Abstract
Recent advances in DNA synthesis technology have enabled the construction of novel genetic pathways and genomic elements, furthering our understanding of system-level phenomena1,2,3,4,5,6,7. The ability to synthesize large segments of DNA allows the engineering of pathways and genomes according to arbitrary sets of design principles. Here we describe a synthetic yeast genome project, Sc2.0, and the first partially synthetic eukaryotic chromosomes, Saccharomyces cerevisiae chromosome synIXR, and semi-synVIL. We defined three design principles for a synthetic genome as follows: first, it should result in a (near) wild-type phenotype and fitness; second, it should lack destabilizing elements such as tRNA genes or transposons8,9; and third, it should have genetic flexibility to facilitate future studies. The synthetic genome features several systemic modifications complying with the design principles, including an inducible evolution system, SCRaMbLE (synthetic chromosome rearrangement and modification by loxP-mediated evolution). We show the utility of SCRaMbLE as a novel method of combinatorial mutagenesis, capable of generating complex genotypes and a broad variety of phenotypes. When complete, the fully synthetic genome will allow massive restructuring of the yeast genome, and may open the door to a new type of combinatorial genetics based entirely on variations in gene content and copy number.
Similar content being viewed by others
Main
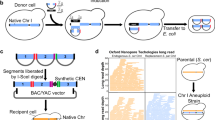
The first phase of any genome engineering project is design (Supplementary Text 1). We designed the right arm of chromosome IX (IXR) according to the three principles outlined above and in Box 1. IXR is the smallest chromosome arm in the genome and features several genomic elements of interest (Fig. 1a), making it suitable for a pilot study. The designed sequence, synIXR, is based on a native IXR sequence extending from open reading frame (ORF) YIL002W through the centromere and the remainder of chromosome IXR, an 89,299-base-pair (bp) sequence (native IXR position 350,585–438,993 (ref. 10)). In accordance with the second design principle, a transfer RNA gene, a Ty1 long terminal repeat (LTR), and telomeric sequences were removed. The final synIXR sequence, 91,010 bp, is slightly longer than the native sequence owing to the inclusion of 43 loxPsym sites, and it replaces 20.3% of the native chromosome. A 30-kilobase (kb) telomeric segment of the left arm of chromosome VI (semi-synVIL) was similarly designed (Fig. 1b and Supplementary Text 2), and replaced 15.7% of the native chromosome. Of the original sequence lengths, 17% was changed by base substitution, deleted, or inserted during design of the two synthetic segments (Supplementary Table 1). Sequences were submitted to GenBank (sequences synIXR:JN020955 and semi-synVIL:JN020956 are also available in Supplementary Information).
Boxed text indicates elements deleted in the synthetic chromosomes. Vertical green bars inside ORFs indicate PCRTag amplicons; only sequences at the outside edges of these are recoded. ARS, autonomously replicating sequence. a, SynIXR. Vector is circular. b, Semi-synVIL.
We systematically introduced two sets of changes in silico using the genome editing suite BioStudio (S.M.R., J.S.D., J.D.B. and J.S.B., unpublished data): TAG/TAA stop-codon swaps and PCRTag sequences (see Supplementary Text 1). In recognition of the third design principle, the elimination of the TAG stop codon by recoding to TAA frees a codon for future expansion of the genetic code (for example, by adding a twenty-first, unnatural amino acid11,12), and could serve as a future mechanism of reproductive isolation and control. PCRTags are short pairs of recoded sequences, unique to either the wild-type or synthetic genome. They serve as convenient, low-cost, closely spaced genetic markers for verifying the introduction of synthetic sequence and the removal of native sequence by allowing the design of PCR primers for rapid evaluation of the presence of synthetic sequences and absence of native sequences. This is critical for evaluating the incorporation of synthetic DNA (see below and Supplementary Text 2). PCRTags, designed in silico, were tested in triplicate to verify specificity (Supplementary Fig. 1 and Supplementary Tables 2 and 3).
LoxPsym sequences are nondirectional loxP sites that are capable of recombining in either orientation13. Theoretically, they produce inversions or deletions with equal probability. Under the third design principle, these sites form the substrate for the inducible SCRaMbLE system and are intended to generate combinatorial diversity. We inserted loxPsym sites 3 bp after the stop codon of each nonessential gene and at major landmarks, such as sites of LTR and tRNA deletions, flanking the centromere CEN9, and adjacent to telomeres (Fig. 1 and Supplementary Text 1). LoxPsym sites inserted at equivalent positions genome-wide will allow the formation of many structurally distinct genomes.
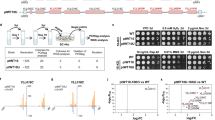
After completion of chromosome design and construction, ‘arm-swap’ strains, wherein the wild-type sequence was replaced with synthetic sequence, were generated. The synIXR chromosome, cloned in a circular bacterial artificial chromosome (BAC) vector, includes all sequences needed for propagation in yeast and bacteria (Fig. 1a). We introduced synIXR into a diploid strain by transformation (Fig. 2a); typically, about 10–15% of the synIXR transformants obtained were positive for all PCRTag pairs tested (Fig. 2d). We chose one such transformant, strain A (Fig. 2a), and truncated one native IXR homologue (IXΔR) by transforming with a suitably designed linear DNA fragment14, introducing a selectable marker (URA3) and a telomere seed sequence, generating strain C (Fig. 2b). Chromosome truncation was confirmed by pulsed-field gel electrophoresis analysis (Fig. 2c), and strain C was sporulated to generate haploids carrying synIXR and IXΔR. We observed more spore lethality than in control crosses, presumably owing to segregation of synIXR away from IXΔR; cells bearing only synIXR or only IXΔR would lack many essential genes and would not survive. PCRTag analysis of 14 synIXR candidate arm-swap strains revealed ten haploids with all synthetic PCRTags and no native PCRTags present (Fig. 2d and Supplementary Fig. 2). The remaining four strains carried BACs with patchworks of synthetic and native sequences indicative of meiotic gene-conversion events (Supplementary Fig. 2). Sanger sequencing and structural analyses (Supplementary Fig. 3, Supplementary Table 4 and Supplementary Text 3) of recovered synIXR BACs revealed that no mutations had occurred in the synthetic chromosome. Thus, the synthetic sequence is replicated faithfully.
a, Generation of synIXR haploids. The synIXR BAC (L) was transformed into the wild-type strain BY4743 (WT, step I) to generate strain A (step II). One copy of native IXR in A was replaced with a URA3–telomere seed cassette (U), generating IXΔR in strain B (step III). B was sporulated to produce haploids (step IV). Circle, centromere; small square, LEU2 gene. b, Structure of IXΔR. c, Electrophoretic karyotype (top panel) and Southern blot of NotI digest (bottom panel) of the wild-type, strain A, strain B and synIXR-1D genomes. Linearized synIXR migrates as a discrete band of ∼100 kb. The probe (YIL002C) detects all isoforms of chromosome IX. *, native IXR; **, IXΔR. d, PCRTag analysis. SYN, synIXR BAC; V, vector amplicon.
Whereas synIXR was incorporated in a circular form, we used an alternate strategy to integrate the semi-synVIL chromosome fragment into native chromosome VI (Supplementary Fig. 4): a linear synthetic fragment marked with LEU2 was transformed into a YFL054C::kanMX strain. Approximately 13% of transformants (75 of 586) had the Leu+G418S phenotype expected for the desired integrant. PCRTag analysis showed that 10 of 12 such strains contained only synthetic PCRTags, as expected for full replacement (Supplementary Fig. 5).
The first design principle prioritizes a wild-type phenotype and a high level of fitness despite the incorporated modifications. SynIXR has a designed sequence alteration approximately every 500 bp, 2.64% of total sequence is altered, and it carries 43 loxPsym sites. To check for negative effects of modifications on fitness, we examined colony size and morphology under various conditions, and also performed transcript profiling. We inspected colony size and morphology of synIXR swap strains under six distinct growth conditions. It was impossible to distinguish swap strains from the wild type (BY4741) under these conditions, indicating that any fitness defect attributable to synIXR is modest; fitness tests on semi-synVIL gave similar results (Supplementary Fig. 6).
Synonymous substitutions, introduction of loxPsym sites or other changes might change gene expression. We performed transcript profiling on the swap strains synIXR-1D, synIXR-6B, and synIXR-22D (Supplementary Text 4); these studies revealed notable but predictable trends (Fig. 3). As expected, genes present in two copies (YIL001W and YIL002C, present on both synIXR and IXΔR) were approximately doubled in transcript abundance. Most genes showed no substantial expression change, although a few showed modest decreases; however, the subtelomeric genes YIR039C and YIR042C showed increased expression. We speculate that in the circular synthetic chromosome, these are released from telomeric silencing, resulting in their overexpression. Overall, synIXR genes show relatively normal expression, indicating that loxPsym sites and PCRTags affect expression only minimally. Similarly, no substantial changes were observed by RNA blotting (Supplementary Fig. 7a). To detect possible compensatory transcriptome changes, we profiled transcripts genome-wide. Except for trivial differences attributable to slightly different configurations of selectable markers in the strains, there were no consistent, statistically significant differences outside IXR itself (Supplementary Fig. 7b). Thus, modifications present in synIXR and semi-synVIL do not produce major fitness effects or compensatory transcriptomic alterations.
Transcript profiling of synIXR-1D, -6B, and -22D. The log2 ratio of RNA abundance relative to wild type (BY4741 or BY4742) is shown. YIL002C and YIL001W (blue) exist in two copies. Essential genes are labelled in red. Error bars, s.d.
A central feature of the synthetic yeast genome is the incorporated conditional genome instability system, SCRaMbLE. The design principles dictate that SCRaMbLE should be available for use on demand, yet should lie dormant until intentional Cre recombinase induction, at which point generation of genetic diversity is desirable. To complete the SCRaMbLE toolkit, we incorporated an engineered Cre recombinase fused to the murine oestrogen binding domain (EBD). This recently described Cre-EBD variant15 is oestradiol-inducible, has low basal activity and is controlled by the daughter-cell-specific promoter SCW11 (Supplementary Fig. 8). The plasmid pSCW11-Cre-EBD should produce a pulse of recombinase activity once and only once in each cell’s lifetime, and should depend on oestradiol exposure. The uninduced, integrated construct is well tolerated even in swap strains, which, with 43 loxPsym sites, are expected to be Cre-hypersensitive. Upon oestradiol addition, rearrangements were induced at the loxPsym sites and viability dropped by 100-fold in synIXR strains (Fig. 4a and Supplementary Fig. 9). This loss of viability probably results from loss of synIXR essential genes. In contrast, viability in semi-synVIL, which lacks essential genes, is not affected by Cre induction (Fig. 1b and Supplementary Fig. 9d).
a, Cre induction reduces the fitness of the synIXR strain (SYN) but not the wild type (WT; BY4741). EST, oestradiol; time, oestradiol exposure time. b, PCR analysis of semi-synVIL SCRaMbLE. The map shows primer positions. Amplicon 13 is spurious (wrong size). SCR, SCRaMbLE. c, Shifted colony-size distribution in SCRaMbLE survivors (wild type and the swap strain synIXR-1D). d, PCRTag analysis of Met− (red), Lys− (blue) and Met− Lys− (green) auxotrophs using PCRTags. PCRTag pairs are numbered for each column (see Supplementary Table 2); MET28, pair 25; LYS1, pair 45. Each row represents one clone. Shaded boxes indicate presumed deletions. Panels a–c show strains with integrated Cre-EBD; d shows episomal Cre-EBD.
Semi-synVIL contains just five loxPsym sites, including one immediately adjacent to the telomeric TG1–3 repeats (Fig. 1b). This simple configuration allows comprehensive PCR-based mapping of rearrangements of four of the loxPsym sites in SCRaMbLEd strains. A SCRaMbLEd semi-synVIL population was analysed by PCR for most of the possible rearranged configurations, revealing a large variety of deletions and inversions (Fig. 4b); most predicted rearrangements were readily detected.
The symmetry of loxPsym sites allows alignment in two orientations, theoretically giving rise to deletions and inversions with equal frequency. SynIXR contains 43 loxPsym sites, allowing more than 3,600 potential pairwise interactions between synIXR loxPsym sites. We reasoned that SCRaMbLEd synIXR clones should display high phenotypic diversity. Indeed, SCRaMbLEd swap strains show more growth-rate heterogeneity than wild-type controls (Fig. 4c and Supplementary Fig. 10). These SCRaMbLEd clones show many different phenotypes (Supplementary Fig. 11 and Supplementary Text 5). In summary, SCRaMbLE is sufficient to generate substantial genetic heterogeneity and complex phenotypes.
To characterize the utility of SCRaMbLE further, we performed a mutagenesis study. SynIXR encodes both MET28 and LYS1, genes required for biosynthesis of amino acids16,17. Null mutants result in auxotrophy, and can be detected easily by replica-plating. We introduced episomal Cre-EBD (pSCW11-Cre-EBD-URA3MX cloned in a CEN plasmid) into strain C that was previously made LYS2+ (strain D, yJS587), and performed SCRaMbLE. We screened 20,242 colonies and 3% (604 of 20,242) were candidate lys1 and/or met28 auxotrophs. Of 360 candidates tested more rigorously, 295 (81.9%) were confirmed: we found 212 Lys− auxotrophs (1.37%), 66 Met– auxotrophs (0.43%) and, notably, 17 Lys– Met– double auxotrophs (0.11%). PCRTag profiles of 24 Met– auxotrophs, 35 Lys– auxotrophs and seven double auxotrophs (Fig. 4d) showed that all Met– auxotrophs had deletions in the loxPsym-flanked segment containing MET28 and YAP5, whereas all Lys– auxotrophs had deletions in the loxPsym-flanked segment containing LYS1. The deletion profiles of many SCRaMbLEd auxotrophs were highly variable and more than one segment was often missing.
To confirm that the observed SCRaMbLE phenotypes resulted solely from deletions in synIXR, we recovered the synIXR chromosomes from two Met– auxotrophs into Escherichia coli, and then introduced them to a clean genetic background. In both cases, the auxotrophic phenotype was associated with the presence of the SCRaMbLEd chromosomes (Supplementary Fig. 12 and Supplementary Text 6). Thus, the SCRaMbLE system is a highly effective method of mutagenesis, giving rise to mutants with different genetic backgrounds and generating a wide variety of double mutants.
We have shown there does not seem to be any major theoretical impediment to extending the design strategy outlined here to the entire yeast genome, apart from the challenge of 12-megabase DNA synthesis. Whether or not fitness defects will accumulate as design and synthesis are scaled up remains to be seen; however, the overall high fitness of the swap strains described here validates the design strategy. Furthermore, the iterative, bottom-up approach will allow identification of potential ‘problem regions’ in synthetic sequences as synthesis moves forward. If a given swap experiment results in only transformants with reduced fitness (or if no transformants are obtainable), then the underlying defect can be mapped by introducing sub-segments, facilitated by strategic placement of unique restriction sites throughout synthetic chromosome arms. Also, because a subset of transformants consist of patchworks of native and synthetic sequence (Supplementary Figs 2 and 5), analysis of such strains can be used to map phenotypic defects rapidly. The stability and sequence fidelity of large circular chromosomes seen here and elsewhere5,6,7 bode well for the use of yeast as a host platform for synthetic biology.
SCRaMbLE may become a useful general strategy for analysing genome structure, content and function. One important feature of SCRaMbLE is its potential for customization: expression of different Cre-EBD variants from various promoters at distinct levels of inducer (oestradiol) should produce distinct SCRaMbLE dynamics. Use of weaker promoters than pSCW11, use of promoters expressed at different phases of the cell cycle, performing SCRaMbLE in diploids, and lowering the inducer concentration should all contribute to decreased lethality of SCRaMbLE strains, an important consideration as additional segments of the genome are replaced with synthetic counterparts and the proportion of essential genes that can be lost by SCRaMbLEing increases. As shown here, SCRaMbLE mutagenesis is efficient and generates mutants with a wide variety of different genetic backgrounds. It is possible that different combinations of gene deletions will give rise to a variety of subtly different phenotypes that can be mapped rapidly by PCRTag analysis; more extensive analysis by deep sequencing will reveal changes in genome structure and content. As the synthetic yeast genome grows, opportunities for genome rearrangement will increase exponentially. In principle, changes in chromosome number, ploidy, content and structure are all possible, increasing the utility of the SCRaMbLE system. For example, there may be many different routes to a minimal genome, and exploring all of them by a hit or miss predictive approach is impractical and unlikely to yield comprehensive results. Using SCRaMbLE, many independent routes of genome minimization can be explored at one time, under many environmental conditions, for instance by growing yeast cells long-term in serially transferred batch cultures, or in a chemostat or turbidistat under conditions in which Cre is minimally active. Such an approach may also lead to derivatives that are more fit than the parent, for example, by gene duplication events facilitated by the Cre-EBD/loxPsym system.
Methods Summary
DNA preparation
BAC DNA was prepared using the Qiagen plasmid midi kit or alkaline lysis18. The following protocol modifications were made: cells were diluted 1:100 from an overnight culture into 50 ml, grown in Luria broth with 50 μg ml−1 carbenicillin, and grown at 30 °C for 14–16 h. Qiagen-purified DNA was treated with 60 μg ml−1 proteinase K at 37 °C overnight, then extracted with phenol/chloroform. DNAs prepared without a column were phenol/chloroform extracted, and then treated with RNase immediately before use.
Yeast genomic DNA for use in PCRTag analysis was prepared by standard methods19. DNA preparation for recovery of the synIXR BAC into bacteria was as previously reported20.
PCR conditions
PCRTags were amplified using Taq polymerase (New England Biolabs). Template concentrations were 1 ng μl−1 for genomic DNA and 10 pg μl−1 for purified BAC DNA. The following program was used: 94 °C 3 min; 30 cycles of 94 °C 30 s, 65 °C 30 s, 72 °C 30 s; 72 °C 3 min.
RNA analysis
Total RNA was isolated by hot acid phenol extraction. Microarray hybridization and data analysis were performed at the Johns Hopkins Microarray Core Facility (http://www.microarray.jhmi.edu). Dubious ORFs and pseudogenes were omitted from synIXR transcript analysis.
Pulsed-field gels
DNAs were prepared as described elsewhere21. The identity of the chromosomes was inferred from the known molecular karyotype of wild type (BY4743), and from lambda ladders run on the same gel.
Online Methods
DNA preparation
BAC DNA was prepared using the Qiagen plasmid midi kit or alkaline lysis18. The following protocol modifications were made: cells were diluted 1:100 from an overnight culture into 50 ml, grown in Luria broth with 50 μg ml−1 carbenicillin, and grown at 30 °C for 14–16 h. Qiagen-purified DNA was treated with 60 μg ml−1 proteinase K at 37 °C overnight, then extracted with phenol/chloroform. DNAs prepared without a column were phenol/chloroform extracted, and then treated with RNase immediately before use.
Yeast genomic DNA for use in PCRTag analysis was prepared by standard methods19. DNA preparation for recovery of the synIXR BAC into bacteria was as previously reported20.
PCR conditions
PCRTags were amplified using Taq polymerase (New England Biolabs). Template concentrations were 1 ng μl−1 for genomic DNA and 10 pg μl−1 for purified BAC DNA. The following program was used: 94 °C 3 min; 30 cycles of 94 °C 30 s, 65 °C 30 s, 72 °C 30 s; 72 °C 3 min.
RNA analysis
Total RNA was isolated by hot acid phenol extraction. Microarray hybridization and data analysis were performed at the Johns Hopkins Microarray Core Facility (http://www.microarray.jhmi.edu). Dubious ORFs and pseudogenes were omitted from synIXR transcript analysis.
Pulsed-field gels
DNAs were prepared as described elsewhere21. The identity of the chromosomes was inferred from the known molecular karyotype of wild type (BY4743), and from lambda ladders run on the same gel.
Yeast strains, transformation and tetrad analysis
Strains ABY7 and ABY8 were derived from strain BY4743; ABY7 (MAT a) and ABY7 (MATα) otherwise share the genotype his3Δ1 leu2Δ0 ura3Δ0 lys2Δ0 met15Δ0 yil001::URA3 yir039::kanMX. All strain genotypes are listed in Supplementary Table 8.
BY4743 spheroplasts were transformed with synIXR. The strain YFL054C::kanMX was transformed with synVIL restriction fragments by standard lithium acetate transformation.
The synIXR-1D strain and others were backcrossed to strains ABY7 and ABY8; the resultant diploids were sporulated and genotyped to identify synIXR segregants.
Phenotypic screening
Single colonies were picked into 96-well plates and grown for 48 h in yeast peptone dextrose (YPD) at 30 °C. (SCRaMbLE strains were grown for 72 h in YPD at 30 °C, diluted 1:10 and grown for 4 h before plating.) Tenfold dilutions were spotted on various types of agar medium and selective conditions in OmniTrays (NUNC), as previously described27. Most cells were grown for 72 h (except those grown on yeast extract/peptone/glycerol/ethanol (YPGE) plates, which were grown for 108 h), then scored for growth and photographed.
Yeast growth and media
Unless otherwise indicated, all experiments were performed at 30 °C. YPGE was supplemented with 2% ethanol and 2% glycerol. Concentrations of drugs were as follows: hydroxyurea, 0.2 M; methylmethane sulphonate, 0.05%; 6-azauracil, 100 μg ml−1; benomyl, 15 μg ml−1; hydrogen peroxide, 1 mM; cycloheximide, 10 μg ml−1. Resistance to cycloheximide and hydrogen peroxide was assayed by growing cells in treated medium for 2 h, then plating on YPD. Other phenotypes were assayed by growing cells to mid-log phase in rich media, then spotting tenfold dilutions on selective media.
Colony size measurements
Cells were plated at various dilutions so that similar numbers of colonies were observed on control and experimental (oestradiol-treated) plates. Colony size was measured using ImageJ software28, and normalized against the total number of colonies on each plate. Sample sizes for data presented in Fig. 4c are as follows: wild-type, n = 488 colonies; wild-type + Cre + oestradiol, n = 486; 1D, n = 395; 1D + Cre, n = 251; 1D + oestradiol, n = 416; 1D + Cre + oestradiol, n = 394.
SynIXR BAC sequence analysis
The original synIXR BAC was sequenced by the manufacturer, Codon Devices29. SynIXR BACs were recovered into bacteria and sequenced by Agencourt (Beckman Coulter Genomics), using sequencing primers listed in Supplementary Table 5. Repetitive sequences, including the highly internally repetitive MUC1 open reading frame, were PCR-amplified before sequencing when necessary.
Pulsed-field gels
Samples were run on a 1.0% agarose gel in ×0.5 TBE (pH 8.0) for 20 h at 14 °C on a clamped homogenous electric field (CHEF) gel apparatus. The voltage was 3.5 V cm−1, at an angle of 120° and a switch time of 60–120 s, ramped over 20 h.
NotI (Promega) digests were performed on whole chromosomes embedded in agarose plugs. Agarose plugs were removed from the 0.5 M EDTA storage buffer, washed with 0.05 M EDTA for 1 h at room temperature (∼23°C), and then washed with ×0.1 restriction enzyme buffer, followed by ×1 buffer, under the same conditions.
Probe preparation for northern and Southern blots
Probes were prepared using the Prime-It II kit (Stratagene) and hybridized using Ultrahyb hybridization solution (Ambion) according to the manufacturer’s instructions.
SCRaMbLE
Cre activity was induced by exposure to 1 μM β-oestradiol (Sigma-Aldrich) in rich media for either 48 h (integrated Cre) or 4 h (episomal Cre), except where indicated otherwise. PCRTag analysis of Met− and Lys− auxotrophs was performed with a non-redundant array, using one primer pair per loxPsym-flanked segment.
Accession codes
References
Han, J. S. & Boeke, J. D. A highly active synthetic mammalian retrotransposon. Nature 429, 314–318 (2004)
Richardson, S. M., Wheelan, S. J., Yarrington, R. M. & Boeke, J. D. GeneDesign: rapid, automated design of multikilobase synthetic genes. Genome Res. 16, 550–556 (2006)
Chan, L. Y., Kosuri, S. & Endy, D. Refactoring bacteriophage T7. Mol. Syst. Biol. 1, 2005.0018 (2005)
Stricker, J. et al. A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008)
Gibson, D. G. et al. One-step assembly in yeast of 25 overlapping DNA fragments to form a complete synthetic Mycoplasma genitalium genome. Proc. Natl Acad. Sci. USA 105, 20404–20409 (2008)
Gibson, D. G. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010)
Lartigue, C. et al. Creating bacterial strains from genomes that have been cloned and engineered in yeast. Science 325, 1693–1696 (2009)
Ji, H. et al. Hotspots for unselected Ty1 transposition events on yeast chromosome III are near tRNA genes and LTR sequences. Cell 73, 1007–1018 (1993)
Admire, A. et al. Cycles of chromosome instability are associated with a fragile site and are increased by defects in DNA replication and checkpoint controls in yeast. Genes Dev. 20, 159–173 (2006)
Churcher, C. et al. The nucleotide sequence of Saccharomyces cerevisiae chromosome IX. Nature 387, 84–87 (1997)
Park, H. et al. Expanding the genetic code of Escherichia coli with phosphoserine. Science 333, 1151–1154 (2011)
Isaacs, F. J. et al. Precise manipulation of chromosomes in vivo enables genome-wide codon replacement. Science 333, 348–353 (2011)
Hoess, R. H., Wierzbicki, A. & Abremski, K. The role of the loxP spacer region in P1 site-specific recombination. Nucleic Acids Res. 14, 2287–2300 (1986)
Vollrath, D., Davis, R. W., Connelly, C. & Hieter, P. Physical mapping of large DNA by chromosome fragmentation. Proc. Natl Acad. Sci. USA 85, 6027–6031 (1988)
Lindstrom, D. L. & Gottschling, D. E. The mother enrichment program: a genetic system for facile replicative life span analysis in Saccharomyces cerevisiae. Genetics 183, 413–422 (2009)
Kuras, L., Cherest, H., Surdin-Kerjan, Y. & Thomas, D. A heteromeric complex containing the centromere binding factor 1 and two basic leucine zipper factors, Met4 and Met28, mediates the transcription activation of yeast sulfur metabolism. EMBO J. 15, 2519–2529 (1996)
Ogawa, H. & Fujioka, M. Purification and characterization of saccharopine dehydrogenase from baker’s yeast. J. Biol. Chem. 253, 3666–3670 (1978)
Sambrook, J. & Russell, D. W. Isolation of BAC DNA from small-scale cultures. Cold Spring Harb. Protoc. 10.1101/pdb.prot4006 (2006)
Hoffman, C. S. Preparation of yeast DNA. Curr. Protoc. Mol. Biol. Ch. 13, Unit 13.11. (2001)
Boeke, J. D., Garfinkel, D. J., Styles, C. A. & Fink, G. R. Ty elements transpose through an RNA intermediate. Cell 40, 491–500 (1985)
Schwartz, D. C. & Cantor, C. R. Separation of yeast chromosome-sized DNAs by pulsed field gradient gel electrophoresis. Cell 37, 67–75 (1984)
Lemoine, F. J., Degtyareva, N. P., Lobachev, K. & Petes, T. D. Chromosomal translocations in yeast induced by low levels of DNA polymerase: a model for chromosome fragile sites. Cell 120, 587–598 (2005)
Louis, E. J. The chromosome ends of Saccharomyces cerevisiae. Yeast 11, 1553–1573 (1995)
Parenteau, J. et al. Deletion of many yeast introns reveals a minority of genes that require splicing for function. Mol. Biol. Cell 19, 1932–1941 (2008)
Percudani, R., Pavesi, A. & Ottonello, S. Transfer RNA gene redundancy and translational selection in Saccharomyces cerevisiae. J. Mol. Biol. 268, 322–330 (1997)
Dymond, J. S. et al. Teaching synthetic biology, bioinformatics and engineering to undergraduates: the interdisciplinary build-a-genome course. Genetics 181, 13–21 (2009)
Hampsey, M. A review of phenotypes in Saccharomyces cerevisiae. Yeast 13, 1099–1133 (1997)
Abramoff, M. D., Magelhaes, P. J. & Ram, S. J. Image processing with ImageJ. Biophotonics Int. 11, 36–42 (2004)
Blake, W. J. et al. Pairwise selection assembly for sequence-independent construction of long-length DNA. Nucleic Acids Res. 38, 2594–2602 (2010)
Acknowledgements
We thank G. Church for suggesting the global substitution of TAG codons with TAA codons, C. Connelly for sharing technical expertise and V. Huang for generating a sequence visualizer. We are grateful to B. Cormack, G. Seydoux and J. Nathans for offering helpful advice, to Y. Cai and J. Peccoud for suggesting methods to validate the sequence data, and to E. Louis for providing expert advice on telomeres. The work was supported by National Science Foundation grant MCB0718846 to J.D.B., J.S.B. and S.C.; by a grant from Microsoft to J.S.B. and J.D.B.; by Department of Energy Fellowship DE-FG02097ER25308 to S.M.R.; by National Institutes of Health grant AG023779 to D.E.G.; and by a fellowship from Fondation pour la Recherche Médicale to H.M.
Author information
Authors and Affiliations
Contributions
J.S.D., S.M.R., S.C., J.S.B. and J.D.B. designed experiments. J.S.D., S.M.R., C.E.C., T.B., H.M., N.A., J.W.S., J.D. and A.C.B. performed experiments. W.J.B. built the synIXR chromosome. D.L.L. and D.E.G. generated the integrated CRE-EBD cassette. J.S.D., S.M.R., J.S.B. and J.D.B. analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text 1-8, Supplementary References and Supplementary Figures 1-13 with legends. (PDF 1729 kb)
Supplementary Tables
This file contains Supplementary Tables 1-8. This file was corrected on 22 September 2011 due to an error in one of the tables. (PDF 908 kb)
Rights and permissions
About this article
Cite this article
Dymond, J., Richardson, S., Coombes, C. et al. Synthetic chromosome arms function in yeast and generate phenotypic diversity by design. Nature 477, 471–476 (2011). https://doi.org/10.1038/nature10403
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10403
This article is cited by
-
A designer synthetic chromosome fragment functions in moss
Nature Plants (2024)
-
A robust yeast biocontainment system with two-layered regulation switch dependent on unnatural amino acid
Nature Communications (2023)
-
Trimming the genomic fat: minimising and re-functionalising genomes using synthetic biology
Nature Communications (2023)
-
Engineering the gut microbiome
Nature Reviews Bioengineering (2023)
-
A framework to efficiently describe and share reproducible DNA materials and construction protocols
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.