Abstract
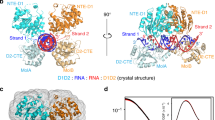
DEAD-box proteins are the largest family of nucleic acid helicases, and are crucial to RNA metabolism throughout all domains of life1,2. They contain a conserved ‘helicase core’ of two RecA-like domains (domains (D)1 and D2), which uses ATP to catalyse the unwinding of short RNA duplexes by non-processive, local strand separation3. This mode of action differs from that of translocating helicases and allows DEAD-box proteins to remodel large RNAs and RNA–protein complexes without globally disrupting RNA structure4. However, the structural basis for this distinctive mode of RNA unwinding remains unclear. Here, structural, biochemical and genetic analyses of the yeast DEAD-box protein Mss116p indicate that the helicase core domains have modular functions that enable a novel mechanism for RNA-duplex recognition and unwinding. By investigating D1 and D2 individually and together, we find that D1 acts as an ATP-binding domain and D2 functions as an RNA-duplex recognition domain. D2 contains a nucleic-acid-binding pocket that is formed by conserved DEAD-box protein sequence motifs and accommodates A-form but not B-form duplexes, providing a basis for RNA substrate specificity. Upon a conformational change in which the two core domains join to form a ‘closed state’ with an ATPase active site, conserved motifs in D1 promote the unwinding of duplex substrates bound to D2 by excluding one RNA strand and bending the other. Our results provide a comprehensive structural model for how DEAD-box proteins recognize and unwind RNA duplexes. This model explains key features of DEAD-box protein function and affords a new perspective on how the evolutionarily related cores of other RNA and DNA helicases diverged to use different mechanisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Linder, P. & Jankowsky, E. From unwinding to clamping—the DEAD box RNA helicase family. Nature Rev. Mol. Cell Biol. 12, 505–516 (2011)
Jarmoskaite, I. & Russell, R. DEAD-box proteins as RNA helicases and chaperones. WIREs: RNA 2, 135–152 (2011)
Yang, Q., Del Campo, M., Lambowitz, A. M. & Jankowsky, E. DEAD-box proteins unwind duplexes by local strand separation. Mol. Cell 28, 253–263 (2007)
Pan, C. & Russell, R. Roles of DEAD-box proteins in RNA and RNP folding. RNA Biol. 7, 667–676 (2010)
Huang, H. R. et al. The splicing of yeast mitochondrial group I and group II introns requires a DEAD-box protein with RNA chaperone function. Proc. Natl Acad. Sci. USA 102, 163–168 (2005)
Del Campo, M. et al. Unwinding by local strand separation is critical for the function of DEAD-box proteins as RNA chaperones. J. Mol. Biol. 389, 674–693 (2009)
Potratz, J. P., Del Campo, M., Wolf, R. Z., Lambowitz, A. M. & Russell, R. ATP-dependent roles of the DEAD-box protein Mss116p in group II intron splicing in vitro and in vivo. J. Mol. Biol. 411, 661–679 (2011)
Cao, W. et al. Mechanism of Mss116 ATPase reveals functional diversity of DEAD-box proteins. J. Mol. Biol. 409, 399–414 (2011)
Mohr, G. et al. High-throughput genetic identification of functionally important regions of the yeast DEAD-box protein Mss116p. J. Mol. Biol. 413, 952–972 (2011)
Del Campo, M. & Lambowitz, A. M. Structure of the yeast DEAD-box protein Mss116p reveals two wedges that crimp RNA. Mol. Cell 35, 598–609 (2009)
Mallam, A. L. et al. Solution structures of DEAD-box RNA chaperones reveal conformational changes and nucleic acid tethering by a basic tail. Proc. Natl Acad. Sci. USA 108, 12254–12259 (2011)
Wang, S., Overgaard, M. T., Hu, Y. & McKay, D. B. The Bacillus subtilis RNA helicase YxiN is distended in solution. Biophys. J. 94, L01–L03 (2008)
Sengoku, T., Nureki, O., Nakamura, A., Kobayashi, S. & Yokoyama, S. Structural basis for RNA unwinding by the DEAD-box protein Drosophila Vasa. Cell 125, 287–300 (2006)
Rudolph, M. G., Heissmann, R., Wittmann, J. G. & Klostermeier, D. Crystal structure and nucleotide binding of the Thermus thermophilus RNA helicase Hera N-terminal domain. J. Mol. Biol. 361, 731–743 (2006)
Schütz, P. et al. Comparative structural analysis of human DEAD-box RNA helicases. PLoS ONE 5, e12791 (2010)
Egli, M., Usman, N. & Rich, A. Conformational influence of the ribose 2′-hydroxyl group: crystal structures of DNA-RNA chimeric duplexes. Biochemistry 32, 3221–3237 (1993)
Wahl, M. C. & Sundaralingam, M. B-form to A-form conversion by a 3′-terminal ribose: crystal structure of the chimera d(CCACTAGTG)r(G). Nucleic Acids Res. 28, 4356–4363 (2000)
Halls, C. et al. Involvement of DEAD-box proteins in group I and group II intron splicing. Biochemical characterization of Mss116p, ATP hydrolysis-dependent and -independent mechanisms, and general RNA chaperone activity. J. Mol. Biol. 365, 835–855 (2007)
Henn, A. et al. Pathway of ATP utilization and duplex rRNA unwinding by the DEAD-box helicase, DbpA. Proc. Natl Acad. Sci. USA 107, 4046–4050 (2010)
Rocak, S. & Linder, P. DEAD-box proteins: the driving forces behind RNA metabolism. Nature Rev. Mol. Cell Biol. 5, 232–241 (2004)
Chen, Y. et al. DEAD-box proteins can completely separate an RNA duplex using a single ATP. Proc. Natl Acad. Sci. USA 105, 20203–20208 (2008)
Liu, F., Putnam, A. & Jankowsky, E. ATP hydrolysis is required for DEAD-box protein recycling but not for duplex unwinding. Proc. Natl Acad. Sci. USA 105, 20209–20214 (2008)
Andersen, C. B. F. et al. Structure of the exon junction core complex with a trapped DEAD-box ATPase bound to RNA. Science 313, 1968–1972 (2006)
Fairman-Williams, M. E., Guenther, U. P. & Jankowsky, E. SF1 and SF2 helicases: family matters. Curr. Opin. Struct. Biol. 20, 313–324 (2010)
Singleton, M. R., Dillingham, M. S. & Wigley, D. B. Structure and mechanism of helicases and nucleic acid translocases. Annu. Rev. Biochem. 76, 23–50 (2007)
Gorbalenya, A. E., Koonin, E. V., Donchenko, A. P. & Blinov, V. M. Two related superfamilies of putative helicases involved in replication, recombination, repair and expression of DNA and RNA genomes. Nucleic Acids Res. 17, 4713–4730 (1989)
Kowalinski, E. et al. Structural basis for the activation of innate immune pattern-recognition receptor RIG-I by viral RNA. Cell 147, 423–435 (2011)
Jiang, F. et al. Structural basis of RNA recognition and activation by innate immune receptor RIG-I. Nature 479, 423–427 (2011)
Luo, D. et al. Structural insights into RNA recognition by RIG-I. Cell 147, 409–422 (2011)
Ozalp, V. C., Pedersen, T. R., Nielsen, L. J. & Olsen, L. F. Time-resolved measurements of intracellular ATP in the yeast Saccharomyces cerevisiae using a new type of nanobiosensor. J. Biol. Chem. 285, 37579–37588 (2010)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. D 67, 293–302 (2011)
Pflugrath, J. W. The finer things in X-ray diffraction data collection. Acta Crystallogr. D 55, 1718–1725 (1999)
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D 62, 72–82 (2006)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007)
Wilkins, M. R. et al. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 112, 531–552 (1999)
Tang, G. Q., Bandwar, R. P. & Patel, S. S. Extended upstream A-T sequence increases T7 promoter strength. J. Biol. Chem. 280, 40707–40713 (2005)
Acknowledgements
We thank A. Monzingo (University of Texas at Austin Macromolecular Crystallography Facility) for help with X-ray diffraction data collection, and R. Russell (University of Texas at Austin) and E. Jankowsky (Case Western Reserve University) for comments on the manuscript. X-ray diffraction data were collected at the Berkeley Center for Structural Biology, which is supported in part by the National Institutes of Health (NIH), National Institute of General Medical Sciences and the Howard Hughes Medical Institute. The Advanced Light Source is supported by the Director, Office of Science, Office of Basic Energy Sciences, of the US Department of Energy under contract no. DE-AC02-05CH11231. A.L.M. is the recipient of an EMBO long-term fellowship (ALTF 389-2010). This work was supported by NIH grant GM037951.
Author information
Authors and Affiliations
Contributions
This study was designed by A.L.M. and A.M.L. A.L.M. cloned the individual domain constructs, purified proteins, crystallized the complexes, collected X-ray crystallographic data, and performed binding assays. A.L.M., M.D.C. and D.J.S. processed and refined the X-ray diffraction data. B.G. performed genetic assays. All authors contributed to analysing the results. A.L.M., M.D.C. and A.M.L. wrote the paper, with contributions from D.J.S. and B.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-10, Supplementary Tables 1-2 and Supplementary References. (PDF 10910 kb)
Rights and permissions
About this article
Cite this article
Mallam, A., Del Campo, M., Gilman, B. et al. Structural basis for RNA-duplex recognition and unwinding by the DEAD-box helicase Mss116p. Nature 490, 121–125 (2012). https://doi.org/10.1038/nature11402
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11402
This article is cited by
-
Autoinhibition of a clamp-loader ATPase revealed by deep mutagenesis and cryo-EM
Nature Structural & Molecular Biology (2024)
-
Cellular functions of eukaryotic RNA helicases and their links to human diseases
Nature Reviews Molecular Cell Biology (2023)
-
The mechanism of RNA duplex recognition and unwinding by DEAD-box helicase DDX3X
Nature Communications (2019)
-
Cryo-EM structure of an early precursor of large ribosomal subunit reveals a half-assembled intermediate
Protein & Cell (2019)
-
DEAD-box helicase 27 promotes colorectal cancer growth and metastasis and predicts poor survival in CRC patients
Oncogene (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.