Abstract
Deregulated expression of the MYC oncoprotein contributes to the genesis of many human tumours, yet strategies to exploit this for a rational tumour therapy are scarce. MYC promotes cell growth and proliferation, and alters cellular metabolism to enhance the provision of precursors for phospholipids and cellular macromolecules1,2. Here we show in human and murine cell lines that oncogenic levels of MYC establish a dependence on AMPK-related kinase 5 (ARK5; also known as NUAK1) for maintaining metabolic homeostasis and for cell survival. ARK5 is an upstream regulator of AMPK and limits protein synthesis via inhibition of the mammalian target of rapamycin 1 (mTORC1) signalling pathway. ARK5 also maintains expression of mitochondrial respiratory chain complexes and respiratory capacity, which is required for efficient glutamine metabolism. Inhibition of ARK5 leads to a collapse of cellular ATP levels in cells expressing deregulated MYC, inducing multiple pro-apoptotic responses as a secondary consequence. Depletion of ARK5 prolongs survival in MYC-driven mouse models of hepatocellular carcinoma, demonstrating that targeting cellular energy homeostasis is a valid therapeutic strategy to eliminate tumour cells that express deregulated MYC.
Similar content being viewed by others
Main
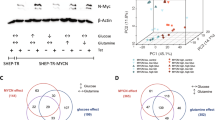
To identify kinases that are specifically required for the viability of cells expressing deregulated MYC, we used U2OS cells expressing c-MYC fused to the oestrogen receptor ligand binding domain (MYC–ER) (Fig. 1a). Activation of MYC–ER by 4-hydroxytamoxifen (OHT) had little effect on apoptosis when cells were grown at low density in the presence of growth factors. Under these conditions, we performed a short interfering (si)RNA screen of the human kinome, using automated microscopy to identify siRNAs that induced poly-ADP-ribose-polymerase cleavage specifically in the presence of OHT. This screen yielded two hits, ARK5 and AMPK (Supplementary Table 1).
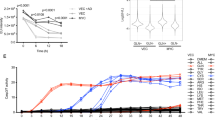
a, Depletion of ARK5 induces MYC-dependent death. U2OS MYC–ER cells were transfected with control siRNA or siARK5 and treated with OHT or solvent. The graph shows the percentage of cells staining positive (pos.) for annexin V only (black) or for both annexin V and propidium iodide (grey). Results were consistent in three independent experiments. Immunoblots document expression of ARK5, MYC–ER and endogenous MYC. Results are shown as mean plus standard deviation (s.d.) of biological triplicates from one representative experiment (here: for cumulative death, annexin V positive plus annexin V and propidium iodide (PI) double positive), except where expressly stated. *P values < 0.01; NS, not statistically significant. b, shRNA depletion of ARK5 induces MYC-dependent death. Top, percentage of U2OS MYC–ER cells with subG1 DNA content after expression of control shRNA (sh con.) or shARK5. Results are averaged (± s.d.) from three independent experiments. Bottom, immunoblots documenting expression of ARK5, MYC–ER and β-actin. c, ARK5 depletion kills cells expressing constitutive MYC. FACS analysis (left) or crystal violet staining (right) of control and MYC-expressing U2OS cells, 4 days after retroviral expression of shARK5 (sequence 3 shown in panel b). EV, empty vector. Results are representative of two independent experiments. d, ARK5 kinase activity is required to prevent MYC-dependent death. U2OS MYC–ER cells expressing the indicated murine ARK5 point mutant proteins, superinfected with retroviruses expressing shArk5. Results are representative of three independent experiments. The immunoblot documents shRNA-resistant expression of murine ARK5. e, Pharmacological inhibition of ARK5 drives MYC-dependent death. U2OS MYC–ER cells were treated once with 50 nM BX795 or solvent, in the presence or absence of OHT and analysed after 5 days. Results are representative of four independent experiments.
Depletion of ARK5 induced the accumulation of MYC-expressing cells that stained positive for annexin V and propidium iodide (Fig. 1a and Supplementary Fig. 1a). Similarly, expressing different short hairpin (sh)RNAs targeting ARK5 induced levels of MYC-dependent death that correlated with the degree of knockdown (Fig. 1b). Titration of OHT revealed that levels of MYC that cause a dependence on ARK5 are higher than those required to promote proliferation (Supplementary Fig. 1b). Depletion of ARK5 induced death in U2OS cells constitutively expressing MYC and suppressed propagation of MRC5 fibroblasts in a MYC-dependent manner (Fig. 1c and Supplementary Fig. 1c). Expression of murine ARK5, which is not targeted by the shRNAs used, prevented death upon depletion of human ARK5 (Fig. 1d). This rescue required LKB1-dependent phosphorylation of T212, but not AKT-dependent phosphorylation of S601 (refs 3, 4). Mutation of K85 within the ATP-binding domain blocked the ability of murine ARK5 to prevent death, demonstrating that rescue requires ARK5 catalytic activity. Accordingly, a small-molecule inhibitor of ARK5, BX795, mimicked the effects of ARK5 depletion (Fig. 1e and Supplementary Fig. 1d–f)5.
Ectopic expression of BCL2 or MCL1, which protect cells from apoptosis induced by growth-factor deprivation, failed to alleviate the dependence on ARK5 (Supplementary Fig. 2a and data not shown)6. Depletion of ARK5 did not enhance the pro-apoptotic activity of E2F1 or E2F2 (Supplementary Figs 2b, c). ARK5 regulates the Hippo pathway, as it destabilizes LATS1 (Supplementary Figs 1d and 2d)7. However, co-depletion of LATS1 had no effect on death upon depletion of ARK5 (Supplementary Fig. 2d). Furthermore, depletion of ARK5 had little effect on MYC-dependent target gene activation (Supplementary Fig. 2e). To identify relevant effector pathways of ARK5, we characterized the effects of ARK5 depletion on cell physiology. Depletion of ARK5 delayed progression through all phases of the cell cycle (Fig. 2a, b). ARK5-depleted cells were larger than controls during S and G2 phase, demonstrating that ARK5 restricts cell growth (Fig. 2c and Supplementary Fig. 2f).
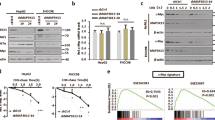
a, Depletion of ARK5 retards cell proliferation. U2OS cells were transfected with siARK5 and counted daily. Results are representative of four independent experiments. si con., control siRNA. b, Length of cell-cycle phases in ARK5-depleted U2OS cells. The percentage of cells in each phase of the cell cycle was determined by BrdU fluorescence-activated cell sorting (FACS). Using the doubling time from the experiment described in a, the length of each cell-cycle phase was calculated. Consistent results were obtained in two independent experiments. c, ARK5-depleted U2OS MYC–ER cells are larger than controls. ARK5-depleted and control cells were stained with Vybrant Dye violet, gated by DNA content and size was determined for G1, S and G2 cells. sh con., control shRNA. FSC, forward scatter. d, MYC-dependent activation of AMPK during culture of U2OS cells. Lysates were prepared from U2OS MYC–ER cells cultured in the presence of OHT and probed with the indicated antibodies. Results are representative of three independent experiments. p, phospho. e, Lysates from U2OS MYC–ER cells cultured with or without OHT were probed with anti-T172-phosphorylated AMPK-α1 and β-actin. f, ARK5 is required for activation of AMPK in response to MYC. U2OS MYC–ER cells transfected with control or siARK5, treated as per panel e. Results are representative of three independent experiments. AMPK-β1 denotes the β1 subunit of AMPK. g, ARK5 restricts global protein synthesis. U2OS MYC–ER cells, expressing shARK5 or control shRNA, treated with or without OHT and/or rapamycin and labelled after 24 h with 3H-leucine. Label incorporation per cell is shown. c.p.m., counts per minute. Consistent results were obtained using 3H-lysine and across three independent experiments in total. Results are shown as mean plus s.d.; *P values < 0.01; NS, not statistically significant.
AMPK inhibits the mTORC1 pathway that controls anabolic cell growth8. Different shRNAs targeting AMPK induced death in a MYC-dependent manner (Supplementary Fig. 3a). MYC promotes multiple anabolic processes and might thereby strain energy supplies and activate AMPK. Indeed, we observed a progressive increase in T172-phosphorylated AMPK9 and a progressive decease in mTORC1 activity in MYC-expressing cells (Fig. 2d, e). Depletion of ARK5 ablated activation of AMPK in response to MYC and to the AMP analogue aminoimidazole carboxamide ribonucleotide (AICAR) (Fig. 2f and Supplementary Fig. 3b). Consistently, depletion of ARK5 enhanced overall protein synthesis in a manner that was sensitive to the mTORC1 inhibitor rapamycin (Fig. 2g). Proteomic analysis demonstrated that ARK5 is required for protecting the β1 subunit of AMPK, which activates AMPK in response to alterations in the ATP:AMP ratio, from proteasomal degradation (Fig. 2f, Supplementary Fig. 3c and Supplementary Table 2)10. These findings suggested that unrestrained mTORC1 activity contributes to the death of MYC-expressing cells when depleted of ARK5 (ref. 11). Accordingly, addition of rapamycin protected cell viability in the absence of ARK5 or AMPK (Fig. 3a and Supplementary Fig. 3d). Similarly, addition of OSI-027, an inhibitor of mTORC1 and mTORC2, prevented death of ARK5-depleted cells, excluding the possibility that AKT signalling accounts for the blockade of death (Supplementary Fig. 3e)12,13.
a, Rapamycin protects MYC-overexpressing cells from ARK5 depletion. U2OS MYC–ER cells, expressing shARK5 or control shRNA (sh con.) and cultured in the presence of OHT with or without 100 nM rapamycin, as indicated. The percentage of apoptotic cells was determined daily by FACS and consistent results obtained in two independent time-course analyses. b, Depletion of ARK5 induces a MYC-dependent collapse of cellular ATP levels. U2OS MYC–ER cells transfected with control siRNA or siARK5 and treated with OHT. Viable cells were harvested daily. Levels of ATP are plotted relative to day 2. rapa., rapamycin. Results are representative of two experiments. c, Gas chromatography–mass spectrometry (GC–MS) analysis of carbon flow in MYC-transformed HCCs with or without ARK5. HCCs expressing MYC, AKT and doxycyline-inducible shARK5 were labelled with 13C-glucose or 13C-glutamine 48 h after induction of shARK5, as indicated. Graphs show the rate of label incorporation into TCA cycle intermediates (picomoles/1 × 106 cells per minute). Mean results and s.d. from three biological replicates, each with two technical replicates, are shown. ND, not detected. d, Proteomic analysis of ARK5 depletion. The chart summarizes results of a stable isotope labelling by amino acids in cell culture (SILAC) experiment measuring the abundance of total cellular proteins before and after depletion of ARK5. Subunit constituents of the mitochondrial respiratory complexes I, III and IV account for 29 of the 40 most suppressed proteins in ARK5-depleted cells. e, Induction of respiratory chain components by MYC is overridden by ARK5 depletion. Immunoblots show U2OS MYC–ER cells expressing shARK5 or control shRNA, treated with or without OHT for 24 h. Results are representative of two independent experiments. f, ARK5 depletion reduces mitochondrial membrane potential. U2OS MYC–ER cells expressing shARK5 or control shRNA, treated for 48 h with OHT, or acutely with carbonyl cyanide m-chlorophenyl hydrazone (CCCP), labelled with 3,3′-diethyloxacarbocyanine iodide (DiOC2(3)) and analysed by FACS. Left, representative FACS distribution of ARK5-depleted, OHT-treated cells (red) compared with untreated control cells (black). Right, percentage of cells with fluorescent intensity above 103 in each condition. Results are representative of two independent experiments. *P values < 0.01; NS, not statistically significant.
Depletion of ARK5 in MYC-expressing cells led to a progressive rapamycin-sensitive decline in ATP levels, which preceded death (Fig. 3a, b and Supplementary Fig. 4). Correlating with increased protein synthesis, depletion of ARK5 enhanced uptake of glutamine (Supplementary Fig. 5a). Because MYC-transformed cells depend on glutamine14,15,16, loss of ATP might be secondary to depletion of glutamine in the medium. However, daily addition of either glutamine or glucose failed to prevent death (Supplementary Fig. 5b). Furthermore, low levels of autophagy were observed in U2OS MYC–ER cells upon activation of MYC, and depletion of ARK5 moderately enhanced the accumulation of LC3 punctae, indicative of autophagy, suggesting that altered autophagy does not account for the loss of ATP (not shown)17. Finally, activation of MYC had only small effects on protein (Fig. 2g) and DNA synthesis (Supplementary Fig. 1b) that are unlikely to alone account for the ATP loss in MYC-expressing cells.
These results were recapitulated in a murine hepatocellular carcinoma line that expresses MYC and inducible shRNAs targeting ARK5 (see later). We pulse-labelled these cells with 13C-glucose or 13C-glutamine and traced the carbon flow. Consistent with previous demonstrations that deregulated MYC diverts glucose away from mitochondrial metabolism, flow from glucose into the tricarboxylic acid (TCA) cycle was low and most glucose was converted to lactate (Fig. 3c and Supplementary Fig. 5c, d)14,16. Depletion of ARK5 had little effect on glycolysis (Supplementary Fig. 5c). In line with enhanced uptake of glutamine, cells depleted of ARK5 exhibited increased flow from 13C-glutamine to α-ketoglutarate (Fig. 3c and Supplementary Fig. 5d). Depletion of ARK5 inhibited the entry of α-ketoglutarate into the TCA cycle, as flow into succinate, fumarate and malate was suppressed (Fig. 3c). Addition of membrane-permeable di-methyl-2-oxoglutarate did not rescue ARK5-depleted cells (data not shown). A reduction in TCA cycling was also evident from reduced flow into citrate, irrespective of the carbon source (Fig. 3c).
Depletion of ARK5 had no effect on the expression of α-ketoglutarate dehydrogenase subunits at either protein or messenger RNA levels (Supplementary Fig. 6a). A proteomic analysis revealed downregulation of multiple subunits of complexes I, III and IV of the mitochondrial respiratory chain (Fig. 3d and Supplementary Table 3; Gene Ontology term Benjamini enrichment score, 2 × 10−48). This regulation is post-transcriptional, specific and independent of the proteasome (Supplementary Tables 2, 4 and Supplementary Figs 3c, 6b). Immunoblot analyses confirmed previous observations that MYC upregulates expression of respiratory chain proteins and showed that depletion of ARK5 overrides this effect (Fig. 3e and Supplementary Fig. 6c)18,19. Depletion of ARK5 also lowered the mitochondrial membrane potential (Fig. 3f). Consistently, depletion of ARK5 reduced oxygen consumption, indicating that an inability to oxidize NADH limits entry of α-ketoglutarate into the TCA cycle (Supplementary Fig. 6d). Therefore, ARK5 is required to maintain sufficient respiratory capacity to sustain glutamine consumption in MYC transformed cells.
To ascertain if the restriction of mitochondrial respiration contributes to death, we limited mitochondrial function by depriving cells of nutrients or by blocking oxidative phosphorylation (Supplementary Fig. 7a). Cells cultured in the absence of nutrients and in the presence of oligomycin had undetectable levels of ATP and underwent cell death regardless of MYC expression. In the absence of nutrients, ATP levels declined more rapidly in MYC-expressing cells and these specifically underwent rapid cell death (Supplementary Fig. 7a–c). Consistently, depletion of ARK5 in MYC-expressing cells induced multiple hallmarks of low cellular ATP levels or limited electron transport, including an unfolded protein response, accumulation of DNA damage and enhanced levels of reactive oxygen species (Supplementary Fig. 7d–g).
Available databases document increased levels of ARK5 mRNA in hepatocellular and pancreatic carcinoma (Supplementary Fig. 8a). Immunohistochemistry revealed elevated ARK5 expression in the majority of colon, pancreatic and hepatocellular carcinoma cases and a high overlap with MYC expression (Fig. 4a and Supplementary Fig. 8b, c). Accordingly, depletion of ARK5 suppressed proliferation of 5/14 human tumour cell lines (Supplementary Fig. 8d). Depletion of ARK5 or AMPK induced cell death in Ls174T colon carcinoma cells, which harbour a mutant β-catenin gene that drives MYC expression20. Death was paralleled by loss of ATP and prevented by rapamycin (Supplementary Figs 9a, b). Co-depletion of MYC also prevented death, demonstrating that deregulated MYC establishes the dependence on ARK5 (Supplementary Fig. 9c). Moreover, depletion of ARK5 suppressed tumour formation by Ls174T cells (Supplementary Fig. 9d).
a, ARK5 is expressed in human cancers. Samples from human colorectal carcinoma (n = 26), pancreatic ductal adenocarcinoma (n = 21) and hepatocellular carcinoma (n = 11) were analysed for expression of ARK5 and MYC by immunohistochemistry (scale bar, 50 μm). b, Depletion of ARK5 suppresses propagation of HCC cells in a rapamycin-dependent manner. Top, immunoblots; bottom, crystal-violet-stained cultures of these cells. Consistent results were obtained in four independent experiments. c, ATP collapses after ARK5 depletion in HCC cells. ATP was measured daily in cells grown with or without doxycycline and/or rapamycin, as indicated. Results are plotted relative to untreated cultures. Consistent results were obtained in two independent experiments. Data are presented as mean ± s.d. d, Suppression of ARK5 prevents tumorigenesis in a model of HCC. Images of representative livers from mice transplanted with HCC cells carrying either of two doxycycline-inducible Ark5 shRNA vectors and treated with (n = 5 mice for each shRNA) or without (n = 4 mice for each shRNA) doxycycline from the time of transplantation. Tumours are outlined with a hashed border. Scale bar, 5 mm. e, Kaplan–Meier diagram documenting survival of mice orthotopically injected with 1 × 106 HCC cells. Expression of shArk5 (n = 4 mice) or control shRNA (n = 4 mice) was induced 9 days after transplantation (arrow). Statistical comparison of Kaplan–Meier curves is based on the log-rank test. f, Selection against shArk5 during HCC development. The panel shows representative examples of the liver of mice expressing shArk5 or control shRNA (left) and GFP staining documenting selection against GFP-positive tumour cells expressing shArk5 (right). Scale bar, 5 mm.
To test the therapeutic efficacy of ARK5 depletion, we transplanted murine p53−/− hepatoma cells that express MYC and AKT together with two different doxycycline-inducible Ark5 shRNAs under the liver capsule to induce orthotopic carcinomas (Supplementary Fig. 10a)21,22. In vitro, depletion of ARK5 led to culture collapse, preceded by loss of ATP (Fig. 4b, c, Supplementary Fig. 10b and data not shown). Addition of rapamycin restored ATP levels and allowed sustained culture of ARK5-depleted cells (Supplementary Fig. 10c). Upon transplantation, hepatocellular carcinomas (HCCs) developed in 8/8 untreated control mice, but only 1/10 mice treated with doxycycline (two-sided P value = 0.0004; Fig. 4d). Depletion of ARK5 also provided a survival advantage relative to cells expressing a shRNA targeting luciferase in an intervention study, in which tumours were allowed to develop before doxycycline was added (P < 0.01; Fig. 4e). The vector that drives the shRNA encodes GFP, reflecting shRNA expression. All control tumours were GFP positive, whereas expression of GFP was barely detectable in tumours containing shArk5, and tumour relapse was accompanied by re-expression of ARK5 mRNA (Fig. 4f and Supplementary Fig. 11a). Acute depletion of ARK5 activated mTOR and resulted in cell death and proliferative arrest in vivo (Supplementary Fig. 11b, c). Similar HCCs generated by co-expression of NRAS and MYC depended on ARK5, whereas HCCs generated by expression of NRAS in Arf−/− cells did not (Supplementary Fig. 12).
Synthetic lethal interactions allow the elimination of tumour cells carrying genetic lesions that cannot be targeted using small molecules23,24,25,26,27,28. We show that elevated energy consumption and addiction to mitochondrial glutaminolysis in cells expressing deregulated MYC establish a dependence on the kinase ARK5, which limits mTORC1 activity and maintains a high respiratory capacity. Our data support the view that oncogene-altered energy metabolism presents a new class of target molecules for tumour therapy.
Methods Summary
High-content screening
U2OS cells stably expressing MYC–ER under the control of a retroviral long terminal repeat (LTR) were transfected with siRNA in a 96-well format. Twenty-four hours after transfection, medium was replaced by medium with or without 200 nM OHT. Twenty-four hours later, cells were subjected to indirect immunofluorescence using cleaved-Parp (51-9000017, BD) followed by Alexa488-conjugated secondary antibody (Invitrogen). To reduce potential off-target effects, a pool of four individual siRNAs was transfected for each kinase. For automated data acquisition, the BD Pathway 855 bioimager (BD Biosciences) was used.
Metabolomic analysis
Cells were incubated with 13C6-glucose (Sigma) for 3 min or with 13C5-glutamine (Sigma) for 7 min. After labelling, cells were quenched with MeOH/water (1:1; −20 °C), extracted with ChCl3/MeOH/water and analysed by gas chromatography–time of flight–mass spectrometry (GC–TOF–MS) as described previously29. Mass isotopomers were extracted from individual mass spectra using the MetMax software tool30 and relative label incorporation was calculated after normalization to total area of measured analytes. For calculation of absolute concentrations of TCA cycle intermediates (all from Sigma) 8-point calibration curves spanning 2–3 orders of magnitudes were measured together with the samples.
Animal experiments
To deplete ARK5 in murine HCC cells, shRNAs were cloned into a retroviral vector for doxycycline-regulatable shRNA expression22. Early passage murine hepatoma cells were outgrown from genetically defined hepatocellular carcinomas (Myc; Akt; p53−/−). Cells were transduced with Ark5-shRNA-expressing retroviruses at low multiplicity of infection to ensure single-copy retroviral integration. After puromycin selection, 1,000,000 cells were transplanted under the liver capsula of female C57BL/6 mice. shRNA expression was induced by administration of doxycycline via the drinking water (1 mg ml−1 with 2% sucrose). Tumour progression was monitored by abdominal palpation and whole-body GFP imaging. All mice were maintained under pathogen-free conditions in accordance with the institutional guidelines of the Helmholtz Centre for Infection Research. All animal experiments were approved by the German legal authorities.
Online Methods
Cell culture
U2OS, MRC5, HCC4977 and Phoenix cells were cultured in DMEM (4.5 g ml−1 glucose; 2 mM glutamine) containing 10% FBS, 100 U ml−1 penicillin and 100 mg ml−1 streptomycin (PAA). Where indicated, 200 nM 4-hydroxytamoxifen (OHT; Sigma), 5 μg ml−1 doxycycline (Sigma), 2 mM caffeine (Sigma), 5 μg ml−1 oligomycin (Sigma), 500 µM AICAR (Sigma), 100 nm rapamycin (LC Labs), 50 nM BX795 (Invivogen) or 10 μM OSI-027 (Active Biochemicals) were added to the culture media. For ATP depletion experiments, cells were cultured in glucose- and glutamine-free DMEM (Sigma #D5030) supplemented with penicillin/streptomycin and 10% dialysed FBS (PAA). Apoptosis was measured by propidium iodide/annexin V (Molecular Probes) labelling and analysed using a BD FACS Canto II. Alternatively, cells were harvested by mild trypsinization and fixed in 80% ethanol, stained with 20 µg ml−1 propidium iodide in 0.1% Triton X-100/PBS containing 0.2 mg ml−1 RNAase A, and cellular DNA content was measured by FACS. siGENOME RNA oligonucleotides were purchased from Thermo Scientific Dharmacon, and cells were transfected using Dharmafect1 (Dharmacon) or Lipofectamine RNAiMAX (Invitrogen). Medium was replenished 24 h after transfection. Cellular ATP levels were determined using a luminescence-based ATP detection kit, ATPlite (PerkinElmer). For measurement of glutamine uptake, U2OS MYC–ER cells were incubated in DMEM medium and pulsed for 1 min with 1 µM 14C-labelled glutamine. The cells were washed three times with PBS and lysed in RIPA buffer for 30 min on ice. The lysates were analysed with a scintillation counter (Wallac 1410, Pharmacia). Mitochondrial membrane potential was measured FACS analysis of DiOC6(3) (Molecular Probes) stained cells, according to the manufacturer’s instructions. Reactive oxygen species were likewise measured using CellROX Deep Red (Invitrogen). Oxygen consumption rates were calculated from parafilm-sealed 10-cm dishes of cells using a Clarke O2 electrode.
High-content screening
U2OS cells stably expressing MYC–ER under the control of a retroviral long terminal repeat (LTR) were transfected with siRNA in a 96-well format. Twenty-four hours after transfection, medium was replaced by medium with or without 200 nM OHT. Twenty-four hours later, cells were subjected to indirect immunofluorescence using cleaved-Parp (51-9000017, BD) followed by Alexa488-conjugated secondary antibody (Invitrogen). To reduce potential off-target effects, a pool of four individual siRNAs was transfected for each kinase. For automated data acquisition, the BD Pathway 855 bioimager (BD Biosciences) was used.
Immunoblotting and antibodies
The following antibodies were purchased from Cell Signaling Technology: ARK5 (#4458), mTOR (#2972), phospho-mTOR (#2971), AMPK (#2532), phospho-AMPKThr172 (#2535), S6 ribosomal protein (#2212), phospho-S6S240/244 (#2215), tuberin/TSC2 (D57A9, #3990), phospho-tuberin/TSC2Ser1387 (#5584), p70 S6 kinase (49D7, #2708), phospho-p70 S6KThr389 (108D2, #9234). Anti-β-actin (AC-15, #A5441) and anti-mouse-ER (M-20) were purchased from Sigma. Cytochrome C Oxidase VIb (ab110266) and NduFS4 (ab87399) antibodies were purchased from Abcam. 9E10 and anti-mouse-ER were used to detect MYC and MYC–ER. Primary antibodies were used at 1:1,000, except for β-actin, which was used at 1:50,000. Secondary antibodies were purchased from Amersham.
Protein synthesis
For protein synthesis measurement U2OS cells were incubated for 2 h with 3H-leucine (50 µCi ml−1 medium) in DMEM. The cells were washed twice with PBS, after which the cells were incubated in 10% TCA for 10 min on ice. The TCA treatment was repeated twice for 5 min each followed by a washing step in MeOH. The cells were air dried and lysed in 0.3 M NaOH, 1% SDS for 30 min at room temperature (20–22 °C). The lysates were mixed with scintillation fluid (Rotiszint exo plus, Carl Roth) and measured with a scintillation counter (Wallac 1410, Pharmacia).
Plasmids
To construct shRNAs for human ARK5 and AMPK, hairpin-encoding oligonucleotides were annealed and ligated into pRetroSuper vector. The following targeting sequences were used: ARK5-1, GATGACAACTGCAATATTA; ARK5-2 GGACAGTAATGATGTGATG; ARK5-3, AGGACAAAATTAAGGATGA; AMPK-1, AAGTCAAAGTCGACCAAAT; AMPK-2, GCATAAAGTAGCTGTGAAG; AMPK-3, CAGCCGAGAAGCAGAAACA; AMPK-4, CCATACCCTTGATGAATTA. ARK5-3 was used for most experiments. To deplete murine ARK5, hairpin encoding nucleotides targeting AGGGATTTACTGGCATGGT (4977) and CGGTGGATGCTGATGGTGA (967) were inserted into pTGMP.
A full-length mouse cDNA encoding Ark5 was obtained from ImaGenes, and subcloned into pBabe vector. Site-directed mutagenesis using the QuikChange XL kit (Stratagene) was performed to generate constructs expressing mutant mouse ARK5. The following primers were used: K85A, forward, 5′-CCGAGTGGTTGCTATAGCATCCATCCGTAAGGAC-3′; K85A, reverse, 5′-GTCCTTACGGATGGATGCTATAGCAACCACTCGG-3′; T212A, forward, 5′-CAGAAGGACAAGTTCTTGCAAGCATTTTGTGGGAGCCCACTC-3′; T212A, reverse, 5′-GAGTGGGCTCCCACAAAATGCTTGCAAGAACTTGTCCTTCTG-3′; S601A, forward, 5′-GCCCGCCAGCGCATCCGCGCTTGCGTCTCTGCTGAAAAC-3′; S601A, reverse, 5′-GTTTTCAGCAGAGACGCAAGCGCGGATGCGCTGGCGGGC-3′.
Human tissues
Paraffin blocks previously used for diagnostic purposes were taken from the files of the Institute of Pathology of the University of Marburg. Tissues were fixed with 10% formalin. In all, 26 cases of colorectal carcinoma, 20 cases of hepatocellular carcinoma (of which 11 were co-examined for MYC expression) and 21 cases of pancreatic ductal adenocarcinoma were examined. All tissues were pseudonymized, without use of any personal data of the patients, in accordance with the local Ethics Committee.
Animal experiments
To deplete ARK5 in murine HCC cells, shRNAs were cloned into a retroviral vector for doxycycline-regulatable shRNA expression22. Early passage murine hepatoma cells were outgrown from genetically defined hepatocellular carcinomas (Myc; Akt; p53−/−). Cells were transduced with Ark5-shRNA-expressing retroviruses at low multiplicity of infection to ensure single-copy retroviral integration. After puromycin selection, 1,000,000 cells were transplanted under the liver capsula of female C57BL/6 mice. shRNA expression was induced by administration of doxycycline via the drinking water (1 mg ml−1 with 2% sucrose). Tumour progression was monitored by abdominal palpation and whole-body GFP imaging. All mice were maintained under pathogen-free conditions in accordance with the institutional guidelines of the Helmholtz Centre for Infection Research. All animal experiments were approved by the German legal authorities.
Immunohistochemistry
3–4-µm-thick paraffin sections were mounted on poly-l-lysine-coated slides, incubated at 58 °C, and deparaffinized. For antigen retrieval, sections were incubated in an antigen retrieval buffer (Tris-EDTA buffer, pH 9.0; Dako) for 30 min in a household steamer. The following incubations, including prior blocking of endogenous peroxidase activity, were performed using an automated immunohistochemistry apparatus (Autostainer plus; Dako): anti-ARK5 (Cell Signaling Technology, #4458) and anti c-MYC (Epitomics, #1472-1) antibodies were used at a dilution of 1:100. Detection was via Dako REAL Detection System Peroxidase/DAB+, rabbit/mouse; Dako) followed by staining with 3,3′-diaminobenzidine (DAB). For mild counterstaining, Mayer’s haematoxylin solution was used. Where indicated, sections of snap frozen tumour tissues were subjected to TdT-mediated dUTP nick end labelling (TUNEL) staining (Roche) and to Ki-67 (Dianova) immunohistochemistry using standard protocols. For negative controls, the primary antibody was replaced by buffer or an irrelevant monoclonal antibody. For phospho-S6, 5-μm sections were subjected to heat-based antigen retrieval in 10 mM sodium citrate, blocked for 1 h in 3% BSA and incubated with primary antibody (Cell Signaling #5364) at 1:5,000 overnight at 4 °C. Detection was with signal stain boost (Cell Signaling) for 45 min at room temperature.
Metabolomic analysis
Cells were incubated with 13C6-glucose (Sigma) for 3 min or with 13C5-glutamine (Sigma) for 7 min. After labelling, cells were quenched with MeOH/water (1:1; −20 °C), extracted with ChCl3/MeOH/water and analysed by gas chromatography–time of flight–mass spectrometry (GC–TOF–MS) as described previously29. Mass isotopomers were extracted from individual mass spectra using the MetMax software tool30 and relative label incorporation was calculated after normalization to total area of measured analytes. For calculation of absolute concentrations of TCA cycle intermediates (all from Sigma) 8-point calibration curves spanning 2–3 orders of magnitudes were measured together with the samples.
Proteome analysis
To achieve quantitative proteome data a ‘heavy’ proteome reference was spiked in equal amounts in every sample. HCC cells were grown in SILAC medium for 5 passages. SILAC medium was prepared as described previously31. In essence, DMEM lacking arginine and lysine was supplemented with 10% dialysed FBS (Sigma, F0392) and antibiotics. Amino acids (84 mg l−1 13C615N4 l-arginine plus 146 mg l−1 13C615N2 l-lysine) were added to obtain ‘heavy’ medium. Harvested cells were lysed in an appropriate amount of urea buffer (8 M urea, 50 mM Tris-HCl, pH 7.4). The lysates were cleared by centrifugation at 14,000 r.p.m. for 15 min at 4 °C. Disulphide bridges were then reduced in DTT 2 mM for 30 min at 25 °C and successively free cysteines were alkylated in 11 mM iodoacetamide for 20 min at room temperature in the darkness. LysC digestion was performed by adding LysC (Wako) in a ratio 1:40 (w/w) to the sample and incubating it for 18 h under gentle shaking at 30 °C. After LysC digestion, the samples were diluted 3 times with 50 mM ammonium bicarbonate solution, 7 µl of immobilized trypsin (Applied Biosystems) was added and samples were incubated for 4 h under rotation at 30 °C. Digestion was stopped by acidification with 10 μl of trifluoroacetic acid and removal of trypsin beads by centrifugation. After digestion peptides were extracted and desalted before analysis by mass spectrometry. 5 μl were injected in duplicate on a LC-MS/MS system (Agilent 1200 (Agilent Technologies) and LTQ-Orbitrap Velos (Thermo)), using a 240 min gradient ranging from 5% to 40% of solvent B (80% acetonitrile, 0.1% formic acid; solvent A = 5% acetonitrile, 0.1% formic acid). For the chromatographic separation, a ∼20-cm-long capillary (75 µm inner diameter) was packed with 3 µm C18 beads (ReprosilPur C18 AQ, Dr. Maisch). On one end of the capillary a nanospray tip was generated using a laser puller (P-2000 Laser Based Micropipette Puller, Sutter Instruments), allowing fretless packing. The nanospray source was operated with a spray voltage of 1.9 kV and an ion transfer tube temperature of 260 °C. Data were acquired in data dependent mode, with one survey MS scan in the Orbitrap mass analyser (resolution 60,000 at m/z 400) followed by up to 20 MS\MS scans in the ion trap on the most intense ions (intensity threshold, 500 counts). Once selected for fragmentation, ions were excluded from further selection for 30 s, to increase new sequencing events.
Data analysis
Raw data were analysed using the MaxQuant proteomics pipeline (v.1.1.1.36) and the built in the Andromeda search engine32 with the International Protein Index Mouse database. Carbamidomethylation of cysteines was chosen as fixed modification, oxidation of methionine and acetylation of N terminus were chosen as variable modifications. The search engine peptide assignments were filtered at 1% FDR and the feature match between runs was not enabled; second peptide feature was enabled, while other parameters were left as default. For SILAC samples, two ratio counts were set as threshold for quantification. Data analysis was performed using custom tools in Microsoft Excel and R. Gene Ontology analysis was performed using David tool33.
Microarray experiments were performed using Agilent whole human genome microarray kit (G4112F; Agilent Technologies) according to the manufacturer’s instructions.
Accession codes
Data deposits
Array results have been deposited in ArrayExpress (http://www.ebi.ac.uk/arrayexpress/) under accession number E-MEXP-3543.
References
Dang, C. V., Le, A. & Gao, P. MYC-induced cancer cell energy metabolism and therapeutic opportunities. Clin. Cancer Res. 15, 6479–6483 (2009)
Eilers, M. & Eisenman, R. N. Myc’s broad reach. Genes Dev. 22, 2755–2766 (2008)
Lizcano, J. M. et al. LKB1 is a master kinase that activates 13 kinases of the AMPK subfamily, including MARK/PAR-1. EMBO J. 23, 833–843 (2004)
Suzuki, A. et al. ARK5 is a tumor invasion-associated factor downstream of Akt signaling. Mol. Cell. Biol. 24, 3526–3535 (2004)
Clark, K., Plater, L., Peggie, M. & Cohen, P. Use of the pharmacological inhibitor BX795 to study the regulation and physiological roles of TBK1 and IκB kinase ε: a distinct upstream kinase mediates Ser-172 phosphorylation and activation. J. Biol. Chem. 284, 14136–14146 (2009)
Bissonnette, R. P., Echeverri, F., Mahboubi, A. & Green, D. R. Apoptotic cell death induced by c-myc is inhibited by bcl-2. Nature 359, 552–554 (1992)
Humbert, N. et al. Regulation of ploidy and senescence by the AMPK-related kinase NUAK1. EMBO J. 29, 376–386 (2010)
Shackelford, D. B. & Shaw, R. J. The LKB1–AMPK pathway: metabolism and growth control in tumour suppression. Nature Rev. Cancer 9, 563–575 (2009)
Hawley, S. A. et al. Use of cells expressing γ subunit variants to identify diverse mechanisms of AMPK activation. Cell Metab. 11, 554–565 (2010)
Oakhill, J. S. et al. β-Subunit myristoylation is the gatekeeper for initiating metabolic stress sensing by AMP-activated protein kinase (AMPK). Proc. Natl Acad. Sci. USA 107, 19237–19241 (2010)
Choo, A. Y. et al. Glucose addiction of TSC null cells is caused by failed mTORC1-dependent balancing of metabolic demand with supply. Mol. Cell 38, 487–499 (2010)
Carayol, N. et al. Critical roles for mTORC2- and rapamycin-insensitive mTORC1-complexes in growth and survival of BCR-ABL-expressing leukemic cells. Proc. Natl Acad. Sci. USA 107, 12469–12474 (2010)
Um, S. H. et al. Absence of S6K1 protects against age- and diet-induced obesity while enhancing insulin sensitivity. Nature 431, 200–205 (2004)
Gao, P. et al. c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 458, 762–765 (2009)
Yuneva, M., Zamboni, N., Oefner, P., Sachidanandam, R. & Lazebnik, Y. Deficiency in glutamine but not glucose induces MYC-dependent apoptosis in human cells. J. Cell Biol. 178, 93–105 (2007)
Wise, D. R. et al. Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc. Natl Acad. Sci. USA 105, 18782–18787 (2008)
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nature Cell Biol. 13, 132–141 (2011)
Zhang, H. et al. HIF-1 inhibits mitochondrial biogenesis and cellular respiration in VHL-deficient renal cell carcinoma by repression of C-MYC activity. Cancer Cell 11, 407–420 (2007)
Li, F. et al. Myc stimulates nuclearly encoded mitochondrial genes and mitochondrial biogenesis. Mol. Cell. Biol. 25, 6225–6234 (2005)
van de Wetering, M. et al. The β-catenin/TCF-4 complex imposes a crypt progenitor phenotype on colorectal cancer cells. Cell 111, 241–250 (2002)
Zender, L. et al. An oncogenomics-based in vivo RNAi screen identifies tumor suppressors in liver cancer. Cell 135, 852–864 (2008)
Zuber, J. et al. Toolkit for evaluating genes required for proliferation and survival using tetracycline-regulated RNAi. Nature Biotechnol. 29, 79–83 (2011)
Campaner, S. et al. Cdk2 suppresses cellular senescence induced by the c-myc oncogene. Nature Cell Biol. 12, 54–59 (2010)
Scholl, C. et al. Synthetic lethal interaction between oncogenic KRAS dependency and STK33 suppression in human cancer cells. Cell 137, 821–834 (2009)
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009)
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005)
Luo, J. et al. A genome-wide RNAi screen identifies multiple synthetic lethal interactions with the Ras oncogene. Cell 137, 835–848 (2009)
Yang, D. et al. Therapeutic potential of a synthetic lethal interaction between the MYC proto-oncogene and inhibition of aurora-B kinase. Proc. Natl Acad. Sci. USA 107, 13836–13841 (2010)
Kempa, S. et al. A plastid-localized glycogen synthase kinase 3 modulates stress tolerance and carbohydrate metabolism. Plant J. 49, 1076–1090 (2007)
Kempa, S. et al. An automated GC×GC-TOF-MS protocol for batch-wise extraction and alignment of mass isotopomer matrixes from differential 13C-labelling experiments: a case study for photoautotrophic-mixotrophic grown Chlamydomonas reinhardtii cells. J. Basic Microbiol. 49, 82–91 (2009)
Ong, S. E. & Mann, M. A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nature Protocols 1, 2650–2660 (2007)
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011)
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009)
Acknowledgements
We thank R. Baumann for immunohistochemical staining, B. Samans, A. Filmer and M. Krause for help with the siRNA screen and expression array analysis, A. Rath for help with analysis of human tumour samples, W. Kolch for siRNA targeting LATS1, S. Lowe and J. Zuber for providing the tTGMP vector, S. Martin for His-tagged annexin V expression vector, P. Schirmacher for human HCC cell lines, V. Ellenrieder for human pancreatic cancer cell lines, H. Riedesel and the staff of the Helmholtz Centre for Infection Research animal facility for technical assistance. We thank P. Gallant and all members of the Eilers laboratory for critical reading of the manuscript. Funding for this work was provided by the GROWTHSTOP consortium of the European Union, the Deutsche Forschungsgemeinschaft via the Transregio 17 (M.E.), the Transregio 77 (L.Z.) and the Emmy Noether Programme ZE 545/2-1 (L.Z.), the Bundesministerium für Bildung und Forschung, the Senate of Berlin, the University of Wuerzburg Graduate School of Life Sciences and the “Rebirth” Cluster of Excellence.
Author information
Authors and Affiliations
Contributions
L.L., J.U., J.M., T.R.K., N.M. and R.R. performed the experiments, R.M. analysed human tumour samples, and T.W. analysed the mouse model of HCC. L.A. and S.K. performed all metabolomic and proteomic analyses and questions regarding the technology should be addressed to S.K. L.R. performed bio-informatic analysis. L.Z., S.K., M.E. and D.J.M. designed experiments, and M.E. and D.J.M. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures1-12 and Supplementary Tables 1-4. (PDF 4734 kb)
Rights and permissions
About this article
Cite this article
Liu, L., Ulbrich, J., Müller, J. et al. Deregulated MYC expression induces dependence upon AMPK-related kinase 5. Nature 483, 608–612 (2012). https://doi.org/10.1038/nature10927
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10927
This article is cited by
-
ARK5 enhances cell survival associated with mitochondrial morphological dynamics from fusion to fission in human multiple myeloma cells
Cell Death Discovery (2024)
-
NUAK1 coordinates growth factor-dependent activation of mTORC2 and Akt signaling
Cell & Bioscience (2023)
-
Derivation of healthy hepatocyte-like cells from a female patient with ornithine transcarbamylase deficiency through X-inactivation selection
Scientific Reports (2022)
-
Loss of LKB1-NUAK1 signalling enhances NF-κB activity in a spheroid model of high-grade serous ovarian cancer
Scientific Reports (2022)
-
MicroRNA-1182 and let-7a exert synergistic inhibition on invasion, migration and autophagy of cholangiocarcinoma cells through down-regulation of NUAK1
Cancer Cell International (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.