Abstract
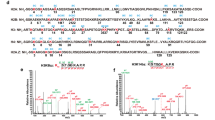
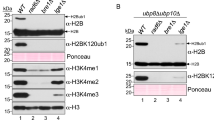
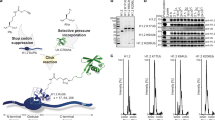
Chromatin reorganization is governed by multiple post-translational modifications of chromosomal proteins and DNA1,2. These histone modifications are reversible, dynamic events that can regulate DNA-driven cellular processes3,4. However, the molecular mechanisms that coordinate histone modification patterns remain largely unknown. In metazoans, reversible protein modification by O-linked N-acetylglucosamine (GlcNAc) is catalysed by two enzymes, O-GlcNAc transferase (OGT) and O-GlcNAcase (OGA)5,6. However, the significance of GlcNAcylation in chromatin reorganization remains elusive. Here we report that histone H2B is GlcNAcylated at residue S112 by OGT in vitro and in living cells. Histone GlcNAcylation fluctuated in response to extracellular glucose through the hexosamine biosynthesis pathway (HBP)5,6. H2B S112 GlcNAcylation promotes K120 monoubiquitination, in which the GlcNAc moiety can serve as an anchor for a histone H2B ubiquitin ligase. H2B S112 GlcNAc was localized to euchromatic areas on fly polytene chromosomes. In a genome-wide analysis, H2B S112 GlcNAcylation sites were observed widely distributed over chromosomes including transcribed gene loci, with some sites co-localizing with H2B K120 monoubiquitination. These findings suggest that H2B S112 GlcNAcylation is a histone modification that facilitates H2BK120 monoubiquitination, presumably for transcriptional activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000)
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007)
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007)
Berger, S. L. The complex language of chromatin regulation during transcription. Nature 447, 407–412 (2007)
Hart, G. W., Housley, M. P. & Slawson, C. Cycling of O-linked β-N-acetylglucosamine on nucleocytoplasmic proteins. Nature 446, 1017–1022 (2007)
Love, D. C. & Hanover, J. A. The hexosamine signaling pathway: deciphering the ‘O-GlcNAc code’. Sci. STKE 2005, re13 (2005)
Yang, X., Zhang, F. & Kudlow, J. E. Recruitment of O-GlcNAc transferase to promoters by corepressor mSin3A: coupling protein O-GlcNAcylation to transcriptional repression. Cell 110, 69–80 (2002)
Gambetta, M. C., Oktaba, K. & Muller, J. Essential role of the glycosyltransferase sxc/Ogt in polycomb repression. Science 325, 93–96 (2009)
Fujiki, R. et al. GlcNAcylation of a histone methyltransferase in retinoic-acid-induced granulopoiesis. Nature 459, 455–459 (2009)
Wang, Z. et al. Extensive crosstalk between O-GlcNAcylation and phosphorylation regulates cytokinesis. Sci. Signal. 3, ra2 (2010)
Sakabe, K., Wang, Z. & Hart, G. W. β-N-acetylglucosamine (O-GlcNAc) is part of the histone code. Proc. Natl Acad. Sci. USA 107, 19915–19920 (2010)
Luger, K. et al. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997)
Das, C., Lucia, M. S., Hansen, K. C. & Tyler, J. K. CBP/p300-mediated acetylation of histone H3 on lysine 56. Nature 459, 113–117 (2009)
Dang, W. et al. Histone H4 lysine 16 acetylation regulates cellular lifespan. Nature 459, 802–807 (2009)
Dong, L. & Xu, C. W. Carbohydrates induce mono-ubiquitination of H2B in yeast. J. Biol. Chem. 279, 1577–1580 (2004)
Yoshida, Y. et al. E3 ubiquitin ligase that recognizes sugar chains. Nature 418, 438–442 (2002)
Kim, J. et al. RAD6-Mediated transcription-coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells. Cell 137, 459–471 (2009)
Dentin, R. et al. Hepatic glucose sensing via the CREB coactivator CRTC2. Science 319, 1402–1405 (2008)
Chikanishi, T. et al. Glucose-induced expression of MIP-1 genes requires O-GlcNAc transferase in monocytes. Biochem. Biophys. Res. Commun. 394, 865–870 (2010)
Jackson, S. P. & Tjian, R. O-glycosylation of eukaryotic transcription factors: implications for mechanisms of transcriptional regulation. Cell 55, 125–133 (1988)
Sinclair, D. A. et al. Drosophila O-GlcNAc transferase (OGT) is encoded by the Polycomb group (PcG) gene, super sex combs (sxc). Proc. Natl Acad. Sci. USA 106, 13427–13432 (2009)
Sawatsubashi, S. et al. A histone chaperone, DEK, transcriptionally coactivates a nuclear receptor. Genes Dev. 24, 159–170 (2009)
Fujiki, R. et al. Ligand-induced transrepression by VDR through association of WSTF with acetylated histones. EMBO J. 24, 3881–3894 (2005)
He, H. H. et al. Nucleosome dynamics define transcriptional enhancers. Nature Genet. 42, 343–347 (2010)
Minsky, N. et al. Monoubiquitinated H2B is associated with the transcribed region of highly expressed genes in human cells. Nature Cell Biol. 10, 483–488 (2008)
Acknowledgements
We thank A. Miyajima, S. Saito and N. Moriyama for experimental support, and M. Yamaki for manuscript preparation. We also thank Y. Maekawa, J. Seta and N. Iwasaki for support with MS. This work was supported in part by The Naito Foundation, the Astellas foundation (to R.F.), the Ministry of Education, Culture, Sports, Science and Technology (MEXT) and the Japan Society for the Promotion of Science (to R.F. and S.K.).
Author information
Authors and Affiliations
Contributions
S.K. planned the study with H.K.; R.G.R. and M.B. provided support and general guidance; R.F. designed the study and performed the experiments with H.S. (α-O-GlcNAc purification), A.Y. (LC–MS/MS), W.H. (O-GlcNAc site mapping), T.C. (in vitro OGT assay), S.I. (Drosophila analysis), Y.I., H.H.H. (ChIP-seq), F.O., J.K. (in vitro monoubiquitination assay), K.I. and J.K (microarray).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Text, Supplementary Figures 1-25 with legends and legends for Supplementary Table 1-3. (PDF 11414 kb)
Supplementary Table 1
This file shows a list of identified GlcNAcylated proteins in chromatin (Please see Supplementary Information file for full legend.) (XLS 168 kb)
Supplementary Table 2
This file shows microarray analysis of HeLa cells cultured under specific experimental conditions (Please see Supplementary Information file for full legend.) (XLS 6550 kb)
Supplementary Table 3
This file shows a list of genes harboring H2B S112 GlcNAc in the promoter or 50 kbp within the gene body, and the expression levels (Please see Supplementary Information file for full legend.) (XLS 1314 kb)
Rights and permissions
About this article
Cite this article
Fujiki, R., Hashiba, W., Sekine, H. et al. GlcNAcylation of histone H2B facilitates its monoubiquitination. Nature 480, 557–560 (2011). https://doi.org/10.1038/nature10656
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10656
This article is cited by
-
O-GlcNAcylation: the sweet side of epigenetics
Epigenetics & Chromatin (2023)
-
Extracellular vesicles are dynamic regulators of maternal glucose homeostasis during pregnancy
Scientific Reports (2023)
-
H4S47 O-GlcNAcylation regulates the activation of mammalian replication origins
Nature Structural & Molecular Biology (2023)
-
Investigation of in vitro histone H3 glycosylation using H3 tail peptides
Scientific Reports (2022)
-
O-GlcNAcylation and stablization of SIRT7 promote pancreatic cancer progression by blocking the SIRT7-REGγ interaction
Cell Death & Differentiation (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.