Abstract
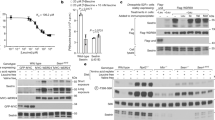
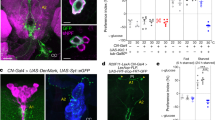
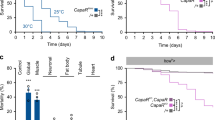
Many stem, progenitor and cancer cells undergo periods of mitotic quiescence from which they can be reactivated1,2,3,4,5. The signals triggering entry into and exit from this reversible dormant state are not well understood. In the developing Drosophila central nervous system, multipotent self-renewing progenitors called neuroblasts6,7,8,9 undergo quiescence in a stereotypical spatiotemporal pattern10. Entry into quiescence is regulated by Hox proteins and an internal neuroblast timer11,12,13. Exit from quiescence (reactivation) is subject to a nutritional checkpoint requiring dietary amino acids14. Organ co-cultures also implicate an unidentified signal from an adipose/hepatic-like tissue called the fat body14. Here we provide in vivo evidence that Slimfast amino-acid sensing and Target of rapamycin (TOR) signalling15 activate a fat-body-derived signal (FDS) required for neuroblast reactivation. Downstream of this signal, Insulin-like receptor signalling and the Phosphatidylinositol 3-kinase (PI3K)/TOR network are required in neuroblasts for exit from quiescence. We demonstrate that nutritionally regulated glial cells provide the source of Insulin-like peptides (ILPs) relevant for timely neuroblast reactivation but not for overall larval growth. Conversely, ILPs secreted into the haemolymph by median neurosecretory cells systemically control organismal size16,17,18 but do not reactivate neuroblasts. Drosophila thus contains two segregated ILP pools, one regulating proliferation within the central nervous system and the other controlling tissue growth systemically. Our findings support a model in which amino acids trigger the cell cycle re-entry of neural progenitors via a fat-body–glia–neuroblasts relay. This mechanism indicates that dietary nutrients and remote organs, as well as local niches, are key regulators of transitions in stem-cell behaviour.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dhawan, J. & Rando, T. A. Stem cells in postnatal myogenesis: molecular mechanisms of satellite cell quiescence, activation and replenishment. Trends Cell Biol. 15, 666–673 (2005)
Coller, H. A. What’s taking so long? S-phase entry from quiescence versus proliferation. Nature Rev. Mol. Cell Biol. 8, 667–670 (2007)
Yanagida, M. Cellular quiescence: are controlling genes conserved? Trends Cell Biol. 19, 705–715 (2009)
Chen, E. & Finkel, T. The tortoise, the hare, and the FoxO. Cell Stem Cell 5, 451–452 (2009)
Sánchez-Garcia, I., Vicente-Duenas, C. & Cobaleda, C. The theoretical basis of cancer-stem-cell-based therapeutics of cancer: can it be put into practice? Bioessays 29, 1269–1280 (2007)
Betschinger, J. & Knoblich, J. A. Dare to be different: asymmetric cell division in Drosophila, C. elegans and vertebrates. Curr. Biol. 14, R674–R685 (2004)
Egger, B., Chell, J. M. & Brand, A. H. Insights into neural stem cell biology from flies. Phil. Trans. R. Soc. Lond. B 363, 39–56 (2008)
Doe, C. Q. Neural stem cells: balancing self-renewal with differentiation. Development 135, 1575–1587 (2008)
Sousa-Nunes, R., Cheng, L. Y. & Gould, A. P. Regulating neural proliferation in the Drosophila CNS. Curr. Opin. Neurobiol. 20, 50–57 (2010)
Truman, J. W. & Bate, M. Spatial and temporal patterns of neurogenesis in the central nervous system of Drosophila melanogaster . Dev. Biol. 125, 145–157 (1988)
Tsuji, T., Hasegawa, E. & Isshiki, T. Neuroblast entry into quiescence is regulated intrinsically by the combined action of spatial Hox proteins and temporal identity factors. Development 135, 3859–3869 (2008)
Kambadur, R. et al. Regulation of POU genes by castor and hunchback establishes layered compartments in the Drosophila CNS. Genes Dev. 12, 246–260 (1998)
Isshiki, T., Pearson, B., Holbrook, S. & Doe, C. Q. Drosophila neuroblasts sequentially express transcription factors which specify the temporal identity of their neuronal progeny. Cell 106, 511–521 (2001)
Britton, J. S. & Edgar, B. A. Environmental control of the cell cycle in Drosophila: nutrition activates mitotic and endoreplicative cells by distinct mechanisms. Development 125, 2149–2158 (1998)
Colombani, J. et al. A nutrient sensor mechanism controls Drosophila growth. Cell 114, 739–749 (2003)
Brogiolo, W. et al. An evolutionarily conserved function of the Drosophila insulin receptor and insulin-like peptides in growth control. Curr. Biol. 11, 213–221 (2001)
Ikeya, T., Galic, M., Belawat, P., Nairz, K. & Hafen, E. Nutrient-dependent expression of insulin-like peptides from neuroendocrine cells in the CNS contributes to growth regulation in Drosophila . Curr. Biol. 12, 1293–1300 (2002)
Rulifson, E. J., Kim, S. K. & Nusse, R. Ablation of insulin-producing neurons in flies: growth and diabetic phenotypes. Science 296, 1118–1120 (2002)
Slaidina, M., Delanoue, R., Gronke, S., Partridge, L. & Leopold, P. A Drosophila insulin-like peptide promotes growth during nonfeeding states. Dev. Cell 17, 874–884 (2009)
Okamoto, N. et al. A fat body-derived IGF-like peptide regulates postfeeding growth in Drosophila . Dev. Cell 17, 885–891 (2009)
Géminard, C., Rulifson, E. J. & Leopold, P. Remote control of insulin secretion by fat cells in Drosophila . Cell Metab. 10, 199–207 (2009)
Polak, P. & Hall, M. N. mTOR and the control of whole body metabolism. Curr. Opin. Cell Biol. 21, 209–218 (2009)
Neufeld, T. P. Body building: regulation of shape and size by PI3K/TOR signaling during development. Mech. Dev. 120, 1283–1296 (2003)
Teleman, A. A. Molecular mechanisms of metabolic regulation by insulin in Drosophila . Biochem. J. 425, 13–26 (2010)
Grönke, S., Clarke, D. F., Broughton, S., Andrews, T. D. & Partridge, L. Molecular evolution and functional characterization of Drosophila insulin-like peptides. PLoS Genet. 6, e1000857 (2010)
Zhang, H. et al. Deletion of Drosophila insulin-like peptides causes growth defects and metabolic abnormalities. Proc. Natl Acad. Sci. USA 106, 19617–19622 (2009)
Chell, J. M. & Brand, A. H. Nutrition-responsive glia control exit of neural stem cells from quiescence. Cell 143, 1161–1173 (2010)
Mangan, S. & Alon, U. Structure and function of the feed-forward loop network motif. Proc. Natl Acad. Sci. USA 100, 11980–11985 (2003)
Fernandez-Almonacid, R. & Rosen, O. M. Structure and ligand specificity of the Drosophila melanogaster insulin receptor. Mol. Cell. Biol. 7, 2718–2727 (1987)
Hodge, R. D., D’Ercole, A. J. & O’Kusky, J. R. Insulin-like growth factor-I accelerates the cell cycle by decreasing G1 phase length and increases cell cycle reentry in the embryonic cerebral cortex. J. Neurosci. 24, 10201–10210 (2004)
Oldham, S. et al. Genetic and biochemical characterization of dTOR, the Drosophila homolog of the target of rapamycin. Genes Dev. 14, 2689–2694 (2000)
Brogiolo, W. et al. An evolutionarily conserved function of the Drosophila insulin receptor and insulin-like peptides in growth control. Curr. Biol. 11, 213–221 (2001)
Zhang, H. et al. Deletion of Drosophila insulin-like peptides causes growth defects and metabolic abnormalities. Proc. Natl Acad. Sci. USA 106, 19617–19622 (2009)
Gronke, S. et al. Molecular evolution and functional characterization of Drosophila insulin-like peptides. PLoS Genet. 6, e1000857 (2010)
Colombani, J. et al. A nutrient sensor mechanism controls Drosophila growth. Cell 114, 739–749 (2003)
Tapon, N. et al. The Drosophila tuberous sclerosis complex gene homologs restrict cell growth and cell proliferation. Cell 105, 345–355 (2001)
Hennig, K. M., Colombani, J. & Neufeld, T. P. TOR coordinates bulk and targeted endocytosis in the Drosophila melanogaster fat body to regulate cell growth. J. Cell Biol. 173, 963–974 (2006)
Xiong, W. C. et al. repo encodes a glial-specific homeo domain protein required in the Drosophila nervous system. Genes Dev. 8, 981–994 (1994)
Connolly, J. B. et al. Associative learning disrupted by impaired Gs signaling in Drosophila mushroom bodies. Science 274, 2104–2107 (1996)
Ito, K., Urban, J. & Technau, G. M. Distribution, classification, and development of Drosophila glial cells in the late embryonic and early larval ventral nerve cord. Rouxs Arch. Dev. Biol. 204, 284–307 (1995)
Hacker, U. et al. piggyBac-based insertional mutagenesis in the presence of stably integrated P elements in Drosophila . Proc. Natl Acad. Sci. USA 100, 7720–7725 (2003)
Shiga, Y., Tanaka-Matakatsu, M. & Hayashi, S. A nuclear GFP/ β-galactosidase fusion protein as a marker for morphogenesis in living Drosophila . Dev. Growth Differ. 38, 99–106 (1996)
Leevers, S. J. et al. The Drosophila phosphoinositide 3-kinase Dp110 promotes cell growth. EMBO J. 15, 6584–6594 (1996)
Huang, H. et al. PTEN affects cell size, cell proliferation and apoptosis during Drosophila eye development. Development 126, 5365–5372 (1999)
Weinkove, D. et al. Regulation of imaginal disc cell size, cell number and organ size by Drosophila class IA phosphoinositide 3-kinase and its adaptor. Curr. Biol. 9, 1019–1029 (1999)
Rintelen, F., Stocker, H., Thomas, G. & Hafen, E. PDK1 regulates growth through Akt and S6K in Drosophila . Proc. Natl Acad. Sci. USA 98, 15020–15025 (2001)
Puig, O., Marr, M. T., Ruhf, M. L. & Tjian, R. Control of cell number by Drosophila FOXO: downstream and feedback regulation of the insulin receptor pathway. Genes Dev. 17, 2006–2020 (2003)
Miguel-Aliaga, I., Thor, S. & Gould, A. P. Postmitotic specification of Drosophila insulinergic neurons from pioneer neurons. PLoS Biol. 6, e58 (2008)
Ikeya, T. et al. Nutrient-dependent expression of insulin-like peptides from neuroendocrine cells in the CNS contributes to growth regulation in Drosophila . Curr. Biol. 12, 1293–1300 (2002)
Moline, M. M., Southern, C. & Bejsovec, A. Directionality of wingless protein transport influences epidermal patterning in the Drosophila embryo. Development 126, 4375–4384 (1999)
Maurange, C., Cheng, L. & Gould, A. P. Temporal transcription factors and their targets schedule the end of neural proliferation in Drosophila . Cell 133, 891–902 (2008)
Awasaki, T., Lai, S.-L., Ito, K. & Lee, T. Organization and postembryonic development of glial cells in the adult central brain of Drosophila . J. Neurosci. 28, 13742–13753 (2008)
Schwabe, T. et al. GPCR Signaling is required for blood-brain barrier formation in Drosophila . Cell 123, 133–144 (2005)
Scholz, H., Sadlowski, E., Klaes, A. & Klambt, C. Control of midline glia development in the embryonic Drosophila CNS. Mech. Dev. 64, 139–151 (1997)
Kidd, T., Bland, K. S. & Goodman, C. S. Slit is the midline repellent for the Robo receptor in Drosophila . Cell 96, 785–794 (1999)
Rulifson, E. J., Kim, S. K. & Nusse, R. Ablation of insulin-producing neurons in flies: growth and diabetic phenotypes. Science 296, 1118–1120 (2002)
Pospisilik, J. A. et al. Drosophila genome-wide obesity screen reveals hedgehog as a determinant of brown versus white adipose cell fate. Cell 140, 148–160 (2010)
de Navas, L. F., Garaulet, D. L. & Sanchez-Herrero, E. The Ultrabithorax Hox gene of Drosophila controls haltere size by regulating the Dpp pathway. Development 133, 4495–4506 (2006)
Pfeiffer, S., Alexandre, C., Calleja, M. & Vincent, J. P. The progeny of wingless-expressing cells deliver the signal at a distance in Drosophila embryos. Curr. Biol. 10, 321–324 (2000)
Silies, M. et al. Glial cell migration in the eye disc. J. Neurosci. 27, 13130–13139 (2007)
Acknowledgements
We are grateful to A. Brand, S. Cohen, B. Edgar, U. Gaul, E. Hafen, C. Klambt, T. Lee, S. Leevers, P. Leopold, F. Matsuzaki, I. Miguel-Aliaga, T. Neufeld, R. Palmer, L. Partridge, L. Pick, E. Sanchez-Herrero, H. Stocker, N. Tapon and T. Xu, and also to the Bloomington stock centre and Kyoto National Institute of Genetics (NIG) for Drosophila stocks, antibodies and plasmids. We also acknowledge I. Salecker, J.-P. Vincent, A. Bailey, E. Cinnamon, L. Cheng, R. Makki, A. Matheu, P. Pachnis, P. Serpente and I. Stefana for providing advice, reagents and critical reading of the manuscript. The authors were supported by the Medical Research Council (U117584237).
Author information
Authors and Affiliations
Contributions
R.S.-N. and A.P.G. designed the experiments, R.S.-N. and L.L.Y. performed the experiments and R.S.-N. and A.P.G. wrote the manuscript. All authors have read and subscribe to the contents of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Tables 1-2 and Supplementary Figures 1-4 with legends. (PDF 1400 kb)
Rights and permissions
About this article
Cite this article
Sousa-Nunes, R., Yee, L. & Gould, A. Fat cells reactivate quiescent neuroblasts via TOR and glial insulin relays in Drosophila. Nature 471, 508–512 (2011). https://doi.org/10.1038/nature09867
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09867
This article is cited by
-
From bristle to brain: embryonic development of topographic projections from basiconic sensilla in the antennal nervous system of the locust Schistocerca gregaria
Development Genes and Evolution (2024)
-
Single cell RNA-seq analysis reveals temporally-regulated and quiescence-regulated gene expression in Drosophila larval neuroblasts
Neural Development (2022)
-
Drosophila septin interacting protein 1 regulates neurogenesis in the early developing larval brain
Scientific Reports (2022)
-
An interplay between cellular growth and atypical fusion defines morphogenesis of a modular glial niche in Drosophila
Nature Communications (2022)
-
Nutrition influences nervous system development by regulating neural stem cell homeostasis
Proceedings of the Indian National Science Academy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.