Abstract
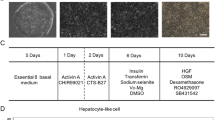
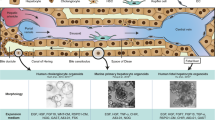
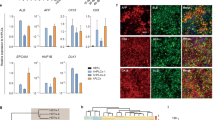
The generation of functional hepatocytes independent of donor liver organs is of great therapeutic interest with regard to regenerative medicine and possible cures for liver disease1. Induced hepatic differentiation has been achieved previously using embryonic stem cells or induced pluripotent stem cells2,3,4,5,6,7,8. Particularly, hepatocytes generated from a patient’s own induced pluripotent stem cells could theoretically avoid immunological rejection. However, the induction of hepatocytes from induced pluripotent stem cells is a complicated process that would probably be replaced with the arrival of improved technology. Overexpression of lineage-specific transcription factors directly converts terminally differentiated cells into some other lineages9,10,11,12, including neurons13, cardiomyocytes14 and blood progenitors15; however, it remains unclear whether these lineage-converted cells could repair damaged tissues in vivo. Here we demonstrate the direct induction of functional hepatocyte-like (iHep) cells from mouse tail-tip fibroblasts by transduction of Gata4, Hnf1α and Foxa3, and inactivation of p19Arf. iHep cells show typical epithelial morphology, express hepatic genes and acquire hepatocyte functions. Notably, transplanted iHep cells repopulate the livers of fumarylacetoacetate-hydrolase-deficient (Fah−/−) mice and rescue almost half of recipients from death by restoring liver functions. Our study provides a novel strategy to generate functional hepatocyte-like cells for the purpose of liver engineering and regenerative medicine.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Zern, M. A. Cell transplantation to replace whole liver transplantation. Gastroenterology 136, 767–769 (2009)
Gouon-Evans, V. et al. BMP-4 is required for hepatic specification of mouse embryonic stem cell–derived definitive endoderm. Nature Biotechnol. 24, 1402–1411 (2006)
Soto-Gutiérrez, A. et al. Reversal of mouse hepatic failure using an implanted liver-assist device containing ES cell-derived hepatocytes. Nature Biotechnol. 24, 1412–1419 (2006)
Duan, Y. et al. Differentiation and enrichment of hepatocyte-like cells from human embryonic stem cells in vitro and in vivo . Stem Cells 25, 3058–3068 (2007)
Basma, H. et al. Differentiation and transplantation of human embryonic stem cell-derived hepatocytes. Gastroenterology 136, 990–999 (2009)
Si-Tayeb, K. et al. Highly efficient generation of human hepatocyte-like cells from induced pluripotent stem cells. Hepatology 51, 297–305 (2010)
Sullivan, G. J. et al. Generation of functional human hepatic endoderm from human induced pluripotent stem cells. Hepatology 51, 329–335 (2010)
Li, F. et al. Hepatoblast-like progenitor cells derived from embryonic stem cells can repopulate livers of mice. Gastroenterology 139, 2158–2169 (2010)
Shen, C. N., Slack, J. M. & Tosh, D. Molecular basis of transdifferentiation of pancreas to liver. Nature Cell Biol. 2, 879–887 (2000)
Xie, H., Ye, M., Feng, R. & Graf, T. Stepwise reprogramming of B cells into macrophages. Cell 117, 663–676 (2004)
Zhou, Q., Brown, J., Kanarek, A., Rajagopal, J. & Melton, D. A. In vivo reprogramming of adult pancreatic exocrine cells to β-cells. Nature 455, 627–632 (2008)
Feng, R. et al. PU.1 and C/EBPα/β convert fibroblasts into macrophage-like cells. Proc. Natl Acad. Sci. USA 105, 6057–6062 (2008)
Vierbuchen, T. et al. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463, 1035–1041 (2010)
Ieda, M. et al. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 142, 375–386 (2010)
Szabo, E. et al. Direct conversion of human fibroblasts to multilineage blood progenitors. Nature 468, 521–526 (2010)
Kyrmizi, I. et al. Plasticity and expanding complexity of the hepatic transcription factor network during liver development. Genes Dev. 20, 2293–2305 (2006)
Zaret, K. S. Genetic programming of liver and pancreas progenitors: lessons for stem-cell differentiation. Nature Rev. Genet. 9, 329–340 (2008)
Schrem, H., Klempnauer, J. & Borlak, J. Liver-enriched transcription factors in liver function and development. Part I: the hepatocyte nuclear factor network and liver-specific gene expression. Pharmacol. Rev. 54, 129–158 (2002)
Schrem, H., Klempnauer, J. & Borlak, J. Liver-enriched transcription factors in liver function and development. Part II: the C/EBPs and D site-binding protein in cell cycle control, carcinogenesis, circadian gene regulation, liver regeneration, apoptosis, and liver-specific gene regulation. Pharmacol. Rev. 56, 291–330 (2004)
Mikula, M. et al. Immortalized p19ARF null hepatocytes restore liver injury and generate hepatic progenitors after transplantation. Hepatology 39, 628–634 (2004)
Grompe, M. et al. Loss of fumarylacetoacetate hydrolase is responsible for the neonatal hepatic dysfunction phenotype of lethal albino mice. Genes Dev. 7 (12A). 2298–2307 (1993)
Wang, X. et al. The origin and liver repopulating capacity of murine oval cells. Proc. Natl Acad. Sci. USA 100 (Suppl. 1). 11881–11888 (2003)
Grompe, M. et al. Pharmacological correction of neonatal lethal hepatic dysfunction in a murine model of hereditary tyrosinaemia type I. Nature Genet. 10, 453–460 (1995)
Overturf, K. et al. Hepatocytes corrected by gene therapy are selected in vivo in a murine model of hereditary tyrosinaemia type I. Nature Genet. 12, 266–273 (1996)
Zaret, K. S. et al. Pioneer factors, genetic competence, and inductive signaling: programming liver and pancreas progenitors from the endoderm. Cold Spring Harb. Symp. Quant. Biol. 73, 119–126 (2008)
Cirillo, L. A. et al. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol. Cell 9, 279–289 (2002)
Odom, D. T. et al. Control of pancreas and liver gene expression by HNF transcription factors. Science 303, 1378–1381 (2004)
Li, H. et al. The Ink4/Arf locus is a barrier for iPS cell reprogramming. Nature 460, 1136–1139 (2009)
Hui, L. et al. p38α suppresses normal and cancer cell proliferation by antagonizing the JNK-c-Jun pathway. Nature Genet. 39, 741–749 (2007)
Acknowledgements
We would like to thank X. Chen, J. Cen, L. Zhang and J. Yuan for technical support, and F. Tang and D. Li for comments. The laboratory of L.H. is funded by the National Science Foundation of China (91019014) and the Chinese Academy of Sciences (KSCX2-YW-R-241, XDA01010402 and the Hundred Talents Program). The laboratory of X.W. is supported by the National Science Foundation of China (30801115), Chinese National 863 Plan Project (2006AA02Z474) and National Key Basic Research and Development Program of China (2007CB947102 and 2009CB941100).
Author information
Authors and Affiliations
Contributions
L.H. conceived the project. P.H. performed most of the experiments. S.J. analysed the in vitro functions of iHep cells. H.S. analysed gene expression of iHep cells. L.H., X.W., P.H. and Z.H. designed the experiments for characterizing in vivo functions of iHep cells. P.H., Z.H., D.X., C.L. and Y.H. performed the in vivo experiments. L.H. and P.H. analysed the data. L.H., P.H. and X.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-3 and Supplementary Figures 1-13 with legends. (PDF 2651 kb)
Rights and permissions
About this article
Cite this article
Huang, P., He, Z., Ji, S. et al. Induction of functional hepatocyte-like cells from mouse fibroblasts by defined factors. Nature 475, 386–389 (2011). https://doi.org/10.1038/nature10116
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10116
This article is cited by
-
A robust reprogramming strategy for generating hepatocyte-like cells usable in pharmaco-toxicological studies
Stem Cell Research & Therapy (2023)
-
ETV2/ER71, the key factor leading the paths to vascular regeneration and angiogenic reprogramming
Stem Cell Research & Therapy (2023)
-
PC3T: a signature-driven predictor of chemical compounds for cellular transition
Communications Biology (2023)
-
Translational landscape of direct cardiac reprogramming reveals a role of Ybx1 in repressing cardiac fate acquisition
Nature Cardiovascular Research (2023)
-
Recent advances and future prospects in direct cardiac reprogramming
Nature Cardiovascular Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.