Abstract
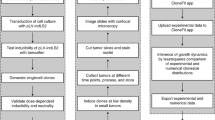
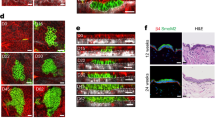
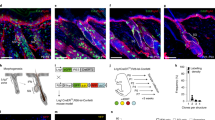
Recent studies using the isolation of a subpopulation of tumour cells followed by their transplantation into immunodeficient mice provide evidence that certain tumours1,2, including squamous skin tumours3,4,5, contain cells with high clonogenic potential that have been referred to as cancer stem cells (CSCs). Until now, CSC properties have only been investigated by transplantation assays, and their existence in unperturbed tumour growth is unproven. Here we make use of clonal analysis of squamous skin tumours using genetic lineage tracing to unravel the mode of tumour growth in vivo in its native environment. To this end, we used a genetic labelling strategy that allows individual tumour cells to be marked and traced over time at different stages of tumour progression. Surprisingly, we found that the majority of labelled tumour cells in benign papilloma have only limited proliferative potential, whereas a fraction has the capacity to persist long term, giving rise to progeny that occupy a significant part of the tumour. As well as confirming the presence of two distinct proliferative cell compartments within the papilloma, mirroring the composition, hierarchy and fate behaviour of normal tissue, quantitative analysis of clonal fate data indicates that the more persistent population has stem-cell-like characteristics and cycles twice per day, whereas the second represents a slower cycling transient population that gives rise to terminally differentiated tumour cells. Such behaviour is shown to be consistent with double-labelling experiments and detailed clonal fate characteristics. By contrast, measurements of clone size and proliferative potential in invasive squamous cell carcinoma show a different pattern of behaviour, consistent with geometric expansion of a single CSC population with limited potential for terminal differentiation. This study presents the first experimental evidence for the existence of CSCs during unperturbed solid tumour growth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shackleton, M., Quintana, E., Fearon, E. R. & Morrison, S. J. Heterogeneity in cancer: cancer stem cells versus clonal evolution. Cell 138, 822–829 (2009)
Lobo, N. A., Shimono, Y., Qian, D. & Clarke, M. F. The biology of cancer stem cells. Annu. Rev. Cell. Dev. Biol. 23, 675–699 (2007)
Malanchi, I. et al. Cutaneous cancer stem cell maintenance is dependent on β-catenin signalling. Nature 452, 650–653 (2008)
Beck, B. et al. A vascular niche and a VEGF–Nrp1 loop regulate the initiation and stemness of skin tumours. Nature 478, 399–403 (2011)
Schober, M. & Fuchs, E. Tumor-initiating stem cells of squamous cell carcinomas and their control by TGF-β and integrin/focal adhesion kinase (FAK) signaling. Proc. Natl Acad. Sci. USA 108, 10544–10549 (2011)
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976)
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012)
Nguyen, L. V., Vanner, R., Dirks, P. & Eaves, C. J. Cancer stem cells: an evolving concept. Nature Rev. Cancer 12, 133–143 (2012)
Kemp, C. J. Multistep skin cancer in mice as a model to study the evolution of cancer cells. Semin. Cancer Biol. 15, 460–473 (2005)
Abel, E. L., Angel, J. M., Kiguchi, K. & DiGiovanni, J. Multi-stage chemical carcinogenesis in mouse skin: fundamentals and applications. Nature Protocols 4, 1350–1362 (2009)
Owens, D. M. & Watt, F. M. Contribution of stem cells and differentiated cells to epidermal tumours. Nature Rev. Cancer 3, 444–451 (2003)
Perez-Losada, J. & Balmain, A. Stem-cell hierarchy in skin cancer. Nature Rev. Cancer 3, 434–443 (2003)
Klein, C. A. Parallel progression of primary tumours and metastases. Nature Rev. Cancer 9, 302–312 (2009)
Clayton, E. et al. A single type of progenitor cell maintains normal epidermis. Nature 446, 185–189 (2007)
Doupé, D. P., Klein, A. M., Simons, B. D. & Jones, P. H. The ordered architecture of murine ear epidermis is maintained by progenitor cells with random fate. Dev. Cell 18, 317–323 (2010)
Lopez-Garcia, C., Klein, A. M., Simons, B. D. & Winton, D. J. Intestinal stem cell replacement follows a pattern of neutral drift. Science 330, 822–825 (2010)
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010)
Simons, B. D. & Clevers, H. Strategies for homeostatic stem cell self-renewal in adult tissues. Cell 145, 851–862 (2011)
Klein, A. M. & Simons, B. D. Universal patterns of stem cell fate in cycling adult tissues. Development 138, 3103–3111 (2011)
Ghazizadeh, S. & Taichman, L. B. Multiple classes of stem cells in cutaneous epithelium: a lineage analysis of adult mouse skin. EMBO J. 20, 1215–1222 (2001)
Potten, C. S. Cell replacement in epidermis (keratopoiesis) via discrete units of proliferation. Int. Rev. Cytol. 69, 271–318 (1981)
Mackenzie, I. C. Retroviral transduction of murine epidermal stem cells demonstrates clonal units of epidermal structure. J. Invest. Dermatol. 109, 377–383 (1997)
Marusyk, A. & Polyak, K. Tumor heterogeneity: causes and consequences. Biochim. Biophys. Acta 1805, 105–117 (2010)
Van Keymeulen, A. et al. Epidermal progenitors give rise to Merkel cells during embryonic development and adult homeostasis. J. Cell Biol. 187, 91–100 (2009)
Rocheteau, P., Gayraud-Morel, B., Siegl-Cachedenier, I., Blasco, M. A. & Tajbakhsh, S. A subpopulation of adult skeletal muscle stem cells retains all template DNA strands after cell division. Cell 148, 112–125 (2012)
Vasioukhin, V., Degenstein, L., Wise, B. & Fuchs, E. The magical touch: genome targeting in epidermal stem cells induced by tamoxifen application to mouse skin. Proc. Natl Acad. Sci. USA 96, 8551–8556 (1999)
Acknowledgements
C.B. is investigator of WELBIO and chercheur qualifié of the FRS/FNRS. B.B. is chargé de recherche of the FRS/FNRS and G.D. is supported by the Brussels Region. This work was supported by the FNRS, the program d’excellence CIBLES of the Wallonia Region, a research grant from the Fondation Contre le Cancer, the ULB foundation and the fond Gaston Ithier, a starting grant of the European Research Council (ERC), and the EMBO Young Investigator Program. We thank F. Bollet-Quivogne and J.-M. Vanderwinden for their help with confocal imaging.
Author information
Authors and Affiliations
Contributions
C.B., G.D., B.B. and B.D.S. designed the experiments and performed data analysis. G.D. and B.B. performed most of the experiments. A.C. provided technical support. C.B. and B.D.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-9 and Supplementary Methods and Data. (PDF 4159 kb)
Rights and permissions
About this article
Cite this article
Driessens, G., Beck, B., Caauwe, A. et al. Defining the mode of tumour growth by clonal analysis. Nature 488, 527–530 (2012). https://doi.org/10.1038/nature11344
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11344
This article is cited by
-
Dynamics of a diffusive model for cancer stem cells with time delay in microRNA-differentiated cancer cell interactions and radiotherapy effects
Scientific Reports (2024)
-
Liquid biopsy techniques and pancreatic cancer: diagnosis, monitoring, and evaluation
Molecular Cancer (2023)
-
Pancreatic cancer stemness: dynamic status in malignant progression
Journal of Experimental & Clinical Cancer Research (2023)
-
Advances in cutaneous squamous cell carcinoma
Nature Reviews Cancer (2023)
-
The emerging role of the gut microbiome in cancer cell plasticity and therapeutic resistance
Cancer and Metastasis Reviews (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.