Abstract
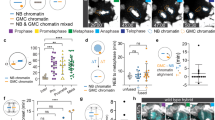
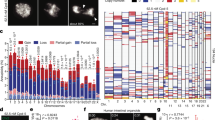
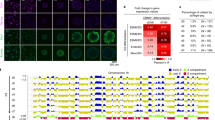
Adult stem cells undergo asymmetric cell division to self-renew and give rise to differentiated cells that comprise mature tissue1. Sister chromatids may be distinguished and segregated nonrandomly in asymmetrically dividing stem cells2, although the underlying mechanism and the purpose it may serve remain elusive. Here we develop the CO-FISH (chromosome orientation fluorescence in situ hybridization) technique3 with single-chromosome resolution and show that sister chromatids of X and Y chromosomes, but not autosomes, are segregated nonrandomly during asymmetric divisions of Drosophila male germline stem cells. This provides the first direct evidence, to our knowledge, that two sister chromatids containing identical genetic information can be distinguished and segregated nonrandomly during asymmetric stem-cell divisions. We further show that the centrosome, SUN–KASH nuclear envelope proteins and Dnmt2 (also known as Mt2) are required for nonrandom sister chromatid segregation. Our data indicate that the information on X and Y chromosomes that enables nonrandom segregation is primed during gametogenesis in the parents. Moreover, we show that sister chromatid segregation is randomized in germline stem cell overproliferation and dedifferentiated germline stem cells. We propose that nonrandom sister chromatid segregation may serve to transmit distinct information carried on two sister chromatids to the daughters of asymmetrically dividing stem cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Morrison, S. J. & Kimble, J. Asymmetric and symmetric stem-cell divisions in development and cancer. Nature 441, 1068–1074 (2006)
Tajbakhsh, S. & Gonzalez, C. Biased segregation of DNA and centrosomes: moving together or drifting apart? Nature Rev. Mol. Cell Biol. 10, 804–810 (2009)
Falconer, E. et al. Identification of sister chromatids by DNA template strand sequences. Nature 463, 93–97 (2010)
Fuller, M. T. & Spradling, A. C. Male and female Drosophila germline stem cells: two versions of immortality. Science 316, 402–404 (2007)
Yamashita, Y. M., Jones, D. L. & Fuller, M. T. Orientation of asymmetric stem cell division by the APC tumor suppressor and centrosome. Science 301, 1547–1550 (2003)
Yamashita, Y. M., Mahowald, A. P., Perlin, J. R. & Fuller, M. T. Asymmetric inheritance of mother versus daughter centrosome in stem cell division. Science 315, 518–521 (2007)
Dernburg, A. F. in Drosophila Protocols (eds Sullivan, W., Ashburner, M. & Hawley, R. S.) Ch. 2 (CSHL Press, 2000)
Fung, J. C., Marshall, W. F., Dernburg, A., Agard, D. A. & Sedat, J. W. Homologous chromosome pairing in Drosophila melanogaster proceeds through multiple independent initiations. J. Cell Biol. 141, 5–20 (1998)
Makunin, I. V. et al. A novel simple satellite DNA is colocalized with the Stalker retrotransposon in Drosophila melanogaster heterochromatin. Mol. Gen. Genet. 261, 381–387 (1999)
Li, K. & Kaufman, T. C. The homeotic target gene centrosomin encodes an essential centrosomal component. Cell 85, 585–596 (1996)
Kracklauer, M. P., Banks, S. M., Xie, X., Wu, Y. & Fischer, J. A. Drosophila klaroid encodes a SUN domain protein required for Klarsicht localization to the nuclear envelope and nuclear migration in the eye. Fly (Austin) 1, 75–85 (2007)
Mosley-Bishop, K. L., Li, Q., Patterson, L. & Fischer, J. A. Molecular analysis of the klarsicht gene and its role in nuclear migration within differentiating cells of the Drosophila eye. Curr. Biol. 9, 1211–1220 (1999)
Razafsky, D. & Hodzic, D. Bringing KASH under the SUN: the many faces of nucleo-cytoskeletal connections. J. Cell Biol. 186, 461–472 (2009)
Phalke, S. et al. Retrotransposon silencing and telomere integrity in somatic cells of Drosophila depends on the cytosine-5 methyltransferase DNMT2. Nature Genet. 41, 696–702 (2009)
Kunert, N., Marhold, J., Stanke, J., Stach, D. & Lyko, F. A. Dnmt2-like protein mediates DNA methylation in Drosophila. Development 130, 5083–5090 (2003)
Schaefer, M. et al. RNA methylation by Dnmt2 protects transfer RNAs against stress-induced cleavage. Genes Dev. 24, 1590–1595 (2010)
Zemach, A., McDaniel, I. E., Silva, P. & Zilberman, D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science 328, 916–919 (2010)
Brawley, C. & Matunis, E. Regeneration of male germline stem cells by spermatogonial dedifferentiation in vivo. Science 304, 1331–1334 (2004)
Kai, T. & Spradling, A. Differentiating germ cells can revert into functional stem cells in Drosophila melanogaster ovaries. Nature 428, 564–569 (2004)
Cheng, J. et al. Centrosome misorientation reduces stem cell division during ageing. Nature 456, 599–604 (2008)
Yadlapalli, S., Cheng, J. & Yamashita, Y. M. Drosophila male germline stem cells do not asymmetrically segregate chromosome strands. J. Cell Sci. 124, 933–939 (2011)
Conrad, T. & Akhtar, A. Dosage compensation in Drosophila melanogaster: epigenetic fine-tuning of chromosome-wide transcription. Nature Rev. Genet. 13, 123–134 (2012)
Hense, W., Baines, J. F. & Parsch, J. X chromosome inactivation during Drosophila spermatogenesis. PLoS Biol. 5, e273 (2007)
Aravin, A. A. et al. Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr. Biol. 11, 1017–1027 (2001)
Tulin, A. V., Kogan, G. L., Filipp, D., Balakireva, M. D. & Gvozdev, V. A. Heterochromatic Stellate gene cluster in Drosophila melanogaster: structure and molecular evolution. Genetics 146, 253–262 (1997)
Tran, V., Lim, C., Xie, J. & Chen, X. Asymmetric division of Drosophila male germline stem cell shows asymmetric histone distribution. Science 338, 679–682 (2012)
Minestrini, G., Mathe, E. & Glover, D. M. Domains of the Pavarotti kinesin-like protein that direct its subcellular distribution: effects of mislocalisation on the tubulin and actin cytoskeleton during Drosophila oogenesis. J. Cell Sci. 115, 725–736 (2002)
Petrella, L. N., Smith-Leiker, T. & Cooley, L. The Ovhts polyprotein is cleaved to produce fusome and ring canal proteins required for Drosophila oogenesis. Development 134, 703–712 (2007)
Sheng, X. R., Brawley, C. M. & Matunis, E. L. Dedifferentiating spermatogonia outcompete somatic stem cells for niche occupancy in the Drosophila testis. Cell Stem Cell 5, 191–203 (2009)
Förstemann, K. et al. Normal microRNA maturation and germ-line stem cell maintenance requires Loquacious, a double-stranded RNA-binding domain protein. PLoS Biol. 3, e236 (2005)
Acknowledgements
We thank F. Lyko, M. Schaefer, G. Reuter, P. Zamore, A. Aravin, D. Glover, L. Cooley, J. Kim, V. Gvozdev, M. Pia Bozzetti, the Bloomington Drosophila Stock Center and the Vienna Drosophila RNAi Center for reagents and helpful information, and Yamashita laboratory members for discussions. This study was supported by the University of Michigan (Life Sciences Institute and Office of the Provost and Executive Vice President for Academic Affairs) (to Y.M.Y.) and AHA (12PRE9630000) and NIH grants (1F31HD071727-01) (to S.Y.). Y.M.Y. is supported by the MacArthur Foundation.
Author information
Authors and Affiliations
Contributions
S.Y. conceived the project and developed the single-chromosome CO-FISH protocol for Drosophila cells. S.Y. and Y.M.Y. designed and conducted experiments, interpreted the data, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-6 and Supplementary Tables 1-3. (PDF 774 kb)
Rights and permissions
About this article
Cite this article
Yadlapalli, S., Yamashita, Y. Chromosome-specific nonrandom sister chromatid segregation during stem-cell division. Nature 498, 251–254 (2013). https://doi.org/10.1038/nature12106
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12106
This article is cited by
-
Asymmetric division of stem cells and its cancer relevance
Cell Regeneration (2024)
-
Asymmetric chromatin retention and nuclear envelopes separate chromosomes in fused cells in vivo
Communications Biology (2022)
-
Interchromosomal interaction of homologous Stat92E alleles regulates transcriptional switch during stem-cell differentiation
Nature Communications (2022)
-
Budding yeast Wee1 distinguishes spindle pole bodies to guide their pattern of age-dependent segregation
Nature Cell Biology (2017)
-
Regulation of Stem Cells in Their Niche
Current Stem Cell Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.