forms a platform for the trimer (Supplementary Fig. 2). The amino (N)-terminal helix of Sra1 links to the rest of the complex through a flexible sequence that lacks electron density (residues 23–56), and contacts an adjacent molecule in the crystal lattice (Supplementary Fig. 3). The trimer contacts the Sra1:Nap1 dimer in a tripartite manner. A long four-helix bundle created by a helix from HSPC300 (residues 14–68), two helices from Abi2 (residues 1–39 and 43–112) and a helix from WAVE1 (residues 26–81) contacts Sra1 extensively and is aligned roughly parallel to the long axis of the dimer (Supplementary Figs 4 and 5). The most extensive contacts are made by HSPC300, which is sandwiched between Sra1 and Abi2:WAVE1 across the entire length of its helix. The ‘homeo-domain homologous region’ of Abi2 (residues 112–155)20 adopts an extended conformation running around the rim of a large cavity on Nap1 (Supplementary Fig. 6). The carboxy (C) terminus of WAVE1, including the V- and the C-regions, forms an irregular, loosely packed chain that lies against a concave surface of Sra1 adjacent to the long side of the four-helix bundle. As detailed below, interactions of elements in the C terminus of WAVE1 with Sra1 and each other are central to the regulation of WRC activity.
The structure reveals that Sra1 and Nap1 have the same domain organization; their coordinates can be superimposed with a root mean square deviation (r.m.s.d.) of 6.9 Å for 681 Cα atoms with Dali Z-score 17.9 (Supplementary Fig. 7)21. Thus, they belong to the same protein family despite their low sequence identity (13%). Homology between Sra1 and Nap1 is also supported by HHpred22, which showed additional human members of this family that are similar in size: KIAA1033/SWIP and Strumpellin, with similarity extending over their entire lengths. When analysed pairwise, SWIP is more similar to Sra1 and Strumpellin is more similar to Nap1. We and others recently reported that SWIP and Strumpellin form a pentameric complex (SHRC, for WASH Regulatory Complex) containing the proteins CCDC53 and Fam21, and another WASP family member, WASH23,24,25. Within the SHRC the WASH VCA is inactive23. HHpred and biochemical analyses suggest that CCDC53, and the N termini of WASH and Fam21 are structurally and/or functionally similar to HSPC300 and the N termini of WAVE and Abi, respectively23. These many similarities, coupled with the similar overall shape of the WRC and SHRC23, suggest that the SHRC is analogously organized as a large SWIP:Strumpellin platform bound to a helical bundle of WASH:CCDC53:Fam21, with the WASH VCA sequestered by a similar mechanism.
We validated the structural organization observed in the crystal by replacing wild-type WAVE2 in HeLa cells with mutants targeting the trimer interface. Mutating the WAVE-HSPC300 interface (I50D/L54DWAVE2) or the WAVE-HSPC300/Abi interface (L40D/F51DWAVE2) appreciably decreased co-immunoprecipitation of WAVE2 with the four other components of the WRC (Supplementary Fig. 4), consistent with structural predictions.
Mechanism of WRC inhibition
The structure explains the inhibited nature of the WRC. In the complex, the WAVE1 VCA is bound by a conserved surface of Sra1 and residues 82–184 of WAVE1, which form five helices (α2–α6) and a series of intervening loops. This element of WAVE1 traces a meandering path across a concave surface of Sra1, and we refer to it as the ‘meander region’ (Fig. 2a and Supplementary Fig. 8). Contacts between the meander region and Sra1 bury over 2,100 Å2 (about 56% of the total WAVE1–Sra1 interface; Supplementary Fig. 9). The meander sequence is highly conserved among the different WAVE proteins (Supplementary Fig. 8), as is its contact surface on Sra1, suggesting that its interactions and irregular structure are conserved.
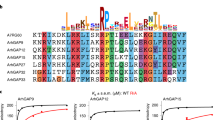
a, MiniWRC (rotated 90°about a horizontal axis from Fig. 1a). Sra1 and Nap1 are grey surfaces with conserved residues shown in green and cyan, respectively. Ribbons coloured as in Fig. 1. The meander region is indicated with a dashed line. b, V-helix–Sra1 interactions. Hydrogen bonds are dashed. Green dots indicate actin-binding residues. c, C-helix binding interface. Green dots indicate residues important for Arp2/3 activation. d, e, Arp2/3-mediated pyrene–actin assembly assays of miniWRC mutants. a.u., arbitrary units. d, L697D/Y704DSra1-miniWRC (light blue), L841A/F844A/W845ASra1-miniWRC (green), with Sra1 mutated at C- and V-helix binding site, respectively; miniWRC (brown); control (dark blue); VCA (magenta). e, W161E/K162DWAVE1-miniWRC (green), with WAVE1 mutated at the C-helix contact site; miniWRC (brown); control (dark blue).
The V- and C-regions of the VCA lie on the surface of Sra1 and form two amphipathic helices (residues 500–514 and 531–543, respectively) that also pack against α2 and α6 of WAVE1, respectively (Supplementary Fig. 10). The A-region of the VCA (residues 545–559) is probably disordered, given that it is not observed in the electron density. During actin filament nucleation, the V-region recruits an initial actin monomer to the nascent filament, while the C- and A-regions contribute binding energy and induce activating conformational changes in the Arp2/3 complex26,27. The structure and complementary experimental data below indicate that sequestration of both the V- and the C-regions by Sra1 and the meander region of WAVE1 underlies VCA inhibition within the WRC.
Inhibition of the V-region involves a combination of contacts to actin-binding residues and induction of structure that is incompatible with actin binding (Fig. 2b). In the complex of the V-region of WAVE2 with actin28, residues equivalent to 497–507 of WAVE1 form a helix that inserts into the cleft between actin subdomains 1 and 3. Residues equivalent to 508–516 are extended, and the Ile 509 and Arg 512 equivalents contact actin. In the structure of miniWRC, the entire V-region is helical, and the side chains of Leu 501, Leu 502, Ile 505, Ile 509 and Arg 512 are buried in the Sra1 interface, making the V-region inaccessible to actin. Sequestration of the V-region is an important contributor to WRC inhibition, because mutating V-helix contact residues Leu 841, Phe 844 and Trp 845 of Sra1 constitutively activates miniWRC towards the Arp2/3 complex (L841A/F844A/W845ASra1-miniWRC, Fig. 2d), producing branched filaments (Supplementary Fig. 11).
The C-helix is also critical for activation of the Arp2/3 complex29, because mutations of Val 531, Leu 535 or Arg 538 in the WAVE1 VCA reduce activity towards the Arp2/3 complex by at least half. In the miniWRC, the C-helix buries its hydrophobic face in the Sra1–α6 interface (Fig. 2c, Supplementary Figs 8 and 9). Intermolecularly, Val 531, Ala 532, Leu 535 and Ile 539 of WAVE1 make van der Waals contacts with Sra1. Intramolecularly, Val 531, Ile 534, Leu 535, Arg 538 and Val 541 of the C-helix pack against α6. Hence, the structure shows that the WRC sequesters C-helix residues that are important for activation of the Arp2/3 complex, resulting in inhibition. This mechanism is also supported by mutagenesis: perturbing contacts of the C-helix with either Sra1 (L697D/Y704DSra1-miniWRC, Fig. 2d) or the α6 helix (W161E/K162DWAVE1-miniWRC, Fig. 2e) leads to constitutive activation of the miniWRC in vitro. Additionally, replacement of wild-type WAVE2 with equivalent levels of the analogous α6 mutant (W160E/K161DWAVE2) in HeLa cells does not alter the integrity of the complex (Supplementary Fig. 4), but causes a dramatic redistribution of actin, with loss of stress fibres and assembly of filaments at the cell periphery, again consistent with constitutive activation of the WRC (Fig. 3a).
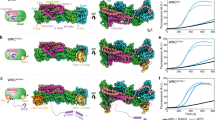
a, HeLa cells were transfected with shVector-yellow fluorescent protein (YFP) control, or vectors simultaneously suppressing WAVE2 and expressing shRNA-resistant YFP-tagged WAVE2 proteins (green) and scored blindly for lamellipodial phenotype. F-actin is visualized with phalloidin (red). Error bars in lower right panel show the standard deviation for at least three independent measurements. b, Fractional saturation of WRC versus free Rac1-GMPPNP measured by equilibrium dialysis. Error bars indicate the standard deviation in at least two independent measurements. KD is estimated from Rac concentration at 50% saturation; curves are binding isotherms to guide the eye. Pink uptriangle, ΔWRC (containing WAVE1(1–186)); black square, L697D/Y704DSra1-miniWRC; gold diamond, miniWRC; green circle, E434K/F626ASra1-ΔWRC; blue cross, R190DSra1-ΔWRC; cyan downtriangle, Δ154WAVE1-WRC (containing WAVE1(1–154)). c, WAVE1 meander region. Sra1 residues involved in binding Rac1 and WAVE1 Y151 are gold and blue sticks, respectively. The dashed oval indicates the proposed Rac1-binding surface. Phosphorylated WAVE1 residues Tyr 125, Thr 138 and Tyr 151 are red sticks. d, Arp2/3-mediated pyrene–actin assembly assays with miniWRC (green) or miniWRC containing Y151EWAVE1 (blue) or F686ESra1 (red). The control assay (orange) lacked WRC.
Interactions of the VCA, and perhaps the structure of the entire meander-VCA element, appear to be highly cooperative, as perturbation of either the V- or C-region contacts produces WRC activity near that of the isolated VCA (Figs 2d and e).
Activation by Rac
Rac plays an important role in controlling actin polymerization and lamellipodia formation through activation of the WRC in vivo8,30,31. Rac1 binds to recombinant WRC and activates it in vitro at micromolar concentrations under optimized assay conditions without disrupting the integrity of the complex13,14 (Supplementary Fig. 12). By analogy to the activation of WASP by Cdc42 (ref. 4), we reasoned that removing the VCA from the WRC would increase its affinity for Rac1. Indeed, in pull-down assays immobilized GST-Rac1:GMPPNP (a GTP analogue) bound a VCA-deleted WRC (ΔWRC) to a greater extent than miniWRC (Supplementary Fig. 13). Using equilibrium dialysis we measured a dissociation constant KD = 1–2 μM for ΔWRC. MiniWRC bound Rac with lower affinity, having KD = 7–10 µM (Fig. 3b). The constitutively active L697D/Y704DSra1-miniWRC (Fig. 3b) had affinity similar to that of ΔWRC, consistent with the idea that sequestration of the VCA motif competes with Rac1 binding. These results imply that Rac1 binds to the body of the WRC competitively with the VCA (either directly or indirectly), leading to activation of the complex.
To define the Rac1 interaction surface better, we searched for ΔWRC mutants with defects in Rac1 binding. Examination of the structure of WRC indentified several conserved surface patches. Mutations L697D/Y704D, L841A/F844A/W845A or E250K/Q399A of Sra1 did not perturb the binding of ΔWRC to Rac1 (data not shown). However, Sra1 mutations C179R, R190D, M632D and E434K/F626A severely impaired binding of ΔWRC to Rac1 without altering the integrity of the recombinant complex (Fig. 3b and Supplementary Fig. 13). These highly conserved Sra1 residues are located in a patch adjacent to α4–α6 of the WAVE1 meander region (Fig. 3c and Supplementary Fig. 14). Furthermore, truncation of α6 (Δ154WAVE1-WRC) also decreases Rac1 binding (Fig. 3c and Supplementary Fig. 13), suggesting that the helix or adjacent parts of the meander may contact the bound GTPase directly or stabilize its interaction sites on Sra1. These data implicate a Rac1 binding site on the WRC involving an Sra1 surface and perhaps part of the meander region of WAVE1. Interactions of Rac1 could then trigger conformational changes in the meander region and/or its contact site on Sra1. Because the meander appears to cooperatively stabilize the V- and C-regions of WAVE1 (see above), these perturbations could drive WRC activation by causing release of the VCA.
Activation by phosphorylation
In cells, phosphorylation of the meander region modulates WRC activity. Several groups report that phosphorylation of the strictly conserved Tyr 150 of WAVE2 (Tyr 151 in WAVE1 and WAVE3) by the Abl kinase is important for WRC-mediated actin assembly and lamellipodia formation15,16,17. Additionally, phosphorylation of WAVE Tyr 125 by Src or Thr 138 by Cdk5 alters cellular actin dynamics18,19.
Tyr 151 of WAVE1 is located in the α5–α6 loop of the meander region, and is buried in a hydrophobic pocket formed by Sra1 and WAVE1 (Fig. 3c and Supplementary Fig. 9). Phosphorylation of Tyr 151 would thus disrupt the contacts between the meander region and Sra1, leading to destabilization of the C-helix of the VCA motif and WRC activation. To test this idea, we reconstituted a miniWRC containing a phospho-mimicking Y151E mutation in vitro, and generated the analogous full-length WRC in HeLa cells using a knockdown/re-expression strategy. Consistent with the structural predictions, the mutant complexes displayed high actin assembly activity both in vivo and in vitro (Fig. 3a and d). Mutation of the Y151-binding pocket in Sra1 (F686ESra1-miniWRC) equivalently activates the WRC (Fig. 3d).
Tyr 125 of WAVE1 is also strictly conserved from animals to plants (Supplementary Fig. 8). This residue is located in α3 and its side chain packs against Gln 685 and makes a hydrogen bond to Asp 689 of Sra1 (Fig. 3c and Supplementary Fig. 9). Phosphorylation of Tyr 125 should disrupt the contact with Asp 689, and could destabilize the meander region of WAVE1, leading to release of the VCA. Consistent with this, replacement of WAVE2 with a Y124D mutant in cells increased lamellipodia formation (Fig. 3a). Thr 138 of WAVE1 makes intramolecular contacts with α4 and α5; its hydroxyl group is part of a network of hydrogen bonds that span these secondary elements (Fig. 3c and Supplementary Fig. 9). Phosphorylation of Thr 138 may thus also perturb the structure of the meander region, again contributing to activation of the WRC18,19.
Together, the data suggest that, analogous to Rac1 activation, phosphorylation could destabilize the meander region and/or its interactions with Sra1, leading to release of the VCA and activation of the WRC.
Discussion
The WRC is typically densely clustered at its sites of action in cells. This is believed to be necessary for spatially restricted actin assembly during, for example, polarized cell movement1. Clustering is mediated by the combined actions of phosphoinositide lipids and Rac, as well as various SH3-containing proteins1,3. The polybasic region of WAVE2 (equivalent to residues 172–184 of WAVE1) can bind phosphoinositide lipids in vitro, and is essential for membrane recruitment of the WRC and formation of lamellipodia in cells12. Surface electrostatic calculations show that the face containing the WAVE1:Abi2:HSPC300 four-helix bundle is negatively charged (Fig. 4a), whereas much of the face of the complex adjacent to the polybasic region is positively charged (Fig. 4b). This polar distribution suggests that when the WRC is recruited to the plasma membrane, the side covered by the four-helix bundle is exposed to the cytoplasm, and the opposite side contacts the membrane. In this orientation, Rac would bind approximately to the side of the WRC (Fig. 4c), and then its C-terminal isoprene group, the polybasic region of WAVE and the basic surface of the Sra1/Nap1 dimer could all be directed towards the plasma membrane. The meander region and the VCA motif of WAVE would face the cytoplasm, making them accessible to other regulators (for example, kinases), and to the Arp2/3 complex and actin. This organization would allow simultaneous phosphoinositide and Rac binding, cooperatively recruiting the WRC to membranes and enhancing allosteric activation. Self-association of the WRC at membranes14, and consequent enhanced activity5, could be mediated by intercomplex binding of the N-terminal helix of Sra1 with the WAVE/Abi/HSPC300 trimer, as observed in the crystal lattice (Supplementary Fig. 3).
a, b, Electrostatic surface of miniWRC (−5 kT per electron (red) to +5 kT per electron (blue)), oriented as in Fig. 1a and b, respectively. c, Schematic illustrating proposed WRC orientation at the plasma membrane and cooperative recruitment and activation by Rac and phospholipids. Plus and minus signs indicate regions of positive and negative surface charge. Phosphorylation sites in WAVE1 meander are indicated in red. Disordered elements of WAVE1 are indicated with dotted purple lines. It remains unclear what portions of the meander are disrupted by different stimuli.
Sra1 was recently reported to support translation inhibition through simultaneous binding to the translational regulator FMRP and the translation initiation factor eIF4E9,10. However, the putative mode of eIF4E binding is incompatible with the WRC structure (Fig. 1b, Supplementary Fig. 15). Thus, eIF4E may bind to isolated Sra1, but not the WRC, consistent with the finding that eIF4E co-immunoprecipitates with Sra1 but not WAVE9. These observations suggest that Sra1 may partition between the WRC, which regulates actin dynamics, and a free (or alternatively complexed) state that regulates translation. Similar arguments have also been made regarding different pools of Nap1 (ref. 32) and Abi (ref. 33). Interestingly, defects in Sra1 or its ligands in both pathways—protocadherin-10, which binds the WRC34, and FMRP—are implicated in autism and other mental disorders35,36, suggesting that an appropriate balance of these pathways or their joint action may be needed for proper neuronal development and function. Future studies of the intact WRC and its separate components will reveal how this system coordinates multiple processes in normal and abnormal cellular function.
Methods Summary
Sra1, Nap1, WAVE1, Abi2 and HSPC300 were overexpressed separately, partially purified, assembled into an Sra1:Nap1 dimer and a WAVE1/Abi2/HSPC300 trimer, respectively, and then assembled into the miniWRC pentamer. Further purification produced homogeneous samples. Crystals of miniWRC were obtained by hanging-drop vapour diffusion at 4 °C. All the data sets were collected at the ID-19 beamline (Advanced Photon Source) and processed with the HKL3000 suite37 and CCP4 suites38. Experimental phases were determined from selenium-MIRAS data collected on samples containing either SeMet-Sra1 or SeMet-Nap1, and analysed using ShelxD39. Phases were improved using MLPHARE40 and Parrot41. The atomic model of the complex was built using Buccaneer42 and Coot43, and refined using Refmac 5 (ref. 44). Equilibrium dialysis was done at room temperature and protein concentrations were determined using Deep Purple gel staining (GE Healthcare). Actin polymerization and GST-Rac1 pull-down assays were performed as described previously13. HeLa cells were grown directly on coverslips, fixed in 4% paraformaldehyde, and prepared for immunofluorescence as described25.
Online Methods
Protein expression and purification
The proteins were handled with a modified version of the earlier protocol13. Mutants were generated using QuikChange (Stratagene). To achieve maximal expression, His-tagged Sra1 (1–1253) and Nap1 (1–1128) were separately overexpressed in sf9 cells and Hi5 cells, respectively. Cells were lysed together and the Sra1:Nap1 complex was partially purified by a Ni-affinity column. For miniWRC, WAVE1–186VCA (WAVE1(1–186)-(GlyGlySer)6-VCA(485–559)), Abi2 (1–158) and full-length HSPC300 (1–75) were expressed separately in Escherichia coli as MBP-fusion proteins, purified with amylose-affinity chromatography and assembled into the trimer subcomplex by incubation in the presence of 1% NP-40 (Sigma) for 48 h. The trimer was isolated using Source 15Q chromatography, and then mixed with the Sra1:Nap1 dimer at a 1:1.5 molar ratio overnight on ice to form the pentameric complex. Excess Sra1:Nap1 dimer was removed by an amylose-affinity column. The eluted complex was further purified using Mono Q chromatography. The fusion tags were removed by treatment with TEV protease at room temperature for 4–6 h. The sample was brought to homogeneity by a Superdex200 column equilibrated with protein buffer (200 mM NaCl, 12% glycerol (w/v), 10 mM Tris-HCl, 5 mM DTT, pH 8.0), concentrated to about 10 mg ml−1 and stored at −80 °C. Selenomethionine-labelled Sra1 and Nap1 were obtained by culturing insect cells in methionine-free medium according to standard protocols (Expression Systems). ΔWRC and Δ154WAVE1-WRC were produced in the same way as miniWRC, but contained WAVE1(1–186) and WAVE1(1–154), respectively.
Crystallization
Crystals were grown at 4 °C by hanging-drop vapour diffusion methods. The native miniWRC crystals grew from 10% (w/v) glycerol, 4% PEG 10,000, 12–20% PEG 300, 100 mM Tris-HCl, 2 mM TCEP, 2 mM EDTA, pH 8.5 with protein to reservoir volume ratio 1:1.8. Crystals form in space group P212121, with a = 103.5 Å, b = 113.8 Å, c = 323.0 Å and diffracted only to 4.2 Å. Extensive optimization failed to improve crystal quality. The crystals with selenomethionine-labelled Sra1 and Nap1 grew from similar conditions in the same space group, with a = 97.0 Å, b = 114.0 Å, c = 327.2 Å, and diffracted to 2.3 Å.
Data processing and structure solution
The structure of miniWRC was solved by experimental phasing using data processed with the HKL3000 suite37,40 (Supplementary Table 1). The initial phases were obtained by combining data from three crystals: the first two contained Se-labelled Sra1 and the third contained Se-labelled Nap1. There was a significant level of non-isomorphism between crystals. In the case of the structure with Se-labelled Sra1, there was an additional component of non-isomorphism due to the breaking of crystal symmetry (P21 versus P212121). The symmetry violation was larger than experimental uncertainty, but still small enough to use the higher-symmetry space group (P212121) in structure solution. So the two data sets for the Sra1-labelled crystals scaled together were used for structure refinement in the P212121 space group. Radiation-induced non-isomorphism present in the collected data sets was corrected with the use of novel procedures45. The data were anisotropic, with the best direction diffracting to 2.29 Å. To use the data fully, structure factors were anisotropically sharpened to make the resolution fall-off isotropic. In consequence, the shell-based Rmerge values did not have statistical meaning in this case. To define the resolution properly we used an I/σ(I) criterion, where I represents averaged intensity after correcting for anisotropicity, and σ(I) is defined for each reflection separately. The very small change of Rfree at higher resolutions is an indicator that the procedure worked correctly.
After all these corrections, the heavy atom substructure for the averaged Sra1-P212121 was determined by ShelxD39 called within the HKL3000 suite37,40. Using initial phases, the positions of heavy atoms for the other data set (Se-labelled Nap1) were determined by searching for peaks in the anomalous difference map. The complete set of heavy atom positions was refined using MLPHARE40 with anomalous differences only. Density modification was performed with Parrot41, and initial model building with Buccaneer42 (Supplementary Table 1). The rest of the model was built manually, using Coot43 and Refmac44, where experimental phases were used as an additional restraint.
Refinement was performed with Refmac, using the TLSMD server to generate TLS bodies for refinement46, and the Molprobity server47 to check the validity of the structure. The final refinement is summarized in Supplementary Table 1, which shows that 98.26%, 99.96% and 0.04% of the amino acid residues are in the favoured, additional allowed and disallowed region of the Ramachandran plots, respectively.
Equilibrium dialysis
Equilibrium dialysis was performed at room temperature using a fast micro-equilibrium dialyser (Harvard Apparatus). Rac1 (Q61L) was charged with GMP-PNP in pH 7.5 buffer containing 20 mM Tris-HCl, 150 mM NaCl, 14% (w/v) glycerol and 2 mM MgCl2. WRC constructs were used at 4 μM. After reaching equilibrium (about 40 h), the concentrations of free Rac1 in one chamber and total Rac1 in the other chamber of the dialyser were analysed by SDS–PAGE gels, stained with Deep Purple (GE Healthcare) and quantified with ImageGauge (Fujifilm) by comparison to a standard curve generated from gels containing known concentrations of Rac1. Nonlinear curve fitting to extract KD was performed using Prism 5. At Rac1 concentrations above about 10 µM, technical limitations—such as bubble formation, protein instability over an extended dialysis time (required by high solution viscosity) and loss—prevented us from obtaining reproducible data in this regime. Thus, KD was estimated from the approximate 50% saturation point in the assay.
Actin polymerization and GST pulldown assays
Arp2/3-complex-mediated actin polymerization assays were performed as described13 using 4 μM actin (5% pyrene labelled) and 10 nM Arp2/3 complex in KMEI-20G buffer (20% (w/v) glycerol, 50 mM KCl, 1 mM MgCl2, 1 mM EGTA and 10 mM imidazole pH 7.0). The data in Supplementary Fig. 12 used 1 μM actin (5% pyrene labelled) and 30 nM Arp2/3 complex in buffer lacking glycerol. 5 nM WAVE1-Abi2-HSPC300 trimer was used as the aggregated VCA. Different concentrations of WAVE1 VCA (residues 485–559) were also used.
GST pulldown experiments were performed using 40 μM GST-Rac1 (20 μM for ΔWRC and Δ154-WRC comparison), 1.5 μM WRC constructs and 60 μl glutathione sepharose 4B resin in 0.2 ml pulldown buffer (10 mM Na-Hepes pH 7.0, 100 mM NaCl, 2 mM MgCl2, 10% glycerol (w/v) and 2 mM DTT). After gentle mixing at room temperature for 30 min, the resin was spun down, washed three times with 0.4 ml pulldown buffer, and eluted with 30 mM reduced glutathione. The eluted proteins were resolved by SDS–PAGE and visualized with Coomassie blue.
Cell biology studies
HeLa cells were transfected with shVector-YFP control and various short hairpin (sh)WAVE2/HA-YFP-WAVE2 reconstitution vectors based on established protocols25,48. We used shWAVE2 (GAGAAGAGAAAGCACAGGA), and made shRNA-resistant WAVE2 complementary DNA (GAaAAaAGgAAaCACAGGA) to generate HA-YFP suppression/reconstitution vectors as described25. Transfectants were analysed 72 h post transfection by immunoprecipitation or immunofluorescence. Anti-HA affinity matrix and anti-HA-HRP were from Roche. Rabbit anti-Nap1 was generated using a synthetic peptide corresponding to amino acids 1117–1128 of human Nap1. Anti-Sra1/PIR121 was previously described47. Anti-WAVE1 was obtained from Upstate Biotechnology. Alexa Fluor-647 phalloidin was used (Invitrogen) to stain the actin filament. Images were obtained with an LSM-710 laser scanning confocal microscope (Carl Zeiss) and analysed for the presence of lammelipodia formation. For quantification, more than 200 cells for each transfected cell population, in at least three independent experiments, were blindly scored.
Electron microscopy
For negative staining electron microscopy, 4 µl of protein solution (10 µg ml−1) was applied to glow-discharged carbon-coated 300-mesh Cu/Rh grids (Emsdiasum) and incubated for 30–60 s. Excess solution was blotted off with filter paper (Whatman #1), the grid was washed with 4 µl of protein buffer and stained with 2% uranyl acetate. Grids were imaged under low-dose conditions (10–25 electrons per Å2) on an FEI Tecnai G2 Spirit BioTwin electron microscope (FEI) with a LaB6 filament operated at 120 kV at a nominal magnification of 30,000×. Images were recorded with a Gatan 2,048 × 2,048-pixel charge-coupled device (CCD) camera (Gatan) using 0.8–2.5 μm underfocus, with a final resolution of 3.63 Å per pixel on the object. Particles were picked manually using the boxer application in EMAN49, normalized, and filtered to 22 Å. Ten class averages were generated using nine iterations of reference-free classification (refine2d.py), using a common reference to orient the classes to show the ‘upright’ view. Classes with fewer than eight particles were discarded automatically after each iteration.
Calculation of surface conservation
The conservation scores were calculated using the Consurf Sever50. Increasing conservation (scored from 1 to 9) was colour-coded in the figures by the spectra of white-to-green and white-to-cyan for Sra1 and Nap1, respectively. Residues that were scored 7–9 (green in Sra1 and cyan in Nap1) are considered to be conserved, which typically have a single amino acid in about 80% of the sequences we examined. The sequences of Sra1 orthologues used in the calculation are: NP_055423.1(Hs_Sra1), AAH72814.1, AAU05773.1, NP_974801.2, XP_001790637.1, EEN67132.1, NP_499949.2, Q6UK63.1, NP_650447.1, NP_997924.1, CAQ17050.1, NP_035500.2, XP_001379666.1, XP_001745727.1, EDO41734.1, NP_001048941.1, XP_001753041.1, XP_002468523.1, XP_002198076.1, EDV20545.1 and XP_002268225.1. The sequences of Nap1 orthologues used are: NP_038464.1(Hs_Nap1), NP_181056.2, ABN04850.1, NP_001062406.1, EDQ74364.1, XP_001369085.1, XP_001232275.1, NP_058661.1, NP_001087969.1, XP_690388.2, CAA56333.1, NP_001137348.1, XP_002221623.1, NP_524214.1, XP_971119.1, EEC12517.1, XP_001184421.1, EDO45022.1, NP_502368.1, CAZ27842.1, XP_644083.1 and EAY21283.1.
Total internal reflection fluorescence microscopy
Actin (1 µM) was polymerized in the presence of Arp2/3 complex (10 nM), N-WASP VCA (0.1 µM) or active L841A/F844A/W845ASra1-miniWRC (0.1 µM) in KMEI buffer for 20 min, before adding Alexa-488-phalloidin (1:50 dilution). As a negative control, actin (1 µM) alone was polymerized for 1 h. Alexa 488-phalloidin-bound filaments were diluted 1,000-fold before adsorbing onto poly-D-lysine-coated glass-bottom dishes (Ted Pella) for 10 min. Filaments were imaged using a laser-based total internal reflection fluorescence microscope (Olympus IX-71 base microscope), Micro-Manager 1.3 (Vale lab), and a Photometrics Cascade II 512 EMCCD camera.
References
Takenawa, T. & Suetsugu, S. The WASP-WAVE protein network: connecting the membrane to the cytoskeleton. Nature Rev. Mol. Cell Biol. 8, 37–48 (2007)
Pollitt, A. Y. & Insall, R. H. WASP and SCAR/WAVE proteins: the drivers of actin assembly. J. Cell Sci. 122, 2575–2578 (2009)
Padrick, S. B. & Rosen, M. K. Physical mechanisms of signal integration by WASP family proteins. Annu. Rev. Biochem. 79, 707–735 (2010)
Kim, A. S., Kakalis, L. T., Abdul-Manan, N., Liu, G. A. & Rosen, M. K. Autoinhibition and activation mechanisms of the Wiskott-Aldrich syndrome protein. Nature 404, 151–158 (2000)
Padrick, S. B. et al. Hierarchical regulation of WASP/WAVE proteins. Mol. Cell 32, 426–438 (2008)
Eden, S., Rohatgi, R., Podtelejnikov, A. V., Mann, M. & Kirschner, M. W. Mechanism of regulation of WAVE1-induced actin nucleation by Rac1 and Nck. Nature 418, 790–793 (2002)
Stovold, C. F., Millard, T. H. & Machesky, L. M. Inclusion of Scar/WAVE3 in a similar complex to Scar/WAVE1 and 2. BMC Cell Biol. 6, 11 (2005)
Vartiainen, M. K. & Machesky, L. M. The WASP-Arp2/3 pathway: genetic insights. Curr. Opin. Cell Biol. 16, 174–181 (2004)
Napoli, I. et al. The fragile X syndrome protein represses activity-dependent translation through CYFIP1, a new 4E-BP. Cell 134, 1042–1054 (2008)
Schenck, A. et al. CYFIP/Sra-1 controls neuronal connectivity in Drosophila and links the Rac1 GTPase pathway to the fragile X protein. Neuron 38, 887–898 (2003)
Kobayashi, K. et al. p140Sra-1 (specifically Rac1-associated protein) is a novel specific target for Rac1 small GTPase. J. Biol. Chem. 273, 291–295 (1998)
Oikawa, T. et al. PtdIns(3,4,5)P3 binding is necessary for WAVE2-induced formation of lamellipodia. Nature Cell Biol. 6, 420–426 (2004)
Ismail, A. M., Padrick, S. B., Chen, B., Umetani, J. & Rosen, M. K. The WAVE regulatory complex is inhibited. Nature Struct. Mol. Biol. 16, 561–563 (2009)
Lebensohn, A. M. & Kirschner, M. W. Activation of the WAVE complex by coincident signals controls actin assembly. Mol. Cell 36, 512–524 (2009)
Sossey-Alaoui, K., Li, X. & Cowell, J. K. c-Abl-mediated phosphorylation of WAVE3 is required for lamellipodia formation and cell migration. J. Biol. Chem. 282, 26257–26265 (2007)
Stuart, J. R., Gonzalez, F. H., Kawai, H. & Yuan, Z. M. c-Abl interacts with the WAVE2 signaling complex to induce membrane ruffling and cell spreading. J. Biol. Chem. 281, 31290–31297 (2006)
Leng, Y. et al. Abelson-interactor-1 promotes WAVE2 membrane translocation and Abelson-mediated tyrosine phosphorylation required for WAVE2 activation. Proc. Natl Acad. Sci. USA 102, 1098–1103 (2005)
Miyamoto, Y., Yamauchi, J. & Tanoue, A. Cdk5 phosphorylation of WAVE2 regulates oligodendrocyte precursor cell migration through nonreceptor tyrosine kinase Fyn. J. Neurosci. 28, 8326–8337 (2008)
Ardern, H. et al. Src-dependent phosphorylation of Scar1 promotes its association with the Arp2/3 complex. Cell Motil. Cytoskeleton 63, 6–13 (2006)
Dai, Z. & Pendergast, A. M. Abi-2, a novel SH3-containing protein interacts with the c-Abl tyrosine kinase and modulates c-Abl transforming activity. Genes Dev. 9, 2569–2582 (1995)
Holm, L., Kaariainen, S., Rosenstrom, P. & Schenkel, A. Searching protein structure databases with DaliLite v.3. Bioinformatics 24, 2780–2781 (2008)
Soding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, 244–248 (2005)
Jia, D. et al. WASH and WAVE actin regulators of the Wiskott-Aldrich syndrome protein (WASP) family are controlled by analogous structurally related complexes. Proc. Natl Acad. Sci. USA 107, 10442–10447 (2010)
Derivery, E. et al. The Arp2/3 activator WASH controls the fission of endosomes through a large multiprotein complex. Dev. Cell 17, 712–723 (2009)
Gomez, T. S. & Billadeau, D. D. A. FAM21-containing WASH complex regulates retromer-dependent sorting. Dev. Cell 17, 699–711 (2009)
Goley, E. D., Rodenbusch, S. E., Martin, A. C. & Welch, M. D. Critical conformational changes in the Arp2/3 complex are induced by nucleotide and nucleation promoting factor. Mol. Cell 16, 269–279 (2004)
Marchand, J. B., Kaiser, D. A., Pollard, T. D. & Higgs, H. N. Interaction of WASP/Scar proteins with actin and vertebrate Arp2/3 complex. Nature Cell Biol. 3, 76–82 (2001)
Chereau, D. et al. Actin-bound structures of Wiskott-Aldrich syndrome protein (WASP)-homology domain 2 and the implications for filament assembly. Proc. Natl Acad. Sci. USA 102, 16644–16649 (2005)
Panchal, S. C., Kaiser, D. A., Torres, E., Pollard, T. D. & Rosen, M. K. A conserved amphipathic helix in WASP/Scar proteins is essential for activation of Arp2/3 complex. Nature Struct. Biol. 10, 591–598 (2003)
Stradal, T. E. et al. Regulation of actin dynamics by WASP and WAVE family proteins. Trends Cell Biol. 14, 303–311 (2004)
Steffen, A. et al. Sra-1 and Nap1 link Rac to actin assembly driving lamellipodia formation. EMBO J. 23, 749–759 (2004)
Weiner, O. D. et al. Hem-1 complexes are essential for Rac activation, actin polymerization, and myosin regulation during neutrophil chemotaxis. PLoS Biol. 4, e38 (2006)
Ryu, J. R., Echarri, A., Li, R. & Pendergast, A. M. Regulation of cell-cell adhesion by Abi/Diaphanous complexes. Mol. Cell. Biol. 29, 1735–1748 (2009)
Nakao, S., Platek, A., Hirano, S. & Takeichi, M. Contact-dependent promotion of cell migration by the OL-protocadherin-Nap1 interaction. J. Cell Biol. 182, 395–410 (2008)
Nowicki, S. T. et al. The Prader-Willi phenotype of fragile X syndrome. J. Dev. Behav. Pediatr. 28, 133–138 (2007)
Stefansson, H. et al. Large recurrent microdeletions associated with schizophrenia. Nature 455, 232–236 (2008)
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr. D 62, 859–866 (2006)
Dodson, E. J., Winn, M. & Ralph, A. Collaborative Computational Project number 4: providing programs for protein crystallography. Methods Enzymol. 277, 620–633 (1997)
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008)
Otwinowski, Z. M. W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Zhang, K. Y., Cowtan, K. & Main, P. Combining constraints for electron-density modification. Methods Enzymol. 277, 53–64 (1997)
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D Biol. Crystallogr. 62, 1002–1011 (2006)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Borek, D., Ginell, S. L., Cymborowski, M., Minor, W. & Otwinowski, Z. The many faces of radiation-induced changes. J. Synchrotron Radiat. 14, 24–33 (2007)
Painter, J. & Merritt, E. A. Optimal description of a protein structure in terms of multiple groups undergoing TLS motion. Acta Crystallogr. D 62, 439–450 (2006)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, 375–383 (2007)
Nolz, J. C. et al. The WAVE2 complex regulates actin cytoskeletal reorganization and CRAC-mediated calcium entry during T cell activation. Curr. Biol. 16, 24–34 (2006)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, 299–302 (2005)
Acknowledgements
We thank B. Chen for providing samples of full-length WRC and VCA polypeptide for electron microscopy and some biochemical analyses, C. Pak for helping with the total internal reflection fluorescence experiment, D. Tomchick and C. Brautigam for technical assistance and N. Grishin for assistance with sequence analysis and discussion. Research was supported by fellowships from the Cancer Research Institute and the NIH (1F32-GM06917902) to Z.C. and S.B.P., respectively, an Allergic Diseases Training grant (AI07047) to T.S.G., grants from the NIH to D.D.B. (R01-AI065474), Z.O. (R01-GM053163) and M.K.R. (R01-GM056322), a grant from the Welch Foundation to M.K.R. (I-1544) and the Howard Hughes Medical Institute. D.D.B. is a Leukemia and Lymphoma Society Scholar. Use of the Argonne National Laboratory Structural Biology Center beamlines at the Advanced Photon Source was supported by the US DOE under contract DE-AC02-06CH11357.
Author information
Authors and Affiliations
Contributions
M.K.R. oversaw the project. Z.C., A.I., S.B.P. and J.U. developed the WRC reconstitution. D.B., Z.C. and Z.O. determined the structure of the WRC. Z.C. performed the biochemical experiments. D.D.B. and T.S.G. performed the cellular experiments. Z.M. performed the electron microscopy experiments. .Z.C., Z.O. and M.K.R. analysed the WRC structure. D.B., Z.C., Z.O., S.B.P. and M.K.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, Supplementary Figures 1-15 with legends and additional references. (PDF 17463 kb)
Rights and permissions
About this article
Cite this article
Chen, Z., Borek, D., Padrick, S. et al. Structure and control of the actin regulatory WAVE complex. Nature 468, 533–538 (2010). https://doi.org/10.1038/nature09623
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09623
This article is cited by
-
Orchestration of synaptic functions by WAVE regulatory complex-mediated actin reorganization
Experimental & Molecular Medicine (2023)
-
Time-dependent inhibition of Rac1 in the VTA enhances long-term aversive memory: implications in active forgetting mechanisms
Scientific Reports (2023)
-
Small extracellular vesicles promote invadopodia activity in glioblastoma cells in a therapy-dependent manner
Cellular Oncology (2023)
-
Regulation of the Scar/WAVE complex in migrating cells: A summary of our understanding
Journal of Biosciences (2023)
-
Role of NCKAP1 in the Defective Phagocytic Function of Microglia-Like Cells Derived from Rapidly Progressing Sporadic ALS
Molecular Neurobiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.