Abstract
Although thousands of large intergenic non-coding RNAs (lincRNAs) have been identified in mammals, few have been functionally characterized, leading to debate about their biological role. To address this, we performed loss-of-function studies on most lincRNAs expressed in mouse embryonic stem (ES) cells and characterized the effects on gene expression. Here we show that knockdown of lincRNAs has major consequences on gene expression patterns, comparable to knockdown of well-known ES cell regulators. Notably, lincRNAs primarily affect gene expression in trans. Knockdown of dozens of lincRNAs causes either exit from the pluripotent state or upregulation of lineage commitment programs. We integrate lincRNAs into the molecular circuitry of ES cells and show that lincRNA genes are regulated by key transcription factors and that lincRNA transcripts bind to multiple chromatin regulatory proteins to affect shared gene expression programs. Together, the results demonstrate that lincRNAs have key roles in the circuitry controlling ES cell state.
Similar content being viewed by others
Main
The mammalian genome encodes many thousands of large non-coding transcripts1 including a class of ∼3,500 lincRNAs identified using a chromatin signature of actively transcribed genes2,3,4. These lincRNA genes have been shown to have interesting properties, including clear evolutionary conservation2,3,4,5, expression patterns correlated with various cellular processes2,6 and binding of key transcription factors to their promoters2,6, and the lincRNAs themselves physically associate with chromatin regulatory proteins4,7. Yet, it remains unclear whether the RNA transcripts themselves have biological functions8,9,10. Few have been demonstrated to have phenotypic consequences by loss-of-function experiments6. As a result, the functional role of lincRNA genes has been widely debated. Various proposals include that lincRNA genes act as enhancer regions, with the RNA transcript simply being an incidental by-product8,9, that lincRNA transcripts act in cis to activate transcription11, and that lincRNA transcripts can act in trans to repress transcription12,13.
We therefore sought to undertake systematic loss-of-function experiments on all lincRNAs known to be expressed in mouse embryonic stem (ES) cells2,3. ES cells are pluripotent cells that can self-renew in culture and can give rise to cells of any of the three primary germ layers including the germ line14. The signalling14, transcriptional15,16,17 and chromatin15,18,19,20,21 regulatory networks controlling pluripotency have been well characterized, providing an ideal system to determine how lincRNAs may integrate into these processes.
Here we show that knockdown of the vast majority of ES-cell-expressed lincRNAs has a strong effect on gene expression patterns in ES cells, of comparable magnitude to that seen for the well-known ES cell regulatory proteins. We identify dozens of lincRNAs that upon loss-of-function cause an exit from the pluripotent state and dozens of additional lincRNAs that, although not essential for the maintenance of pluripotency, act to repress lineage-specific gene expression programs in ES cells. We integrate the lincRNAs into the molecular circuitry of ES cells by demonstrating that most lincRNAs are directly regulated by critical pluripotency-associated transcription factors and ∼30% of lincRNAs physically interact with specific chromatin regulatory proteins to affect gene expression. Together, these results demonstrate a regulatory network in ES cells whereby transcription factors directly regulate the expression of lincRNA genes, many of which can physically interact with chromatin proteins, affect gene expression programs and maintain the ES cell state.
lincRNAs affect global gene expression
To perform loss-of-function experiments, we generated five lentiviral-based short hairpin RNAs (shRNAs)22 targeting each of the 226 lincRNAs previously identified in ES cells2,3 (see Methods and Supplementary Table 1). These shRNAs successfully targeted 147 lincRNAs and reduced their expression by an average of ∼75% compared to endogenous levels in ES cells (see Methods, Fig. 1a, Supplementary Fig. 1 and Supplementary Table 2). As positive controls, we generated shRNAs targeting ∼50 genes encoding regulatory proteins, including both transcription and chromatin factors that have been shown to play critical roles in ES cell regulation17,20,23; validated hairpins were obtained against 40 of these genes (Supplementary Table 2). As negative controls, we performed independent infections with lentiviruses containing 27 different shRNAs with no known cellular target RNA.
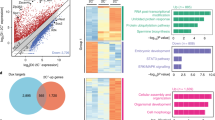
a, A schematic of lincRNA perturbation experiments. ES cells are infected with shRNAs, knockdown level is computed, the best hairpin is selected and profiled on expression arrays, and differential gene expression is computed relative to negative control hairpins. b, Example of a lincRNA knockdown. Top: genomic locus containing the lincRNA. Bottom: heat-map of the 95 genes affected by knockdown of the lincRNA, expression for control hairpins (red line) and expression for lincRNA hairpins (blue line) are shown. c, Distribution of number of affected genes upon knockdown of 147 lincRNAs (blue) and 40 well-known ES cell regulatory proteins (red). Points corresponding to five specific ES cell regulatory proteins are marked.
We infected each shRNA into ES cells, isolated RNA after 4 days, and profiled their effects on global transcription by hybridization to genome-wide microarrays (Fig. 1a, see Methods). We used a stringent procedure to control for nonspecific effects due to viral infection, generic RNA interference (RNAi) responses, or ‘off-target’ effects. Expression changes were deemed significant only if they exceeded the maximum levels observed in any of the negative controls, showed a twofold change in expression compared to the negative controls, and had a low false discovery rate (FDR) assessed across all genes based on permutation tests (Fig. 1b, see Methods). This approach controls for the overall rate of nonspecific effects by estimating the number and magnitude of observed effects in the negative control hairpins, where all effects are nonspecific.
For 137 of the 147 lincRNAs (93%), knockdown caused a significant impact on gene expression (Supplementary Table 3), with an average of 175 protein-coding transcripts affected (range: 20–936) (Fig. 1c, Supplementary Fig. 2 and Supplementary Table 4). These results were similar to those obtained upon knockdown of the 40 well-studied ES cell regulatory proteins: 38 (95%) showed significant effects on gene expression, with an average of 207 genes affected (range: 28 (for Dnmt3l) to 1,187 (for Oct4)) (Fig. 1c, Supplementary Fig. 2 and Supplementary Table 4). Although some individual lincRNAs have been found to lead primarily to gene repression12,13, we find that knockdown of the lincRNAs studied here largely led to comparable numbers of activated and repressed genes (Supplementary Fig. 2 and Supplementary Table 4). To assess off-target effects further, we also profiled the effects of the second-best validated shRNA targeting 10 randomly selected lincRNA genes. In all cases, second shRNAs against the same target produced significantly similar expression changes (see Methods and Supplementary Table 5). These results indicate that the vast majority of lincRNAs have functional consequences on overall gene expression of comparable magnitude (in terms of number of affected genes and impact on levels) to the known transcriptional regulators in ES cells.
lincRNAs affect gene expression in trans
Following the observation that a few lincRNAs act in cis24,25, some recent papers have claimed that most lincRNAs act primarily in cis8,11,26. We found no evidence to support this latter notion: knockdown of only 2 lincRNAs showed effects on a neighbouring gene, only 13 showed effects within a window of 10 genes on either side, and only 8 showed effects on genes within 300 kb; these proportions are no greater than observed for protein-coding genes (Supplementary Fig. 3 and Supplementary Table 6). In short, lincRNAs seem to affect expression largely in trans.
Our results contrast with a recent study that concluded that lincRNAs act in cis, based on the observation that knockdown of 7 out of 12 lincRNAs affected expression of a gene within 300 kb11. The explanation seems to be that the threshold in the previous study failed to account for multiple hypothesis testing within the local region. Accounting for this, the effects on neighbouring genes are no greater than expected by chance and are consistent with our observations here (see Methods).
Although some lincRNAs can regulate gene expression in cis11,24,25, determining the precise proportion of cis regulators requires more direct experimental approaches. We note that our results are consistent with observed correlations between lincRNAs and neighbouring genes2,26, which may represent shared upstream regulation2,12 or local transcriptional effects10,27. In addition, the lincRNAs studied here should be distinguished from transcripts that are produced at enhancer sites8,9, the function of which has yet to be determined.
lincRNAs maintain the pluripotent state
We next sought to investigate whether lincRNAs have a role in regulating the ES cell state. Regulation of the ES cell state involves two components: maintaining the pluripotency program and repressing differentiation programs15. To determine whether lincRNAs have a role in the maintenance of the pluripotency program, we studied their effects on the expression of Nanog, a key transcription factor that is required to establish28 and uniquely marks the pluripotent state29,30. We infected ES cells carrying a luciferase reporter gene expressed from the endogenous Nanog promoter31 with shRNAs targeting lincRNAs or protein-coding genes. We monitored loss of reporter activity after 8 days relative to 25 negative control hairpins across biological replicates (see Methods). To ensure that the observed effects were not simply due to a reduction in cell viability, we excluded shRNAs that caused a reduction in cell numbers (see Methods, Supplementary Fig. 4 and Supplementary Table 7). Altogether, we identified 26 lincRNAs that had major effects on endogenous Nanog levels with many at comparable levels to the knockdown of the known protein-coding regulators of pluripotency such as Oct4 and Nanog (Fig. 2a and Supplementary Table 7). This establishes that these lincRNAs have a role in maintaining the pluripotent state.
a, Activity from a Nanog promoter driving luciferase, following treatment with control hairpins (black) or hairpins targeting luciferase (green), selected protein-coding regulators (red), and lincRNAs (blue). b, Relative mRNA expression levels of Oct4 after knockdown of selected protein-coding (red) and lincRNA (blue) genes affecting Nanog-luciferase levels. The best hairpin (Hairpin 1) and second best hairpin (Hairpin 2) are shown. All knockdowns are significant with a P-value <0.01. Error bars represent standard error (n = 4). c, Morphology of ES cells and immunofluorescence staining of Oct4 for a negative control hairpin (black line) and hairpins targeting Oct4 (red line), and two lincRNAs (blue line). The first row shows bright-field images, the second row shows immunofluorescence staining of the Oct4 protein, and the third row shows DAPI staining of the nuclei.
To validate further the role of these 26 lincRNAs in regulating the pluripotent state, we knocked down these lincRNAs in wild-type ES cells and measured mRNA levels of pluripotency marker genes Oct4 (also called Pou5f1), Sox2, Nanog, Klf4 and Zfp42 after 8 days. In all cases we observed a significant reduction in the expression of multiple pluripotency markers with >90% showing a significant decrease in both Oct4 and Nanog levels (Supplementary Fig. 5 and Supplementary Tables 8 and 9). To control for off-target effects, we studied additional hairpins targeting these lincRNAs. For 15 lincRNAs we had an effective second hairpin. In all 15 cases, the second hairpin produced comparable reductions in Oct4 expression levels, showing that the observations were not due to off-target effects (Fig. 2b and Supplementary Table 10). Notably, >90% of lincRNA knockdowns affecting Nanog reporter levels led to loss of ES cell morphology (Fig. 2c and Supplementary Figs 6 and 7). Thus, inhibition of these 26 lincRNAs lead to an increased exit from the pluripotent state.
lincRNAs repress lineage programs
To determine if lincRNAs act in repressing differentiation programs we compared the overall gene expression patterns resulting from knockdown of the lincRNAs to published gene expression patterns resulting from induced differentiation of ES cells32,33 and assessed significance using a permutation-derived FDR34 (see Methods). These states include differentiation into endoderm, ectoderm, neuroectoderm, mesoderm and trophectoderm lineages. As a positive control for our analytical method, we confirmed the expected results that the expression pattern caused by Oct4 knockdown was strongly associated with the trophoectoderm lineage35 and the pattern caused by Nanog knockdown was strongly associated with endoderm differentiation30 (Fig. 3a).
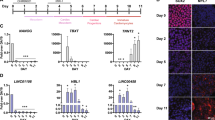
a, Expression changes for each lincRNA compared to gene expression of five differentiation patterns. Each box shows significant positive association (red, FDR <0.01) for Oct4 and Nanog (left) and for lincRNAs (right). b, Expression changes upon knockdown of Oct4 and Nanog (black bars) and representative lincRNAs (grey bars) for five lineage marker genes. The expression changes (FDR <0.05) are displayed on a log scale as the t-statistic compared to a panel of negative control hairpins.
Using this approach, we identified 30 lincRNAs for which knockdown produced expression patterns similar to differentiation into specific lineages (Supplementary Table 11). Among these lincRNAs, 13 are associated with endoderm differentiation, 7 with ectoderm differentiation, 5 with neuroectoderm differentiation, 7 with mesoderm differentiation and 2 with the trophectoderm lineage (Fig. 3a). Consistent with these functional assignments, we observed that most (>85%) of the 30 lincRNAs associated with specific differentiation lineages showed upregulation of the well-known marker genes for the identified states17,32 upon knockdown (such as Sox17 (endoderm), Fgf5 (ectoderm), Pax6 (neuroectoderm), brachyury (mesoderm) and Cdx2 (trophectoderm)) (Fig. 3b, Supplementary Figs 8 and 9 and Supplementary Tables 12 and 13).
The fact that knockdown of these 30 lincRNAs induces gene expression programs associated with specific early differentiation lineages indicates that these lincRNAs normally are a barrier to such differentiation. Interestingly, most of the lincRNA knockdowns (∼85%) that induce gene expression patterns associated with these lineages did not cause the cells to differentiate as determined by Nanog reporter levels (Supplementary Table 7) and Oct4 expression (Supplementary Fig. 10). This is consistent with observations for several critical ES cell chromatin regulators, such as the polycomb complex; loss-of-function of these regulators similarly induces lineage-specific markers without causing differentiation18,36,37.
Together, these data indicate that many lincRNAs have important roles in regulating the ES cell state, including maintaining the pluripotent state and repressing specific differentiation lineages.
lincRNAs are targets of ES cell transcription factors
Having demonstrated a functional role for lincRNAs in ES cells, we sought to integrate the lincRNAs into the molecular circuitry controlling the pluripotent state. First, we explored how lincRNA expression is regulated in ES cells. Towards this end, we used published genome-wide maps of 9 pluripotency-associated transcription factors16,38 and determined whether they bind to the promoters of lincRNA genes. Of the 226 lincRNA promoters ∼75% are bound by at least 1 of 9 pluripotency-associated transcription factors (including Oct4, Sox2, Nanog, c-Myc, n-Myc, Klf4, Zfx, Smad and Tcf3) with a median of 3 factors bound to each promoter (Fig. 4a, Supplementary Fig. 11 and Supplementary Table 14), comparable to the proportion reported for protein-coding genes16. Interestingly, the three core factors (Oct4, Sox2 and Nanog) bind to the promoters of ∼12% of all ES cell lincRNAs and ∼50% of lincRNAs involved in the regulation of the pluripotent state.
a, A heat-map representing ChIP-Seq enrichments for nine transcription factors (columns) at lincRNA promoters (rows). The percentage of bound lincRNAs downregulated upon knockdown of the transcription factor is indicated in boxes. NA, not measured. Right: examples of lincRNAs from two clusters (‘core regulated’ and ‘Myc regulated’) showing their genomic neighbourhood and transcription factor binding. b, Left: a heat-map representing changes in lincRNA expression (rows) after knockdown of 11 transcription factors (columns). Middle: effect of knockdown of Sox2, Oct4 and Nanog on expression levels of linc1405 (grey) and Oct4 (black). Right: effect of knockdown of Klf2, Klf4, n-Myc and Esrrb on expression levels of linc1428.
To determine if lincRNA expression is functionally regulated by the pluripotency-associated transcription factors, we used shRNAs to knockdown the expression of 5 of the 9 pluripotency-associated transcription factor genes for which we could obtain validated hairpins and profiled the resulting changes in lincRNA expression after 4 days. Upon knockdown of a transcription factor, ∼50% of lincRNA genes whose promoters are bound by the transcription factor exhibit expression changes (Fig. 4a); this proportion is comparable to that seen for protein-coding genes whose promoters are bound by the transcription factor (Supplementary Fig. 12). The strong but imperfect correlation between transcription-factor-binding and effect of transcription-factor knockdown is consistent with previous observations39 and may reflect regulatory redundancy in the pluripotency network40. In addition, we profiled the knockdown of an additional 7 pluripotency-associated transcription factors (including Esrrb, Zfp42 and Stat3). Altogether, for ∼60% of the ES cell lincRNAs, we identified a significant downregulation upon knockdown of 1 of these 11 transcription factors (Fig. 4b and Supplementary Table 15).
After retinoic-acid-induced differentiation of ES cells, the ES cell lincRNAs show temporal changes across the time course with ∼75% showing a decrease in expression compared to untreated ES cells (Supplementary Fig. 13 and Supplementary Table 16). Notably, all of the lincRNAs shown to regulate pluripotency are downregulated upon retinoic acid treatment (Supplementary Fig. 13). Our results establish that lincRNAs are direct transcriptional targets of pluripotency-associated transcription factors and are dynamically expressed across differentiation. Collectively, these results demonstrate that lincRNAs are an important regulatory component within the ES cell circuitry.
lincRNAs bind diverse chromatin proteins
To explore how lincRNAs carry out their regulatory roles, we studied whether lincRNAs physically associate with chromatin regulatory proteins in ES cells. We previously showed that many human lincRNAs can interact with the polycomb repressive complex4, a complex that has a critical functional role in the regulation of ES cells18,19. To determine whether the ES cell lincRNAs physically associate with the polycomb complex, we crosslinked RNA–protein complexes using formaldehyde, immunoprecipitated the complex using antibodies specific to both the Suz12 and Ezh2 components of polycomb, and profiled the co-precipitated lincRNAs using a direct RNA quantification method41 (see Methods). We performed immunoprecipitation of the polycomb complex across five biological replicates and eight mock-IgG controls, and we assessed significance using a permutation test (see Methods and Supplementary Fig. 16). Altogether, we identified 24 lincRNAs (∼10% of the ES cell lincRNAs) that were strongly enriched for both polycomb components (Fig. 5b and Supplementary Table 17).
a, A schematic of the classes of chromatin regulators profiled: readers (blue), writers (orange) and erasers (green). b, A heat-map showing the enrichment of 74 lincRNAs (rows) for 1 of 12 chromatin regulatory complexes (columns). The names are colour-coded by chromatin-regulatory mechanism. Major clusters are indicated by vertical lines with a description of the chromatin components.
To determine if lincRNAs interact with additional chromatin proteins, we systematically analysed chromatin-modifying proteins that have been shown to have critical roles in ES cells18,19,20,21,42. Specifically, we screened antibodies against 28 chromatin complexes (see Methods, Supplementary Fig. 14 and Supplementary Table 18) and identified 11 additional chromatin complexes that are strongly and reproducibly associated with lincRNAs (see Methods and Supplementary Figs 15 and 16). These chromatin complexes are involved in ‘reading’ (Prc1, Cbx1 and Cbx3), ‘writing’ (Tip60/P400, Prc2, Setd8, Eset and Suv39h1) and ‘erasing’ (Jarid1b, Jarid1c, and Hdac1) histone modifications, as well as a chromatin-associated DNA binding protein (Yy1) (Fig. 5a). Altogether, we found that 74 (∼30%) of the ES cell lincRNAs are associated with at least 1 of these 12 chromatin complexes (Fig. 5b and Supplementary Table 17). Although most of the identified interactions are with repressive chromatin regulators, this is probably due to limitations of our selection criteria and available antibodies.
Many lincRNAs are strongly associated with multiple chromatin complexes (Fig. 5b). For example, we identified 8 lincRNAs that bind to the Prc2 H3K27 and Eset H3K9 methyltransferase complexes (writers of repressive marks) and the Jarid1c H3K4 demethylase complex (an eraser of activating marks). Consistent with this, the Prc2 and Eset complexes have been reported to bind at many of the same ‘bivalent’ domains21 and to associate functionally with the Jarid1c complex43. Similarly, we identified a distinct set of 17 lincRNAs that bind to the Prc2 complex (‘writer’ of K27 repressive marks), Prc1 complex (‘reader’ of K27 repressive marks) and Jarid1b complex (‘eraser’ of K4 activating marks) (Fig. 5b), as well as other functionally consistent reader, writer and eraser combinations (Supplementary Fig. 17). One of several potential models consistent with these data are that lincRNAs may bind to multiple distinct protein complexes, perhaps serving as ‘flexible scaffolds’ to bridge functionally related complexes as previously described for telomerase RNA44.
To determine if the identified lincRNA–protein interactions have a functional role, we examined the effects on gene expression resulting from knockdown of individual lincRNAs that are physically associated with particular chromatin complexes and from knockdown of genes encoding the associated complex itself (see Methods). For >40% of these lincRNA–protein interactions, we identified a highly significant overlap in affected gene expression programs compared to just ∼6% for random lincRNA–protein pairs (see Methods and Supplementary Table 19). Other cases may reflect the limited power to detect the overlaps, because specific lincRNA–protein complexes may be related to only a fraction of the overall expression pattern mediated by the chromatin complex.
Together, these data demonstrate that many ES cell lincRNAs physically associate with multiple different chromatin regulatory proteins and that these interactions are probably important for the regulation of gene expression programs.
Discussion
Although the mammalian genome encodes thousands of lincRNA genes, few have been functionally characterized. We performed an unbiased loss-of-function analysis of lincRNAs expressed in ES cells and show that lincRNAs are clearly functional and primarily act in trans to affect global gene expression. We establish that lincRNAs are key components of the ES cell transcriptional network that are functionally important for maintaining the pluripotent state, and that many are downregulated upon differentiation. The ES cell lincRNAs physically interact with chromatin proteins, many of which have been previously implicated in the maintenance of the pluripotent state18,20,21. In addition to chromatin proteins, lincRNAs interact with other protein complexes including many RNA-binding proteins (data not shown).
Our data suggest a model whereby a distinct set of lincRNAs is transcribed in a given cell type and interacts with ubiquitous regulatory protein complexes to form cell-type-specific RNA–protein complexes that coordinate cell-type-specific gene expression programs (Fig. 6). Because many of the lincRNAs studied here interact with multiple different protein complexes, they may act as cell-type-specific ‘flexible scaffolds’44 to bring together protein complexes into larger functional units (Fig. 6). This model has been previously demonstrated for the yeast telomerase RNA44 and suggested for the XIST45 and HOTAIR46 lincRNAs. The hypothesis that lincRNAs serve as flexible scaffolds could explain the uneven patterns of evolutionary conservation seen across the length of lincRNA genes3: the more highly conserved patches could correspond to regions of interaction with protein complexes.
ES-cell-specific transcription factors (such as Oct4, Sox2 and Nanog) bind to the promoter of a lincRNA gene and drive its transcription. The lincRNA binds to ubiquitous regulatory proteins, giving rise to cell-type-specific RNA–protein complexes. Through different combinations of protein interactions, the lincRNA–protein complex can give rise to unique transcriptional programs. Right: a similar process may also work in other cell types with specific transcription factors regulating lincRNAs, creating cell-type-specific RNA–protein complexes and regulating cell-type-specific expression programs.
Although a model of lincRNAs acting as ‘flexible scaffolds’ is attractive, it is far from proven. Testing the hypothesis for lincRNAs will require systematic studies, including defining all protein complexes with which lincRNAs interact, determining where these protein interactions assemble on RNA, and ascertaining whether they bind simultaneously or alternatively. Moreover, understanding how lincRNA–protein interactions give rise to specific patterns of gene expression will require determination of the functional contribution of each interaction and possible localization of the complex to its genomic targets.
Methods Summary
RNAi expression effects
We cloned five shRNAs targeting each lincRNA into a puromycin-resistant lentiviral vector22. ES cells were plated on pre-gelatinized 96-well plates and infected with lentivirus before addition of irradiated DR4 mouse embryonic fibroblasts (MEFs). Media containing 1 μg ml−1 puromycin was added 24 h after infection. On-target knockdown was assessed after 4 days and the best hairpin showing a knockdown >60% was selected. RNA from 147 lincRNAs, 40 protein-coding genes and 27 negative controls were hybridized to Agilent microarrays. Differentially expressed genes were defined as having an FDR <5% and fold-change >2-fold compared to controls.
Screening for pluripotency effects
Nanog-luciferase ES cells31 were infected and measured after 8 days. Hits were identified if they reduced luciferase levels (z < −6) across all replicates and did not reduce AlamarBlue levels. Hits were validated in wild-type ES cells by measuring mRNA levels of Oct4, Nanog, Sox2, Klf4 and Zfp42. Oct4 expression was assessed using immunofluorescence staining and morphology was visually assessed.
Lineage expression effects
Lineage expression programs were defined using published data sets (Gene Expression Omnibus GSE12982, GSE11523, and GSE4082) and curated gene expression signatures32,33. Overlaps in gene expression effects were assessed using a modified GSEA34. Expression changes in lineage markers were determined using qPCR.
Transcription factor binding and regulation
ChIP-Seq data was downloaded (GSE11724 and GSE11431), aligned and analysed. lincRNA promoters were previously defined using H3K4me3 peaks3. Changes in expression of the lincRNAs upon knockdown of the transcription factors were analysed using Agilent microarrays.
Chromatin binding and overlap in expression
ES cells were crosslinked with formaldehyde, lysed, immunoprecipitated, washed and reverse crosslinked. RNA was hybridized to the Nanostring code set. We tested antibodies for 28 chromatin complexes and selected successful antibodies that had >10 lincRNAs exceeding a fivefold change and had significant enrichments across 3 replicates. We compared the overlap in gene expression using a modified GSEA34.
Online Methods
ES cell culture
V6.5 (genotype 129SvJae × C57BL/6) and Nanog-luciferase31 ES cells were co-cultured with irradiated C57BL/6 MEFs (GlobalStem; GSC-6002C) on pre-gelatinized plates as previously described47. Briefly, cells were cultured in mES media consisting of knockout DMEM (Invitrogen; 10829018) supplemented with 10% FBS (GlobalStem; GSM-6002), 1% penicillin-streptomycin (Invitrogen; 15140-163), 1% l-glutamine (Invitrogen; 25030-164), 0.001% β-mercaptoethanol (Sigma; M3148-100ML) and 0.01% ESGRO (Millipore; ESG1106).
Picking lincRNA gene candidates
Using our previous catalogue of K4-K36 defined lincRNAs2 along with the reconstructed full-length sequences we determined using RNA-Seq3, we designed shRNA hairpins targeting each lincRNA identified in both sets. Specifically, we used the conservative K4-K36 definitions from our previous work2 that were expressed in mouse ES cells. We further filtered the list to include only multi-exonic lincRNAs that were reconstructed in mouse ES cells3. Together, this yielded 226 lincRNA genes.
Picking protein-coding gene candidates
We selected protein coding gene controls consisting of both transcription factors and chromatin proteins. These proteins were selected based on their well-characterized role in regulating mouse ES cells and include Oct4 (Pou5f1)35,48, Sox2 (refs 17, 49) Nanog (refs 29, 30), Stat3 (ref. 50), Klf4 (ref. 51) and Zfp42 (Rex1)52. In addition, we selected additional transcriptional and chromatin regulators that were identified by RNAi screens as regulators of pluripotency17,20,23 and/or were found in smaller focused studies to have critical roles in the maintenance of the pluripotent state (such as Carm1 (ref. 53), Chd1 (ref. 54), Thap11 (ref. 55), Suz12 (refs 18, 19, 36) and Setdb1 (refs 21, 56)). A full list is provided in Supplementary Table 2.
shRNA design rules
For each lincRNA we designed five hairpins by extending the previously described design rules22 accounting for the sequence content of the hairpin, miRNA seed matches, uniqueness to the target compared to the transcriptome and the genome, and number of lincRNA isoforms covered.
For each lincRNA we enumerated all 21-mer sub-sequences and scored them as follows: (1) a ‘clamp score’ was computed by looking at the nucleotides at positions 18, 19 and 20. If all three positions contained an A/T it was assigned a score of 4, if two positions were A/T it was assigned a score of 1.5 and if one was A/T it was assigned a score of 0.8. We then looked at positions 16, 17, and 21; if all three were A/T it was assigned a score of 1.25, if two were A/T it was assigned a score of 1.1, and if one was A/T is was assigned a score of 0.8. The clamp score was computed as the product of these two scores. (2) A ‘GC score’ was computed by looking at the total GC percentage of the 21-mer sequence. If the sequence was <25% GC it was assigned a score of 0.01, if it was <55% it was assigned a score of 3, if it was <60% it was assigned a score of 1, and if >60% it was assigned a score of 0.01. (3) A ‘4-mer penalty’ of 0.01 was assigned for any hairpin containing the same nucleotide in 4 subsequent nucleotides. (4) A ‘7 GC penalty’ of 0.01 was assigned to any hairpin containing any 7 consecutive G/C nucleotides. (5) We removed all hairpins containing an A in either position 1 or position 2 of the hairpin. (6) We removed all hairpins containing a repeat masked nucleotide. (7) Finally, we computed a ‘miRNA-seed penalty’ by looking at the forward positions 11–17, 12–20 and 13–19 of the hairpin as well as the reverse complement of positions 14–20, 15–21, or 16–21 plus a 3′ C. We then looked up whether these positions matched known miRNA seeds and with what frequency. We computed the scores for the forward and reverse positions and defined the score as the product of the forward and reverse scores. The final score for each hairpin sequence is defined as the product of all seven scores.
We then sorted the candidate hairpin sequences by score, breaking high-scoring ties by the total number of lincRNA isoforms that are covered by the hairpin. We then aligned each hairpin sequence against both the genome and the RefSeq-defined transcriptome (NCBI Release 39), and filtered any hairpin with fewer than three mismatches to any other gene or position in the genome. Candidate sequences were chosen for shRNA production by first picking the highest scoring candidate and then proceeding to successively lower scores. As each hairpin was selected, all other hairpins overlapping this hairpin were removed. We repeated this process until we identified five hairpins that covered each lincRNA.
shRNA cloning and virus prep
We designed 1,143 hairpins targeting 226 lincRNA genes. Of these, we successfully cloned 1,010 hairpins targeting 214 lincRNAs. These hairpins were cloned into a vector containing a puromycin resistance gene and incorporated into a lentiviral vector as previously described22. Briefly, synthetic double-stranded oligos that represent a stem-loop hairpin structure were cloned into the second-generation TRC (the RNAi Consortium) lentiviral vector, pLKO.5; the expression of a given hairpin produces a shRNA that targets the gene of interest. Lentivirus was prepared as previously described22. Briefly, 100 ng of shRNA plasmid, 100 ng of packaging plasmid (psPAX2) and 10 ng of envelope plasmid (VSV-G) were used to transfect packaging cells (293T) with TransIT-LT1 (Mirus Bio). Virus was harvested 48 and 70 h after transfection. Two harvests were combined. Virus titres were measured as previously described22. Briefly, we measured virus titres by infecting A549 cells with appropriately diluted viruses. Twenty-four hours after infection, puromycin was added to a final concentration of 5 μg ml−1 and the selection proceeded for 48 h. The number of surviving cells, which is correlated to virus titre, was measured by AlamarBlue (BioSource) staining using the Envision 2103 Multilabel plate reader (PerkinElmer).
Infection and selection protocol
V6.5 ES cells or Nanog-luciferase ES cells were plated at a density of 5,000 cells per well (8-day time point) or 25,000 cells per well (4-day time point) in 100 μl mES media onto pre-gelatinized 96-well dishes (VWR; BD356689). Cells were infected with 5 μl of a lentiviral shRNA stock and incubated at 37 °C for 30 min. Puromycin-resistant DR4 MEFs (GlobalStem; GSC-6004G) were then added to the plates at a density of ∼6,000 cells per well and incubated overnight at 37 °C, 5% CO2. After 24 h, all media was removed from the cells and replaced with media containing 1 μg ml−1 puromycin. Media was then changed every other day with fresh media containing 1 μg ml−1 puromycin. The end-point depended on the assay and was either 4 days after infection (knockdown validation and microarrays) or 8 days (reporters and qPCR of marker genes).
RNA extraction
ES cells were infected and lysed at day 4 with 150 μl of Qiagen’s RLT buffer and three replicates of each virus plate were pooled for RNA extraction using Qiagen’s RNeasy 96-well columns (74181). RNA extraction was completed following Qiagen’s RNeasy 96-well protocol with the following modifications: 450 μl of 70% ethanol was added to 450 μl total lysate before the first spin. An additional RPE wash was added to the protocol, for a total of three RPE washes.
lincRNA primer design and pre-screen
lincRNA primers were designed using primer3 (http://frodo.wi.mit.edu/primer3/). Specifically, we designed primers spanning exon–exon junctions by specifying each of the regions as preferred inclusion regions in the primer3 program. When a low-scoring primer pair (primer penalty <1) was available it was used. If none was available, we then identified all primers that contained amplicons that spanned an exon–exon junction. In a few cases, when we could not identify a primer pair spanning an exon–exon junction, we designed primers within an exon of the lincRNA. For each primer pair, we tested the specificity against the transcriptome57 (RefSeq NCBI Release 39) and the genome (Mouse MM9) using the isPCR (http://genome.ucsc.edu/cgi-bin/hgPcr) program. Specifically, we required that the primer pair amplify the lincRNA gene and no other genomic of gene amplicon.
For each primer pair, we validated the quantification and specificity before use. Specifically, we tested primers in qPCR reactions using a dilution series of mouse ES cDNA including a no reverse transcriptase (RT) sample. We excluded any primer that did not have robust quantification across a 64-fold dilution curve, had high signal in the no RT sample, or had low detectable expression in the undiluted sample (cycle number >34). For primers that failed this validation we redesigned and tested new primers.
Knockdown validation using qPCR
To determine if lincRNA hairpins were effective at knocking down the lincRNA of interest, we infected each hairpin into mouse embryonic stem cells, selected for lentiviral integration, and measured changes in the targeted lincRNA expression level. We isolated total cellular RNA after 4 days; this time point was chosen to allow for identification of robust changes while minimizing secondary effects due to differentiation of the ES cells. We reasoned that this would allow us to determine more direct effects due to RNAi rather than to differentiation.
Gene panels were constructed that contained all five hairpins targeting a gene along with an empty vector control pLKO.5-nullT and the GFP-targeting hairpin clonetechGfp_437s1c1. cDNA was generated using 10 μl of RNA and 10 μl of 2× cDNA master mix containing 5× Transcriptor RT Reaction Buffer (Roche), DTT, MMLV-RT (Roche), dNTPs (Agilent; 200415-51), Random 9-mer oligos (IDT), Oligo-dT (IDT) and water. cDNA was diluted 1:9 and quantitative PCR was performed using 250 nM of each primer in 2× Sybr green master mix (Roche) and run on a Roche Light-Cycler 480. Target lincRNA expression and Gapdh levels were computed for each panel. lincRNA expression levels were normalized by Gapdh levels and this normalized value was compared to the reference control hairpins within the panel. Knockdown levels were computed as the average of the fold decrease compared to the two control hairpins. Hairpins showing a knockdown greater than 60% of the endogenous level were considered validated and the best validated hairpin from a lincRNA panel was selected for microarray studies.
Picking candidates for microarray analysis
To assess the effects of a lincRNA on gene expression, we profiled the changes in gene expression after knocking down each lincRNA gene. Specifically, for each lincRNA with at least one validated hairpin we profiled the genome-wide expression level changes after knockdown across two independent infections (see above). To control for expression changes due to viral infection, we performed five independent infections containing no RNAi hairpin (pLKO.5-nullT). This control hairpin was embedded in each RNA preparation plate. To control for effects due to an off-target RNAi effect, we profiled 27 distinct negative control hairpins which do not have a known target in the cell. These hairpins included 6 RFP hairpins, 10 GFP hairpins, 6 luciferase hairpins and 5 LacZ hairpins. These hairpins provide a measurement of the variability of the RNAi response triggered due to nonspecific effects. Furthermore, we profiled hairpins targeting 147 lincRNAs, including 10 with a second best hairpin, and 40 protein-coding genes in biological replicate. The hairpins and their replicates were randomly distributed across 7 96 well plates and prepared in batches. Each RNA preparation batch contained one pLKO hairpin and one clonetechGfp_437s1c1 hairpin in a random location on the plate. To minimize batch effects, the plate locations of the biological replicates were scrambled and the positions within the plates were scrambled for all hairpins and replicates.
Agilent microarray hybridization
Using Agilent’s One-Colour Quick Amp Labelling kit (5190-0442), we amplified and labelled total RNA for hybridization to prototype mouse lincRNA arrays (G4140-90040) according to manufacturer’s instructions with a few variations. The custom Agilent SurePrint G3 8x60K mouse array design used for this study (G4102A, AMADID 025725 G4852A) has probes to 21,503 Entrez genes and 2,230 lincRNA genes. A new updated version of this mouse design is commercially available that contains probes to 34,017 Entrez gene targets as well as 2,230 lincRNA genes (G4825A). The cRNA samples were prepared by diluting 200 ng of RNA in 8.3 μl water and adding positive control one-colour RNA spike-in mix (Agilent, 5188-5282) that was diluted serially 1:20, then 1:25 and finally 1:10. We annealed the T7 promoter primer from the kit by incubating at 65 °C for 10 min. We prepared the cDNA master mix and added it to the annealed RNA and incubated at 40 °C for 2 h, followed by 65 °C for 15 min. We prepared the cRNA transcription master mix and added it to the cDNA and incubated at 40 °C for 2 h protected from light. We purified the labelled cRNA using Qiagen’s RNeasy 96-well columns (Qiagen, 74181) by adding 350 μl of Qiagen RLT (without BME) to the cRNA followed by the addition of 250 μl of 95% ethanol before applying to the plate column. After a 4 min spin at 6,000 r.p.m., we washed the columns three times with 800 μl buffer RPE. We dried the columns by spinning for 10 min and eluted the cRNA with 50 μl of water. We measured the cRNA yield and dye incorporation using the Nanodrop 8000 Microarray measurement setting. We mixed 600 ng of cRNA with blocking agent and fragmentation buffer (Agilent, 5190-0404) and fragmented for 30 min in the dark at 60 °C. We added 2× hybridization buffer to each sample and loaded 40 μl onto an 8-pack Hybridization gasket. We placed the microarray slides on top, sealed in the hybridization chamber, and incubated for 18 h at 65 °C. We washed the slides for 1 min in room temperature GE Wash Buffer 1 and then for 1 min in 37 °C GE Wash Buffer 2 (Agilent 5188-5327, no triton addition). We scanned the microarrays using an Agilent Scanner C (G2565CA) using the following settings: dye channel = red & green, scan region = scan area (61 × 21.6 mm), scan resolution = 3 μm. We prepared all of the samples simultaneously using homogenous master mixes to limit variability. Fragmentation and hybridization was staggered over time in batches of 3 to 4 slides (24 to 32 samples).
Array filtering, normalization and probe filtering
Each array was processed and data extracted using the Agilent feature extraction software (G4462AA, Version 10.7.3). Samples were retained if they passed all the following quality control statistics: AnyColourPrcntFeatNonUnifOL <1; eQCOneColourSpikeDetectionLimit >0.01 and <2.0; Metric_absGE1E1aSlope between 0.9 and 1.2; Metric_gE1aMedCVProcSignal <8; gNegCtrlAveBGSubSig >−10 and <5; Metric_gNegCtrlAveNetSig <40; gNegCtrlSDevBGSubSig <10; Metric_gNonCntrlMedCVProcSignal <8; Metric_gSpatialDetrendRMSFilterMinusFit <15; SpotAnalysis_PixelSkewCookiePct >0.8 and <1.2.
Gene expression values were determined using the gProcessedSignal intensity values. Probes were flagged if they were not detectable well above background or had an expression level lower than the lowest detectable spike-in control value. The values were floored across all samples by taking the maximum of the minimum non-flagged values across all experiments. Any value less than this maximum value was set to the maximum. This conservatively eliminates any detection variability across the samples due to stringency or other array variables.
The result of this is a single value for each probe per array. To normalize expression values across arrays, we performed quantile normalization as previously described58. Briefly, we ranked each array from lowest to highest expression. For each rank, we computed the average expression and each experiment with this value at the associated rank. For each probe, we computed the difference between the second smallest expression value and the second largest expression value. If this difference was less than 2, we filtered the probe. This metric was chosen to eliminate bias due to single sample outliers.
Identifying significant gene expression hits from RNAi knockdowns
To control for effects due to nonspecific effects of shRNAs, we profiled 27 distinct negative control hairpins which do not have a known target in the cell. These hairpins provide a measurement of the variability of the expression profiles due to random variability or triggered by ‘off-target’ effects of the shRNA lentiviruses. Assuming that any observed effects in the negative control hairpins are due to off-target effects and observed effects in the targeting hairpins include a mix of both off-target effects and on-target effects, we use permutations of the negative controls to assign a FDR confidence level for being an on-target hit to each gene. As such, a gene would only reach genome-wide significance if the number of genes and scale of the effect was much larger than would be observed randomly among all of the expression changes found for the negative control hairpin.
Specifically, for each gene we computed a t-statistic between shRNAs targeting the lincRNA and control shRNA samples. To assess the significance of each gene we permuted the sample and control groups retaining the relative sizes of the groups and computing the same t-statistic. We then assigned an FDR value to each gene by computing the average number of values in the permuted t-statistics that were greater than the observed value of interest and divided this by the number of all observed t-statistics that were greater than the observed value. We defined genes as significantly differentially expressed if the FDR was <5% and the fold-change compared to the negative controls was >2-fold. Using this approach, an effect would only reach a significant FDR if the scale is significantly larger than would be observed in the negative controls. Knockdown of a lincRNA was considered to have a significant effect on gene expression if we identified at least 10 genes that had an effect that passed all of the criteria.
Gene-neighbour analysis
We identified neighbouring genes based on the RefSeq genome annotation57 (NCBI Release 39). We excluded from analysis all RefSeq genes that corresponded to our lincRNA of interest but included all other coding and non-coding transcripts. We identified a significant hit as any lincRNA affecting a neighbour within 10 genes on either side with an FDR<0.05 and twofold expression change. To compute the closest affected neighbour, we classified all genes affected upon knockdown of the lincRNAs using the same criteria above. We computed the distance between each affected gene and the locus of the lincRNA gene (and protein-coding gene) that was perturbed and took the minimum absolute distance across all affected genes.
Analysis of expected number of neighbouring genes that will change by chance
To determine the expected number of differentially expressed ‘neighbouring’ genes occurring by chance assuming that the knockdown has no effect on gene expression, we calculated the average number of genes in a 300-kb window around a randomly selected gene in the human and mouse genome. We calculated this to be 11.2 (human) and 11.8 (mouse). For simplicity, we will conservatively round this down to 11. Assuming that no genes are changing between the knockdown and control, using a nominal P-value, which has a uniform distribution under the null hypothesis (nothing effected), we would expect to see a difference called in 5% of cases at a P-value of 0.05. If we test one locus, which has on average 11 neighbours, we would expect to identify 0.55 hits by chance (11 × 0.05 = 0.55). However, if we now test 12 loci we would expect to see 6.6 (12 × 0.55) knockdowns that appear to have an effect under the null hypothesis.
Luciferase analysis of Nanog ES lines
ES cells containing a Nanog-luciferase construct31 were infected in biological duplicate and monitored after 7 days. Luciferase activity was measured using Bright-Glo (Promega). All reagents and cells were equilibrated to room temperature. 100 μl Bright-Glo solution was added to each plate well. Plates were incubated in the dark at room temperature for 10 min and luciferase was measured on a plate reader. The luciferase units were normalized to the control hairpins and a Z-score compared to the negative controls (excluding luciferase hairpins) was computed. For each hairpin, we computed a Z-score relative to the negative control hairpins and identified hits reducing luciferase levels more than 6 standard deviations (Z < −6) for both independent replicates. In all cases we were able to identify a significant reduction in luciferase levels when using distinct hairpins targeting luciferase. To exclude hits that were due to an overall reduction in proliferation (which would also cause a reduction of Nanog positive cells in this read-out) we excluded all hairpins that caused a reduction in proliferation as measured by AlamarBlue incorporation (described below). AlamarBlue incorporation was measured in the same cells immediately before reading out Nanog-luciferase levels.
AlamarBlue analysis of ES lines
After a 7-day infection, Nanog-luciferase cell viability was measured using AlamarBlue (Invitrogen; DAL1025). AlamarBlue was mixed with mES media in a 1:10 ratio, added to the cells and incubated at 37 °C for 1 h. Absorbance readings at 570 nm were taken. To control for possible effects due to virus titre, we measured AlamarBlue incorporation on both puromycin treated and non-puromycin treated samples for each infection.
mRNA analysis of pluripotency markers
V6.5 ES cells were infected with shRNAs targeting lincRNAs, protein-coding genes, and 21 negative controls. After 8 days, RNA was extracted and mRNA levels of the Oct4, Nanog, Sox2, Klf4 and Zfp42 pluripotency markers were analysed using qPCR. Primer sequences are listed in Supplementary Table 9. Each sample was normalized to Gapdh levels. Significance was assessed compared to the negative control hairpins using a one-tailed t-test.
To control for off-target effects, we analysed additional hairpins against the 26 lincRNAs affecting Nanog-luciferase levels. Of the 26 lincRNAs, we identified 15 lincRNAs that contained an additional hairpin that reduced lincRNA expression by >50%. V6.5 ES cells were infected with the best and additional hairpin across biological replicates for these 15 lincRNAs and 21 negative control hairpins. RNA was extracted after 8 days and Oct4 expression levels were determined using qPCR. Significance was assessed relative to the negative controls using a one-tailed t-test.
Immunofluorescence
We crosslinked cells in 4% paraformaldehyde for 15 min, and washed in 1× PBS three times. To permeabilize the cells, we washed with 1× PBS +0.1% Triton and then blocked in 1× PBS + 0.1% Triton + 1% BSA for 45 min at room temperature. We incubated cells with anti-Pou5f1 antibody (Santa Cruz: SC-9081) at 1:100 dilution in blocking solution for 1.5 h at room temperature and then washed in blocking solution three times. Next, we incubated cells in anti-rabbit secondary antibody coupled to GFP (Jackson ImmunoResearch: 111-486-152) at a dilution of 1:1,000 in blocking solution for 45 min. Finally, we thoroughly washed cells in blocking solution three times, and added vectashield containing DAPI (VWR: 101098-044) to each well.
Public data set curation
Traditionally, lineage markers are used to identify changes in phenotypic states. Although these markers can be good indicators of differentiation potential, there are two major limitations with this approach. First, there are multiple genes that are associated with each lineage so simply looking at one can often be misleading. Second, this approach only works for classifying states with well-characterized marker genes but would not work for a comprehensive characterization of the function in the cell. Therefore, we decided to take a different approach and look at the entire gene expression profile of each lincRNA knockdown to determine what cell state each lincRNA resembles.
We curated a set of ES perturbations and differentiation states from publicly available sources. Specifically, we used the NCBI e-utils (http://eutils.ncbi.nlm.nih.gov/) to programmatically identify all published data sets containing keywords associated with embryonic stem cells. We filtered the list to only include mouse data sets that were generated across one of three commercial array platforms (Affymetrix, Agilent and Illumina). Following this approach, we manually curated the list to include data sets associated with ES cell perturbations (genetic deletions, RNAi, or chemical perturbations) and differentiation or induced differentiation profiles. This curation yielded 41 GEO data sets corresponding to >150 samples.
Specifically, we defined differentiation lineage states using the following data sets. (1) Neuroectoderm: we downloaded a data set (GSE12982) corresponding to mouse ES cells containing a Sox1–GFP reporter construct. Upon differentiation of Sox1–GFP ES cells into embryoid bodies (EBs), Sox1–GFP-positive cells were collected and their global expression was profiled59. In addition, we downloaded a data set (GSE4082)60 corresponding to direct neuroectoderm differentiation61.
(2) Mesoderm: we downloaded the same data set (GSE12982) as above, where the authors differentiated brachyury–GFP reporter ES cells into EBs and sorted and profiled brachyury–GFP-positive cells59.
(3) Endoderm: we downloaded a data set (GSE11523) corresponding to mouse ES cells which were engineered to overexpress GATA633. GATA6 overexpression has been shown to drive ES cells into a primitive endoderm-like state62.
(4) Ectoderm: we downloaded a data set (GSE4082)60 corresponding to mouse ES cells differentiated into primitive ectoderm-like cells with defined media61.
(5) Trophectoderm: we downloaded a data set (GSE11523)33 corresponding to mouse ES cells which were engineered to deplete Oct435. These cells have been shown to enter a trophectoderm-like state35. To ensure specificity to the trophectoderm state, we also compared the expression effects to trophoblast stem cells33. For all lincRNAs identified, we required a significant enrichment for both induced Oct4 knockout and trophoblast stem cell programs.
In addition, for all lineage states we used a curated discrete gene expression signature of differentiation which was previously functionally tested and shown to correspond specifically to differentiation into the associated states63.
Continuous enrichment analysis and phenotype-projection analysis
To determine relationships between lincRNA knockdowns and functional states, we used a modified Gene Set Enrichment Analysis34 approach that accounts for the continuous nature of the two data sets, similar to previously described extensions34,64,65. For each lincRNA knockdown by functional pair we compute a continuous enrichment score. Specifically, (1) for each lincRNA knockdown we compute a normalized score matrix compared to a panel of negative control hairpins by computing a t-statistic for each gene between the replicate lincRNA knockdown expression values and the control knockdown values. (2) For each experiment, we sort the matrix by the normalized score such that the most differentially expressed upregulated gene is first and the most differentially expressed downregulated gene is last. Using this ordering we sort the functional data set such that the ordering corresponds to the differential rank of the lincRNA knockdown set. (3) We compute a score Si as the running average of values from the first position to position i. We then define the enrichment score E as the maximum of the absolute value of Si for all values of i > 10. We require i > 10 to avoid small fluctuations in the beginning of the ranked list causing fluctuations in the enrichment score. This score is computed for each lincRNA knockdown by functional set. Because we have many lincRNA knockdowns and functional sets, in reality we have a matrix of scores and we will refer to the enrichment score of the ith knockdown and jth functional set as Eij .
To assess the significance of these scores, we compute a permutation-derived FDR and assign a confidence value for each projection. Specifically, to assess the significance of Eij, we permute the lincRNA knockdown samples and control samples and compute the enrichment score for each pair across all permutations. To account for the FDR associated with many lincRNAs and functional sets, we use the values of all permutations directly to assess the FDR level of Eij . Specifically, to assess the FDR for each enrichment value Eij , we accumulate all the permutation values for all lincRNA knockdowns and functional sets and compute the number of values greater than Eij as well as a vector of values greater than Eij corresponding to each permutation. The FDR is computed as the average number of permuted values greater than Eij divided by the observed number greater than Eij . Using this approach, we assign an FDR value to each lincRNA knockdown by functional set and identify significant hits as those with an FDR <0.01.
To highlight the accuracy of this approach, we observed that for publicly available gene perturbations for which we also perturbed the gene we were able to identify a significant association of target genes in ∼75% of cases. Although the remaining few did not pass our conservative significance criteria, they also showed increased enrichments consistent with their common effects. In addition, the projected effects are highly reproducible across distinct experiments originating from many groups and across multiple expression platforms. Highlighting the specificity of this approach, we note that there are many profiles for which no lincRNA had a similar effect.
Analysis of gene-expression overlaps between independent hairpin knockdowns
To determine whether independent hairpins targeting the same lincRNA gene share common gene targets, we computed a continuous enrichment score described above. Briefly, we computed a t-statistic for both hairpins against the negative controls. We then took the second best hairpin and sorted the genes. We scored the best hairpin affected genes based on this ranked order. We assessed the significance of this enrichment by permuting the samples and controls and assigned an FDR of the overlap of the expression effect (as described above).
Discrete gene set analysis
Discrete gene sets were analysed using the Gene Set Enrichment Analysis with a slight modification to the scoring procedure to be more analogous to our continuous scoring procedure (described above). Specifically, we computed the average of the expression changes (defined by the t-statistic) for all genes within the discrete gene set upon knockdown63. Significance was assessed by permuting the control and sample labels and re-computing the average statistic for each permutation. The FDR was assessed off of these values as described above.
Lineage marker gene analysis
We curated lineage marker gene sets from published work and publicly available sources17,32,63. We identified lineage marker genes as significantly upregulated using the differential expression criteria outlined above. We validated the expression of these lineage marker genes for a selected set of lineage marker genes using qPCR (as described above) after a 4-day infection. Specifically, we looked at the expression of Fgf5 (ectoderm), Sox1 (neuroectoderm), Sox17 (endoderm), brachyury (mesoderm) and Cdx2 (trophectoderm). Primer sequences are listed in Supplementary Table 9. Expression estimates were normalized to Gapdh and compared to a panel of 25 negative control hairpins.
Identifying bound lincRNA promoters
We obtained genome-wide transcription factor binding data in mouse ES cells from two sources. The transcription factors Oct4, Sox2, Nanog and Tcf3 were downloaded from the Gene Expression Omnibus (GSE11724) and c-Myc, n-Myc, Zfx, Stat3, Smad1, Klf4 and Esrrb from GEO (GSE11431). For each ChIP-Seq data set, the raw reads were obtained from the SRA (http://www.ncbi.nlm.nih.gov/sra) and processed as follows. (1) The reads were all aligned to the mouse genome assembly (build MM9) using the Bowtie aligner66, requiring a single best placement of each read. All reads with multiple acceptable placements were removed from the analysis. (2) Binding sites were determined from the aligned reads using the MACS67 (http://liulab.dfci.harvard.edu/MACS/) algorithm using the default parameters with –mfold 8 to account for varying read counts in the libraries. (3) lincRNA promoter regions were defined as previously described2,3 using the location of the K4me3 peaks overlapping or within 5 kb of the transcriptional start site determined by RNA-Seq reconstruction. (4) The transcription factor binding locations and lincRNA promoter locations were intersected and the enrichment level of the peak overlapping a lincRNA promoter was assigned transcription factor binding enrichment for each lincRNA. We defined transcription factor binding locations for protein-coding genes in a comparable way. (5) To exclude the possibility that some of this binding might be due to transcription factor binding at distal enhancers, we excluded all binding events that showed evidence of P300—a protein associated with active enhancers68—localization. Altogether, we only identified ∼5% of promoters overlapping with any P300 enrichment signal, a slightly lower percentage than identified for protein-coding gene promoters with detectable P300 signal.
Identifying transcription-factor-regulated lincRNA genes
lincRNA probes on the Agilent microarray were analysed using the differential expression methodology described above after knockdown of the transcription factor and comparison to the negative control hairpins. To confirm the expression changes of these lincRNAs, we hybridized 12 transcription factor knockdowns on a custom lincRNA codeset using the Nanostring nCounter assay41 (LIN-MES1-96). The knockdowns were profiled in biological duplicate along with 15 negative controls. Regulated lincRNAs were identified using the differential expression approach described above.
Nanostring probe-set design
Nanostring probes against lincRNA genes were designed following the standard nanostring design principles with the following modifications specifically for the lincRNA probes. (1) To exclude possible cross-hybridization, probes were screened for cross-hybridization against both the standard mouse transcriptome as well as a background database constructed from all the lincRNA sequences. (2) To account for isoform coverage, a first pass design attempted to select a probe that would target as many isoforms as possible for each lincRNA. In cases where it was not possible to target all isoforms for a given lincRNA, the probe that targeted the largest number was selected, and additional probes were chosen when possible to target the remaining isoforms. (3) The standard restrictions on melting temperature and sequence composition were relaxed to include probes for as many lincRNAs as possible.
Retinoic acid differentiation
V6.5 cells were cultured on gelatin-coated dishes in mES media in the absence of LIF. 5 μM of retinoic acid was added daily and cell samples were taken daily for 6 days. RNA was extracted using Qiagen’s RNeasy spin columns following the manufacturer’s protocol.
Western blots
30 μg of mESC nuclear protein extracts were run on 10% Bis-Tris gels (Invitrogen NP0316BOX) in MOPS buffer (Invitrogen NP0001) at 75 V for 20 min followed by 120 V for 1 h. Gels were incubated for 30 min in 20% methanol transfer buffer (Invitrogen NP0006-1) and transferred onto PVDF membranes (Invitrogen 831605) at 20 V for 1 h using the Bio-Rad semi-dry transfer system (170-3940). Membranes were blocked in Blotto (Pierce, 37530) at room temperature for 1 h. Antibodies were diluted in Blotto and membranes were incubated overnight at 4 °C. Antibodies were diluted in the following concentrations. Ezh2 1:2,000, Suz12 1:5,000, hnRNPH 1:1,000, Ruvbl2 1:1,000, Jarid1b 1:500, Hdac1 1:250, Cbx6 1:500, Yy1 1:500. All antibodies tested were raised in rabbit. The next day, membranes were washed 3× in 0.1% TBST for 5 min each. The membranes were probed with anti-rabbit-horse radish peroxidase (GE Healthcare; NA9340V) at a 1:10,000 dilution, washed 3× in 0.1% TBST, incubated in ECL reagent (GE Healthcare RPN2132) and exposed.
Crosslinked RNA immunoprecipitation
V6.5 mES cells were fixed with 1% formaldehyde for 10 min at room temperature, quenched with 2.5 M glycine, washed with 1× PBS (3×) harvested by scraping, pelleting, and re-suspended in modified RIPA lysis buffer (150 mM NaCl, 50 mM Tris, 0.5% sodium deoxycholate, 0.2% SDS, 1% NP-40) supplemented with RNase inhibitors (Ambion, AM2694) and protease inhibitors. For UV crosslinking experiments, cells were irradiated with 254 nm UV light. Cells were kept on ice and crosslinked in 1× PBS using 400,000 µjoules cm−2.
Cell suspension was sonicated using a Branson 250 Sonifier for 3 × 20 s cycles at 20% amplitude. 10 μl of Turbo DNase (Ambion, AM2238) was added to sonicated material, incubated at 37 °C for 10 min, and spun down at max speed for 10 min at 4 °C. Protein-G beads were washed and pre-incubated with antibodies for 30 min at room temperature. Lysate and beads were incubated at 4 °C for 2 h. Beads were washed 3× using the following wash buffer (1× PBS, 0.1% SDS, 0.5% NP-40) followed by 2× using a high salt wash buffer (5× PBS, 0.1% SDS, 0.5% NP-40) and crosslinks were reversed and proteins were digested with 5 μl proteinase-K (NEB, P8102S) at 65 °C for 2–4 h. RNA was purified using phenol/chloroform/isoamyl alcohol and RNA was precipitated in isopropanol.
Nanostring hybridization
500 ng of total RNA was hybridized for 17 h using the lincRNA code set. The hybridized material was loaded into the nCounter prep station followed by quantification on the nCounter Digital Analyser following the manufacturer’s protocol. For RNA immunoprecipitation experiments, we used a modified protocol. After reverse crosslinking, RNA was extracted using phenol/chloroform and ethanol precipitation methods and re-suspended in 10 μl of H2O. 5 μl of the eluted material was hybridized for 17 h using the lincRNA code set.
Nanostring analysis
Probe values were normalized to negative control probes by dividing the value of the probe by the maximum negative control probe. Probe values were floored to a normalized value of 3 (threefold higher than maximum negative control). Probes with no value greater than this floor across all samples were removed from the analysis. The values were log transformed. To control for variability between runs and different input material amounts, we normalized all samples simultaneously using the quantile normalization approach described above. The result is a set of normalized log-expression values for each probe normalized across all experiments.
Validation of RNA immunoprecipitation methods
To validate our formaldehyde-based RNA immunoprecipitation method we immunoprecipitated the RNA binding protein hnRNPH, which has a role in mRNA splicing69 and identified the associated RNAs. Consistent with known interactions, we identified a strong enrichment for its binding to intronic regions of mRNA genes. We validated these observed results in mouse ES cells by performing UV-crosslinking experiments70,71,72 and identified nearly identical results. We identified a similar correlation between the UV and formaldehyde crosslinked samples as for biological replicates of UV crosslinked samples and formaldehyde crosslinked samples and highly comparable enrichments (data not shown).
Antibody selection
We selected chromatin proteins that have been implicated in regulation of the pluripotent state along with their known associated ‘reader’, ‘writer’ and ‘eraser’ complexes. Specifically, we tested antibodies against 40 chromatin proteins, corresponding to 28 chromatin complexes. In many cases, we tested multiple antibodies against the same target protein to try to identify an antibody that worked well for immunoprecipitation. A full list of tested complexes and their associated antibodies is listed in Supplementary Table 18.
Determining significant chromatin–lincRNA enrichments
We tested each antibody using formaldehyde crosslinked cells and had a two-step procedure for considering an antibody successful. (1) We tested all selected antibodies in batches, with each batch containing a mock-IgG (Santa Cruz) negative control and hnRNPH (Bethyl) positive control. Batches with variability in either the mock-IgG or hnRNPH controls were excluded and retested. For each successful batch, we computed enrichment for each lincRNA between the tested antibody and mock-IgG. We considered an antibody successful in the first step if the highest enrichment level exceeded a fivefold change compared to the mock-IgG control and more than 10 lincRNAs exceeded this threshold. Although this approach can yield false positives (antibodies that pass but are not efficient) it significantly reduced the number of antibodies to be tested in the next step. (2) For all antibodies that successfully passed the first criterion, we performed immunoprecipitation on two additional biological replicates along with 4 mock-IgG controls. We computed a t-statistic for each lincRNA compared to the controls and assessed the significance using a permutation test, by permuting the samples and IgG samples (as above). Hits were considered significant if they exceed a t-statistic cutoff of 2 (log scale) compared to the controls and had an FDR <0.2. We allowed a slightly higher FDR cutoff because the number of permutations was far smaller yielding lower power to estimate the FDR. Only antibodies yielding significant lincRNAs were considered successful. In total, we identified 12 of the 28 complexes (55 antibodies) with at least one successful antibody.
Determining significant overlaps between lincRNA and chromatin protein knockdown effects
To determine the functional overlap between the lincRNA and the chromatin complexes it physically interacts with, we compared the effects on gene expression upon knockdown of the lincRNA and the associated protein complex. To do this, we used the gene expression profiles determined for each lincRNA knockdown and knockdowns of 9 of the 12 identified chromatin complexes for which we had good hairpins. We defined each interaction between a lincRNA and protein, and computed a continuous enrichment score, generated all permutations of the control hairpins and sample hairpins and assigned an FDR to the scores (as described above). At an FDR <0.05 we identified 43% of the interactions to be significant. For 69% of the interactions, we were able to identify an overlap at an FDR <0.1.
References
The FANTOM Consortium The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005)
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009)
Guttman, M. et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nature Biotechnol. 28, 503–510 (2010)
Khalil, A. M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl Acad. Sci. USA 106, 11667–11672 (2009)
Ponjavic, J., Ponting, C. P. & Lunter, G. Functionality or transcriptional noise? Evidence for selection within long noncoding RNAs. Genome Res. 17, 556–565 (2007)
Mattick, J. S. The genetic signatures of noncoding RNAs. PLoS Genet. 5, e1000459 (2009)
Koziol, M. J. & Rinn, J. L. RNA traffic control of chromatin complexes. Curr. Opin. Genet. Dev. 20, 142–148 (2010)
De Santa, F. et al. A large fraction of extragenic RNA pol II transcription sites overlap enhancers. PLoS Biol. 8, e1000384 (2010)
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010)
Ebisuya, M., Yamamoto, T., Nakajima, M. & Nishida, E. Ripples from neighbouring transcription. Nature Cell Biol. 10, 1106–1113 (2008)
Ørom, U. A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010)
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010)
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007)
Smith, A. G. Embryo-derived stem cells: of mice and men. Annu. Rev. Cell Dev. Biol. 17, 435–462 (2001)
Jaenisch, R. & Young, R. Stem cells, the molecular circuitry of pluripotency and nuclear reprogramming. Cell 132, 567–582 (2008)
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008)
Ivanova, N. et al. Dissecting self-renewal in stem cells with RNA interference. Nature 442, 533–538 (2006)
Boyer, L. A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, 349–353 (2006)
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006)
Fazzio, T. G., Huff, J. T. & Panning, B. An RNAi screen of chromatin proteins identifies Tip60-p400 as a regulator of embryonic stem cell identity. Cell 134, 162–174 (2008)
Bilodeau, S., Kagey, M. H., Frampton, G. M., Rahl, P. B. & Young, R. A. SetDB1 contributes to repression of genes encoding developmental regulators and maintenance of ES cell state. Genes Dev. 23, 2484–2489 (2009)
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006)
Hu, G. et al. A genome-wide RNAi screen identifies a new transcriptional module required for self-renewal. Genes Dev. 23, 837–848 (2009)
Plath, K., Mlynarczyk-Evans, S., Nusinow, D. A. & Panning, B. Xist RNA and the mechanism of X chromosome inactivation. Annu. Rev. Genet. 36, 233–278 (2002)
Koerner, M. V., Pauler, F. M., Huang, R. & Barlow, D. P. The function of non-coding RNAs in genomic imprinting. Development 136, 1771–1783 (2009)
Ponjavic, J., Oliver, P. L., Lunter, G. & Ponting, C. P. Genomic and transcriptional co-localization of protein-coding and long non-coding RNA pairs in the developing brain. PLoS Genet. 5, e1000617 (2009)
Sproul, D., Gilbert, N. & Bickmore, W. A. The role of chromatin structure in regulating the expression of clustered genes. Nature Rev. Genet. 6, 775–781 (2005)
Silva, J. et al. Nanog is the gateway to the pluripotent ground state. Cell 138, 722–737 (2009)
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003)
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003)
Brambrink, T. et al. Sequential expression of pluripotency markers during direct reprogramming of mouse somatic cells. Cell Stem Cell 2, 151–159 (2008)
Sherwood, R. I. et al. Prospective isolation and global gene expression analysis of definitive and visceral endoderm. Dev. Biol. 304, 541–555 (2007)
Aiba, K. et al. Defining developmental potency and cell lineage trajectories by expression profiling of differentiating mouse embryonic stem cells. DNA Res. 16, 73–80 (2009)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nature Genet. 24, 372–376 (2000)
Pasini, D., Bracken, A. P., Hansen, J. B., Capillo, M. & Helin, K. The polycomb group protein Suz12 is required for embryonic stem cell differentiation. Mol. Cell. Biol. 27, 3769–3779 (2007)
Jiang, H. et al. Role for Dpy-30 in ES cell-fate specification by regulation of H3K4 methylation within bivalent domains. Cell 144, 513–525 (2011)
Marson, A. et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134, 521–533 (2008)
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nature Genet. 42, 631–634 (2010)
Jiang, J. et al. A core Klf circuitry regulates self-renewal of embryonic stem cells. Nature Cell Biol. 10, 353–360 (2008)
Geiss, G. K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nature Biotechnol. 26, 317–325 (2008)
Dey, B. K. et al. The histone demethylase KDM5b/JARID1b plays a role in cell fate decisions by blocking terminal differentiation. Mol. Cell. Biol. 28, 5312–5327 (2008)
Cloos, P. A., Christensen, J., Agger, K. & Helin, K. Erasing the methyl mark: histone demethylases at the center of cellular differentiation and disease. Genes Dev. 22, 1115–1140 (2008)
Zappulla, D. C. & Cech, T. R. Yeast telomerase RNA: a flexible scaffold for protein subunits. Proc. Natl Acad. Sci. USA 101, 10024–10029 (2004)
Wutz, A., Rasmussen, T. P. & Jaenisch, R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nature Genet. 30, 167–174 (2002)
Tsai, M. C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010)
Meissner, A., Eminli, S. & Jaenisch, R. Derivation and manipulation of murine embryonic stem cells. Methods Mol. Biol. 482, 3–19 (2009)
Nichols, J. et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 95, 379–391 (1998)
Avilion, A. A. et al. Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev. 17, 126–140 (2003)
Niwa, H., Burdon, T., Chambers, I. & Smith, A. Self-renewal of pluripotent embryonic stem cells is mediated via activation of STAT3. Genes Dev. 12, 2048–2060 (1998)
Nakatake, Y. et al. Klf4 cooperates with Oct3/4 and Sox2 to activate the Lefty1 core promoter in embryonic stem cells. Mol. Cell. Biol. 26, 7772–7782 (2006)
Brons, I. G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007)
Torres-Padilla, M. E., Parfitt, D. E., Kouzarides, T. & Zernicka-Goetz, M. Histone arginine methylation regulates pluripotency in the early mouse embryo. Nature 445, 214–218 (2007)
Gaspar-Maia, A. et al. Chd1 regulates open chromatin and pluripotency of embryonic stem cells. Nature 460, 863–868 (2009)
Dejosez, M. et al. Ronin is essential for embryogenesis and the pluripotency of mouse embryonic stem cells. Cell 133, 1162–1174 (2008)
Yuan, P. et al. Eset partners with Oct4 to restrict extraembryonic trophoblast lineage potential in embryonic stem cells. Genes Dev. 23, 2507–2520 (2009)
Pruitt, K. D., Tatusova, T., Klimke, W. & Maglott, D. R. NCBI Reference Sequences: current status, policy and new initiatives. Nucleic Acids Res. 37, D32–D36 (2009)
Yang, Y. H. et al. Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res. 30, e15 (2002)
Shen, X. et al. EZH1 mediates methylation on histone H3 lysine 27 and complements EZH2 in maintaining stem cell identity and executing pluripotency. Mol. Cell 32, 491–502 (2008)
Aiba, K. et al. Defining a developmental path to neural fate by global expression profiling of mouse embryonic stem cells and adult neural stem/progenitor cells. Stem Cells 24, 889–895 (2006)
Ying, Q. L., Stavridis, M., Griffiths, D., Li, M. & Smith, A. Conversion of embryonic stem cells into neuroectodermal precursors in adherent monoculture. Nature Biotechnol. 21, 183–186 (2003)
Morrisey, E. E. et al. GATA6 regulates HNF4 and is required for differentiation of visceral endoderm in the mouse embryo. Genes Dev. 12, 3579–3590 (1998)
Bock, C. et al. Reference maps of human ES and iPS cell variation enable high-throughput characterization of pluripotent cell lines. Cell 144, 439–452 (2011)
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009)
Lamb, J. et al. The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006)
Langmead, B., Hansen, K. D. & Leek, J. T. Cloud-scale RNA-sequencing differential expression analysis with Myrna. Genome Biol. 11, R83 (2010)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Visel, A. et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009)
Katz, Y., Wang, E. T., Airoldi, E. M. & Burge, C. B. Analysis and design of RNA sequencing experiments for identifying isoform regulation. Nature Methods 7, 1009–1015 (2010)
Licatalosi, D. D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008)
Ule, J. et al. CLIP identifies Nova-regulated RNA networks in the brain. Science 302, 1212–1215 (2003)
Wang, Z., Tollervey, J., Briese, M., Turner, D. & Ule, J. CLIP: construction of cDNA libraries for high-throughput sequencing from RNAs cross-linked to proteins in vivo . Methods 48, 287–293 (2009)
Acknowledgements
We thank D. Rivera, T. Green, T. Bhimdi, G. Verstappen, C. Surka, S. Silver, A. Brown, D. Lam and O. Ram for technical help; C. Gifford, S. Markoulaki and R. Jaenisch for providing cell lines used in this study; P. Tsang, B. Curry, A. Tsalenko and Agilent Technologies for microarray and technical help; B. Challis and Active Motif for antibodies; G. Geiss, R. Boykin and Nanostring technologies for technical help; E. Wang and C. Burge for help with RNA immunoprecipitation experiments and helpful discussions; P. Gupta, A. Gnirke, J. Cassady, E. Lieberman-Aiden, M. Cabili and M. Thompson for discussions and ideas; and L. Gaffney for assistance with figures. M. Guttman is a Vertex scholar. This work was funded by NHGRI, a Center for Excellence for Genomic Science, the Merkin Foundation for Stem Cell Research, and funds from the Broad Institute of MIT and Harvard.
Author information
Authors and Affiliations
Contributions
M. Guttman and E.S.L. conceived and designed the overall project with help from A.M., A.R., J.L.R. and D.E.R.; M. Guttman and J.D. designed experiments with help from J.K.G., X.Y. (RNAi), B.W.C. (pluripotency assays) and I.A. (RNA IP); M. Guttman, J.D., G.M., A.B.L., R.A. and G.Y. performed experiments; M. Guttman, J.D. and M. Garber analysed data; L.B., A.M. and D.E.R. provided reagents; and M. Guttman and E.S.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.B.L., R.A. and L.B. are employees of and own stock in Agilent technologies.
Supplementary information
Supplementary Figures
The file contains Supplementary Figures 1-17 with legends. (PDF 5868 kb)
Supplementary Tables
This file contains Supplementary Tables 1–19 as follows: Table 1 ESC lincRNA names, genomic coordinates, sequences, and primers; Table 2 shRNA sequences and on-target knockdown levels; Table 3 Affected genes upon knockdown of lincRNA and protein-coding genes; Table 4 Number of genes affected upon knockdown; Table 5 Overlapping gene expression affects for second hairpins; Table 6 Distance to closest affected protein-coding gene, Table 7 Nanog-Luciferase levels and Alamar levels after infection; Table 8 mRNA expression levels of pluripotency markers after knockdown of lincRNA and protein-coding genes; Table 9 Primer sequences used for pluripotency and differentiation markers; Table 10 Oct4 expression levels across second best hairpins; Table 11 lincRNA knockdown projections; Table 12 Lineage expression profiles; Table 13 Lineage expression expression changes upon transcription factor knockdown, Table 16 incRNA expression changes upon retinoic acid induced differentiation Table17 lincRNA associations with chromatin proteins; Table 18 Tested chromatin complexes and antibodies used and Table19 Overlap in expression between chromatin proteins and lincRNAs. (ZIP 4048 kb)
Rights and permissions
About this article
Cite this article
Guttman, M., Donaghey, J., Carey, B. et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 477, 295–300 (2011). https://doi.org/10.1038/nature10398
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10398
This article is cited by
-
LncRNA Anxa10-203 enhances Mc1r mRNA stability to promote neuropathic pain by recruiting DHX30 in the trigeminal ganglion
The Journal of Headache and Pain (2024)
-
The pluripotent factor OCT4A enhances the self-renewal of human dental pulp stem cells by targeting lncRNA FTX in an LPS-induced inflammatory microenvironment
Stem Cell Research & Therapy (2023)
-
Integrated analysis of mRNAs and lncRNAs reveals candidate marker genes and potential hub lncRNAs associated with growth regulation of the Pacific Oyster, Crassostrea gigas
BMC Genomics (2023)
-
Profiling mitochondria-polyribosome lncRNAs associated with pluripotency
Scientific Data (2023)
-
Estimating residual undifferentiated cells in human chemically induced pluripotent stem cell derived islets using lncRNA as biomarkers
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.