Abstract
The proteasomal ATPase ring, comprising Rpt1–Rpt6, associates with the heptameric α-ring of the proteasome core particle (CP) in the mature proteasome, with the Rpt carboxy-terminal tails inserting into pockets of the α-ring1,2,3,4. Rpt ring assembly is mediated by four chaperones, each binding a distinct Rpt subunit5,6,7,8,9,10. Here we report that the base subassembly of the Saccharomyces cerevisiae proteasome, which includes the Rpt ring, forms a high-affinity complex with the CP. This complex is subject to active dissociation by the chaperones Hsm3, Nas6 and Rpn14. Chaperone-mediated dissociation was abrogated by a non-hydrolysable ATP analogue, indicating that chaperone action is coupled to nucleotide hydrolysis by the Rpt ring. Unexpectedly, synthetic Rpt tail peptides bound α-pockets with poor specificity, except for Rpt6, which uniquely bound the α2/α3-pocket. Although the Rpt6 tail is not visualized within an α-pocket in mature proteasomes2,3,4, it inserts into the α2/α3-pocket in the base–CP complex and is important for complex formation. Thus, the Rpt–CP interface is reconfigured when the lid complex joins the nascent proteasome to form the mature holoenzyme.
Similar content being viewed by others
Main
The proteasome mediates selective protein degradation in eukaryotes1. It is composed of a 19-subunit regulatory particle (RP; also known as PA700 or the 19S complex) and a 28-subunit proteolytic CP (also known as the 20S complex) composed of four stacked heptameric rings. Ubiquitinated proteins are recognized by the RP, translocated into the CP via a channel in its outer (α)-ring, and degraded. The RP comprises a 10-subunit base and 9-subunit lid1,2,3,4. Central to the base is the Rpt ring, a heterohexameric ATPase complex that abuts the α-ring, with flexible C-terminal tails of multiple Rpt subunits inserting into ‘α-pockets’ of the CP11,12,13,14.
The Rpt ring is formed from three modules1: Rpt3–Rpt6; Rpt4–Rpt5; and Rpt1–Rpt2. Within these modules there are four assembly chaperones. Although unrelated phylogenetically, these ‘RP chaperones’ each bind a CP-proximal C domain within a specific Rpt subunit5,9, as follows: Rpt1–Hsm3; Rpt3–Nas6; Rpt5–Nas2; and Rpt6–Rpn14.
A debated aspect of RP assembly is whether the CP facilitates the process15,16, although CP-dependent and CP-independent assembly pathways are not mutually exclusive. We have proposed that the insertion of Rpt tails into CP α-pockets is important for RP assembly in yeast, and that RP chaperones antagonize Rpt tail insertion into CP α-pockets by steric hindrance, thus promoting temporal order in assembly5,6 (Supplementary Fig. 1).
The base is considered a key assembly intermediate of yeast proteasomes5,6,9,10. To investigate the effects of RP chaperones on the RP–CP interface, and to model the behaviour of early assembly intermediates, we developed a reconstitution assay for the base–CP complex. When purified base and CP were mixed, they formed complexes (base1–CP and base2–CP; collectively termed base–CP) with an apparent dissociation constant (Kd) of ∼3 nM (Supplementary Fig. 2). Base–CP was visualized on native polyacrylamide gel electrophoresis (PAGE) by using an in-gel assay for hydrolysis of the fluorogenic peptide LLVY-AMC (Fig. 1a). When chaperones Hsm3, Nas6 and Rpn14 were added in excess at time zero, the chaperone trio inhibited complex formation beyond the detection limit. Because tail–pocket contacts mediate base–CP association12, this experiment satisfies a key prediction of the model5,6—that RP chaperones antagonize insertion of Rpt tails into CP α-pockets. A fourth RP chaperone, Nas2, dissociates before base assembly17 and is accordingly inactive in these assays (data not shown).
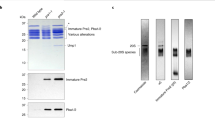
a, Purified base (160 nM) and CP (80 nM) were incubated with or without Rpn14, Nas6 and Hsm3 (trio, 1.6 μM each), and resolved by native PAGE. Top, in-gel peptidase assay (0.02% SDS); bottom, Coomassie stain. For input protein see Supplementary Fig. 2. b, Base (5 nM) and CP (2 nM) were challenged with chaperone trio (amounts in molar excess of base; ATP at 2 mM). In this and all real-time experiments, LLVY-AMC hydrolysis is expressed as relative fluorescence units (r.f.u.) and experiments were performed in triplicate with traces combined for presentation. c, Native gel analysis of base–CP formation as in a, after addition of chaperones to base (160 nM) singly or in combination at tenfold molar excess of base. d, A yeast Rpt hexamer model was built, using the hexameric P97 D1 domain structure as a template (see Supplementary Methods). This model was fit into the EM map2 of yeast Rpt hexamer. Relative positions of Hsm3 (red) and Nas6 (yellow) on the Rpt ring (blue) were assessed by superimposing Hsm3–Rpt1C and Nas6–Rpt3C structures onto the Rpt ring model that had been fit into the EM map. A clipped view of the Rpt ring with bound chaperones and CP (green) is presented. Areas of overlap highlight steric clashes between chaperones and CP.
Antagonism of base–CP association, assessed as described earlier with chaperones in excess, remained strong when chaperones were added at 1:1 stoichiometry versus base, indicating potent interference (Supplementary Fig. 3). Base–CP association can be quantified by real-time fluorometric assays that track LLVY-AMC hydrolysis. Free CP hydrolyses LLVY-AMC slowly, owing to closure of its gated channel1; binding of base opens the channel via Rpt tail–α-pocket interactions12,13. Suppression of base–CP assembly by the chaperone trio is readily observed with the LLVY-AMC assay (Fig. 1b) at low-nanomolar levels, which are comparable to or below their estimated intracellular abundance (ref. 18 and data not shown). Each of these chaperones was found to antagonize the base–CP association individually, although with different potencies (Fig. 1c and Supplementary Fig. 4). In summary, Hsm3, Nas6 and Rpn14 act coordinately, and through a common mechanism, to antagonize the Rpt ring–CP association.
To assess further how RP chaperones regulate proteasomes, we determined the crystal structures of Hsm3 and of Hsm3 complexed with the Rpt1 C domain19,20 (Protein Data Bank accessions 4FP7 and 4JPO; Supplementary Figs 5–9), and modelled this complex into the cryo-electron microscopy (cryoEM) structure of the yeast proteasome holoenzyme2. The Nas6–Rpt3 co-complex21 was similarly modelled into the holoenzyme. The results suggest physical clashing between CP and chaperones in the holoenzyme (Fig. 1d), consistent with the steric interference hypothesis5,6. Although modelling is not completely predictive, these data agree with previous attempts to model chaperones into the holoenzyme5,20. The Rpn14–Rpt6 structure is unsolved and thus the relevance of steric interference to Rpn14 remains conjectural. The RP–CP interface is likely to be dynamic, owing to conformational changes in the Rpt ring during cycles of ATP hydrolysis (see later). Consequently, steric interference may apply to a subset of conformational states.
In the base–CP experimental model, RP chaperones may act by binding to free base, thus preventing association of base with CP. Alternatively, or in addition, chaperones might interact transiently with base–CP to promote dissociation actively. To assess these models, we assayed the time course of base–CP dissociation after chaperone addition. As a control, we examined spontaneous dissociation of base–CP using a ‘CP trap’ that is inactivated by the proteasome inhibitor epoxomicin. The trap captures base that had dissociated from base–CP (Supplementary Fig. 10), thus suppressing LLVY-AMC hydrolysis. Trap addition resulted in slow loss of LLVY-AMC hydrolytic activity over more than 15 min, whereas chaperone addition led to an immediate reduction in hydrolytic activity, as expected from active dissociation (Fig. 2a, left). On chaperone addition, a new, stable steady-state hydrolytic rate was established within approximately 7–8 min (Fig. 2a, right). Chaperone addition to holoenzyme had only weak dissociative effects (Supplementary Fig. 11a). Thus, RP chaperones may work preferentially on assembly intermediates.
a, CP (2 nM) activity stimulated by base (5 nM) was monitored over time (2 mM ATP, 50 mM KCl). At 5.5 min, chaperone trio or CP trap was added in molar excess of base or active CP, respectively. CP trap inhibits re-association of base with active CP. Right, hydrolysis rate (r.f.u. min−1) over time. b, Purified base (5 nM) and CP (2 nM) were assembled in the presence of ATPγS (0.1 mM throughout). At 6 min, chaperone trio or CP trap were added in molar excess. c, CP (15 nM) was immobilized on IgG resin via ProA tag, and incubated with base (∼80 nM) and chaperone trio (160 nM) in the presence of 2 mM ATP or 0.5 mM ATPγS. CP-bound proteins were washed with buffer (50 mM KCl), then eluted with TEV protease while maintaining nucleotide concentration. Immunoblots (IBs) were probed with indicated antibodies. Images are from the same gel and exposure. d, CP (2 nM) activity (LLVY-AMC hydrolysis; r.f.u.) was monitored in the presence of base (5 nM) and 0.1 mM ATPγS for 5 min. Chaperone trio (50 nM) or buffer alone containing 0.1 mM ATPγS was then added. At 10 min, buffer containing either ATPγS or ATP plus ATPγS was added. Final nucleotide concentrations were either 0.1 mM ATPγS or 10 mM ATP plus 0.1 mM ATPγS. See also Supplementary Fig. 12.
To assess the possible relevance of Rpt ring conformational dynamics to base–CP dissociation, we compared the effects of ATP and non-hydrolysable ATPγS in the base–CP dissociation assay. Base–CP assembled normally with ATPγS (Supplementary Fig. 11b), but subsequent chaperone addition had no detectable effect on LLVY-AMC hydrolysis, indicating failure to dissociate the complex (Fig. 2b). Antagonism of base–CP association by chaperones may therefore be finely tuned to the conformational state of the Rpt ring. The simplest interpretation is that ATPγS mimics the ATP-bound state of the Rpt, with a chaperone inhibiting base–CP association when its cognate Rpt is bound to ADP (or is free of nucleotide), but not to ATP. With ATPγS, RP chaperones could fail to antagonize base–CP simply because they cannot bind Rpt proteins under these conditions. However, chaperone–base complexes form comparably with ATP, ATPγS or ADP (Supplementary Fig. 11c).
The results described earlier suggest that a stable base–chaperone–CP co-complex may form in the presence of ATPγS. To test this, we immobilized CP to a resin, added base and chaperones, and assayed resin-bound components after washing. When chaperones were added in the presence of ATP, base dissociated from the complex (Fig. 2c), whereas, with ATPγS, base remained bound to both CP and chaperones. Thus, the chaperones’ lack of effect on LLVY-AMC hydrolysis in the presence of ATPγS reflects their failure to dissociate base–CP, and the chaperones’ capacity to compete with CP for occupancy of the base may be dependent on the Rpt nucleotide hydrolytic cycle.
To test whether preformed base–chaperone–CP complex is primed for dissociation, the complex was formed in the presence of ATPγS, and dissociation was then monitored after ATP addition. ATP produced rapid base–CP dissociation; LLVY-AMC hydrolysing activity decayed with a half-life of approximately 1 min (Fig. 2d and Supplementary Fig. 12). By contrast, ATP and ATPγS produced indistinguishable hydrolytic profiles in the absence of chaperones (Fig. 2d). Thus, RP chaperones actively dissociate base–CP.
To understand chaperone action within the base–CP complex better, we studied the specificity of the insertion of Rpt tails into α-pockets, interactions proposed to be under chaperone control5,6. We previously determined by single-particle cryoEM that the C termini of the homohexameric archaeal PAN ATPases bind the α-pockets of the homoheptameric α-ring of archaeal CP13. This approach is used here to assign distinct α-pocket binding preferences to each yeast Rpt tail. We determined seven subnanometre-resolution three-dimensional reconstructions of the yeast CP, one from CP alone (Supplementary Figs 13 and 14), and the others from CP incubated individually with six different Rpt peptides. Each peptide comprised 8 amino acids from the C terminus of an Rpt.
The pseudo seven-fold symmetry of the heteroheptameric α- and β-rings poses a challenge to single-particle cryoEM. To break this pseudo-symmetry, we fused a glutathione S-transferase (GST) tag to the C terminus of subunit β2 (Supplementary Fig. 13a, e), and verified that the tag does not alter CP function. Differences between maps of peptide–CP and CP alone were calculated as described13 (Supplementary Fig. 15). At proper thresholds, difference densities correspond to peptides bound to α-pockets. Because the pocket between subunits α7 and α1 lacks the Lys residue that is required for binding the C terminus of an Rpt11, no specific binding of any peptide to this pocket is expected. Thresholds were therefore set to show no difference density in the α7/α1-pocket, which in fact always had the lowest difference density corresponding to tail peptide (Fig. 3). As controls, difference maps between two independent three-dimensional reconstructions from two separately collected data sets of the same sample showed no significant difference density, indicating that difference densities assigned to each peptide were not generated by image misalignment or random noise (Supplementary Methods).
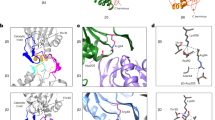
a, Top views of three-dimensional density maps of CP superimposed with difference densities corresponding to C-terminal peptides of each Rpt. Peptides were present at 0.5 mM, CP at 1.6 μM. The amount of each peptide bound is reflected by the size of black densities within each pocket. b, Summary of Rpt tail peptide-binding sites and relative intensities. Sizes of circles represent the volume of difference densities generated by peptides. Grey diagonals denote Rpt tail–α-pocket mapping of intact proteasomes by crosslinking22. Rpt4, Rpt5 and Rpt1 each crosslink to two α-pockets, suggesting an ambiguous register. c, Predominant tail–pocket interactions in yeast holoenzymes as determined by cryoEM2,3,4.
Figure 3a compares binding specificities of the six Rpt peptides for the seven α-pockets of the CP. In an individual difference map, the sizes of the densities in different pockets correlate with peptide-binding affinities for these pockets. Figure 3b summarizes these data. Figure 3b and c also represent previous mapping of Rpt tails to α-pockets in the holoenzyme2,3,4,22. Our data reveal an unexpected lack of specificity in the Rpt tail–α-pocket interaction. We therefore suggest that the specificity of tail–pocket interactions within the mature complex is largely guided by constraints on possible tail–pocket interactions that arise from the defined subunit arrangements of the apposed Rpt and α-rings. For example, within the holoenzyme, the Rpt2 and Rpt3 C termini insert into α3/α4- and α1/α2-pockets, respectively2,3,4,22. However, free forms of Rpt2 and Rpt3 C termini show a marked preference for noncognate pockets (Fig. 3b).
Our results raise the question of how the register of the RP–CP interface is determined. Only the Rpt6 peptide showed high specificity of binding to an established22 cognate pocket (Fig. 3b, c). Thus, the Rpt6 tail has the specificity to serve as an anchor point for either the mature proteasome or an assembly intermediate.
The unique binding specificity of the free Rpt6 tail peptide led us to examine its physiological significance genetically. We generated substitution mutations in Rpt6 tail residues, and assessed effects on proteasome assembly using assays of cell growth (Fig. 4a) and native PAGE (Fig. 4b). Several substitutions resulted in proteasome defects, notably a block substitution of alanines for the terminal LFK sequence (Fig. 4a, b and Supplementary Fig. 16). Deletion of one residue from the C terminus had a similar effect (Fig. 4a, b and Supplementary Figs 16–18).
a, Growth defects of C-terminal rpt6 mutants. Strains were spotted onto plates containing rich media (YPD) in fourfold serial dilutions. Plates were incubated at 30 °C for 2 days. WT, wild type Capital letters indicate the last three amino acids of Rpt6 (LFK) or mutants thereof. b, Whole-cell extracts (100 μg) from rpt6 mutants as in a were resolved by native PAGE and subjected to LLVY-AMC assay in 0.02% SDS. c, Role of Rpt6 tail in base–CP association. Assembly kinetics of wild-type or rpt6-Δ1 base with CP was measured using LLVY-AMC hydrolysis. Purified CP (2 nM) was mixed with the indicated fold excess of base (2 mM ATP). LLVY-AMC hydrolysis is indicated in r.f.u. d, Stability of proteasome holoenzyme (2 nM) from wild-type or rpt6-Δ1 mutants was assessed in 2 mM ATP by adding 50-fold molar excess of CP trap or buffer alone at 15 min.
The exceptional specificity of the free Rpt6 tail for the α2/α3-pocket, together with its role in proteasome assembly, seem to be inconsistent with recent cryoEM studies of the yeast proteasome, which visualize the Rpt2, Rpt3 and Rpt5 tails within α-pockets, with Rpt6 apparently not fixed within an α-pocket, presumably being too flexible to be visualized2,3,4. To resolve this paradox, we first used the base–CP assembly assay to test whether the Rpt6 tail helps to stabilize the base–CP interface. Base complexes were purified from rpt6-Δ1 and wild-type cells, then mixed with CP; the activation of LLVY-AMC hydrolysis was then assayed. At a ratio of 16:1, the rpt6-Δ1 base remained deficient in comparison to a twofold excess of wild-type base over CP (Fig. 4c). The defect in activation can be attributed largely to reduced base–CP association, as shown by native PAGE followed by Coomassie staining (Supplementary Fig. 19).
One scenario to explain the phenotype of rpt6 tail mutants is that their reduced proteasome levels simply reflect a lower affinity between CP and RP. To test this, we purified rpt6-Δ1 proteasome holoenzymes and incubated them in the presence or absence of CP trap to follow their dissociation over time. Mutant and wild-type proteasomes were comparable in stability (Fig. 4d). Thus, the Rpt6 tail influences biosynthetic proteasome assembly rather than holoenzyme stability.
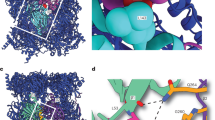
To investigate the base–CP interface more generally, we reconstituted this complex using base from a wild-type (RPT6) strain of yeast, and subjected it to cryoEM (Supplementary Fig. 20). In the three-dimensional reconstruction of base–CP (Fig. 5a), the rotational register between the Rpt ring and the CP, as well as the axial tilt of the Rpt ring from that of CP, are comparable to those in the holoenzyme. Individual Rpt subunits were modelled into the structure as shown, based on previous characterization of the holoenzyme2 (Supplementary Fig. 20c, d).
a, Three-dimensional reconstruction of the singly capped base–CP complex was determined by single-particle cryoEM to a resolution of ∼10 Å. CP subunits are rendered in different colours as indicated. A difference map was calculated between the original three-dimensional reconstruction and one rotated 180° around the two-fold CP symmetry axis. The positive difference density (grey) corresponds to base bound to CP. It shows prominent densities from C termini of Rpt6, Rpt2 and Rpt1, which are clustered on one side of the Rpt ring, bound to specific α-pockets. b, Each panel shows an α-pocket. Arrows indicate densities corresponding to Rpt C termini extending towards the pockets. Asterisks indicate pockets without detectable density from Rpt C termini. Thresholds of CP and base densities are set separately but are identical in all panels. C termini of Rpt6, Rpt2 and Rpt1 are seen to insert into α-pockets.
A key feature of the base–CP complex is the existence of a strong contact between the Rpt6 tail and the α2/α3-pocket (Fig. 5), which is inherently specific for Rpt6 tail peptide. As the holoenzyme does not show a prominent Rpt6 tail contact2,3,4, the structural and functional data both suggest a transient role for the Rpt6 tail in assembly. Distinct contacts were also observed between Rpt1 and the α4/α5-pocket, as well as Rpt2 and the α3/α4-pocket (Fig. 5 and Supplementary Fig. 21), although the Rpt1 contact is relatively superficial. On the basis of the free tail peptide experiments of Fig. 3, the intrinsic specificity of Rpt2 and Rpt1 cannot explain the tail–pocket register of base–CP. This suggests that Rpt6 is a key determinant of the tail–pocket register in base–CP.
The determination of the tail–pocket register is inherently problematic because of the symmetry mismatch between the hexameric Rpt ring and heptameric α-ring, which may underlie a tendency for ambiguous register at this interface3,4,22. Proper register may be achieved through global optimization of tail–pocket interactions, allowing for a subset of incorrect tail–pocket alignments to be rejected despite their being stronger than the correct alignment. However, our data suggest that the tail–pocket register might be largely defined by dominant interactions of high specificity. The Rpt6 tail may perform such a function at an early stage of proteasome maturation, when the RP–CP interface is defined exclusively by Rpt tail–α-pocket contacts. This role of Rpt6 is apparently not sustained in the holoenzyme, perhaps because register is enforced by an alternative mechanism once the lid is incorporated into the complex. Subunit Rpn6 of the lid extends directly past Rpt6 to contact the CP2,23, and may substitute for the anchoring role of Rpt6.
Although the tail–pocket register of base–CP is consistent with that of holoenzyme, the dominant tail–pocket interactions are quite different. For the holoenzyme2,3,4, these are thought to be Rpt3, Rpt2 and Rpt5. These tails alternate across the ring in the holoenzyme (Fig. 3c), whereas in base–CP the dominant tail contacts appear to be collected on one side of the ring (Fig. 5), in an arrangement resembling that of the archaeal PAN complex14.
An interesting feature of base–CP is that the neighbouring17 Rpt6 and Rpt2 tails show strong pocket interactions. Because the tail of Rpt2 has little inherent specificity, its insertion into the α3/α4-pocket may be facilitated by Rpt6. It is consistent with the symmetry mismatch between the Rpt and α-rings that Rpt6 should preferentially promote tail insertion of its nearest neighbour, as more distant tails would fall out of phase with the CP pockets.
Negative regulation of tail–pocket interactions by chaperones may potentially help in temporal ordering of the assembly pathway, in suppressing out-of-register tail–pocket interactions, and in maintaining proteasome assembly intermediates in a highly dynamic state. Our data suggest that chaperone action may be coupled to the ATPase cycle of the Rpt ring, with nucleotide controlling the competition between chaperone and CP for base interaction. The mechanism may involve changes in positioning of the Rpt C domain, which has an integrative role in that it positions the Rpt C-terminal tail, while at the same time binding chaperone on its outer face and contacting nucleotide on its inner face.
The major forms of mature proteasome differ from the ATPγS–base–CP complex in that they are associated with a mixture of ATP and ADP24, whereas early intermediates in RP assembly are reported to have no detectable ATPase activity25, suggesting that an ADP-free species resembling the ATPγS–base–CP complex could potentially function as a transient assembly intermediate. The fate of this complex may be to undergo chaperone-dependent dissociation upon ATP hydrolysis, or alternatively the lid may join the complex before the first round of ATP hydrolysis, to impose new modes of CP binding and suppress complex dissociation.
Methods Summary
The GST-tagged CP used for cryoEM analysis was purified using a 3× Flag tag appended to the Pre1 C terminus (β4). For the structure of CP complexed with peptide, ∼0.5 mM peptide was incubated with 1.6 μM GST-tagged CP for 1 h at 37 °C directly before grid vitrification. Recombinant chaperones were purified from Escherichia coli using a GST tag, which was removed with Prescission protease before biochemical assays. Single-particle cryoEM studies were carried out as described13, with details given in Supplementary Information.
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
Data have been deposited in the Electron Microscopy Data Bank under the following accession numbers: free CP, EMD-5593; Rpt1–CP, EMD-5611; Rpt2–CP, EMD-5612; Rpt3–CP, EMD-5613; Rpt4–CP, EMD-5614; Rpt5–CP, EMD-5615; Rpt6–CP, EMD-5616; and base1–CP: EMD-5617. For the crystal structures, data have been deposited in the Protein Data Bank under accessions 4FP7 (Hsm3) and 4JPO (Hsm3–Rpt1 C domain).
References
Kish-Trier, E. & Hill, C. P. Structural biology of the proteasome. Ann. Rev.. Biophyshttp://dx.doi.org/10.1146/annurev-biophys-083012-130417 (2013)
Lander, G. C. et al. Complete subunit architecture of the proteasome regulatory particle. Nature 482, 186–191 (2012)
Lasker, K. et al. Molecular architecture of the 26S proteasome holocomplex determined by an integrative approach. Proc. Natl Acad. Sci. USA 109, 1380–1387 (2012)
Beck, F. et al. Near-atomic resolution structural model of the yeast 26S proteasome. Proc. Natl Acad. Sci. USA 109, 14870–14875 (2012)
Roelofs, J. et al. Chaperone-mediated pathway of proteasome regulatory particle assembly. Nature 459, 861–865 (2009)
Park, S. et al. Hexameric assembly of the proteasomal ATPases is templated through their C termini. Nature 459, 866–870 (2009)
Le Tallec, B., Barrault, M. B., Guerois, R., Carre, T. & Peyroche, A. Hsm3/S5b participates in the assembly pathway of the 19S regulatory particle of the proteasome. Mol. Cell 33, 389–399 (2009)
Kaneko, T. et al. Assembly pathway of the mammalian proteasome base subcomplex is mediated by multiple specific chaperones. Cell 137, 914–925 (2009)
Saeki, Y., Toh, E. A., Kudo, T., Kawamura, H. & Tanaka, K. Multiple proteasome-interacting proteins assist the assembly of the yeast 19S regulatory particle. Cell 137, 900–913 (2009)
Funakoshi, M., Tomko, R. J., Jr, Kobayashi, H. & Hochstrasser, M. Multiple assembly chaperones govern biogenesis of the proteasome regulatory particle base. Cell 137, 887–899 (2009)
Forster, A., Masters, E. I., Whitby, F. G., Robinson, H. & Hill, C. P. The 1.9 Å structure of a proteasome-11S activator complex and implications for proteasome-PAN/PA700 interactions. Mol. Cell 18, 589–599 (2005)
Smith, D. M. et al. Docking of the proteasomal ATPases’ carboxyl termini in the 20S proteasome’s α ring opens the gate for substrate entry. Mol. Cell 27, 731–744 (2007)
Rabl, J. et al. Mechanism of gate opening in the 20S proteasome by the proteasomal ATPases. Mol. Cell 30, 360–368 (2008)
Smith, D. M. et al. ATP binding to PAN or the 26S ATPases causes association with the 20S proteasome, gate opening, and translocation of unfolded proteins. Mol. Cell 20, 687–698 (2005)
Park, S., Kim, W., Tian, G., Gygi, S. P. & Finley, D. Structural defects in the RP-CP interface induce a novel proteasome stress response. J. Biol. Chem. 286, 36652–36666 (2011)
Kusmierczyk, A. R., Kunjappu, M. J., Funakoshi, M. & Hochstrasser, M. A multimeric assembly factor controls the formation of alternative 20S proteasomes. Nature Struct. Mol. Biol. 15, 237–244 (2008)
Tomko, R. J., Jr, Funakoshi, M., Schneider, K., Wang, J. & Hochstrasser, M. Heterohexameric ring arrangement of the eukaryotic proteasomal ATPases: implications for proteasome structure and assembly. Mol. Cell 38, 393–403 (2010)
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003)
Takagi, K. et al. Structural basis for specific recognition of Rpt1, an ATPase subunit of 26S proteasome, by proteasome-dedicated chaperone Hsm3p. J. Biol. Chem. 287, 12172–12182 (2012)
Barrault, M. B. et al. Dual functions of the Hsm3 protein in chaperoning and scaffolding regulatory particle subunits during the proteasome assembly. Proc. Natl Acad. Sci. USA 109, E1001–E1010 (2012)
Nakamura, Y. et al. Structural basis for the recognition between the regulatory particles Nas6 and Rpt3 of the yeast 26S proteasome. Biochem. Biophys. Res. Commun. 359, 503–509 (2007)
Tian, G. et al. An asymmetric interface between the regulatory particle and core particle of the proteasome. Nature Struct. Mol. Biol. 18, 1259–1267 (2011)
Pathare, G. R. et al. The proteasomal subunit Rpn6 is a molecular clamp holding the core and regulatory subcomplexes together. Proc. Natl Acad. Sci. USA 109, 149–154 (2012)
Smith, D. M., Fraga, H., Reis, C., Kafri, G. & Goldberg, A. L. ATP binds to proteasomal ATPases in pairs with distinct functional effects, implying an ordered reaction cycle. Cell 144, 526–538 (2011)
Thompson, D., Hakala, K. & DeMartino, G. N. Subcomplexes of PA700, the 19S regulator of the 26S proteasome, reveal relative roles of AAA subunits in 26S proteasome assembly and activation and ATPase activity. J. Biol. Chem. 284, 24891–24903 (2009)
Acknowledgements
We thank M. Schmidt, T. Walz, C. Chen, and Finley laboratory members for suggestions, and C. Mann for antibodies. This work was supported in part by grants from the National Institutes of Health (NIH; R01GM082893 and 1S10RR026814-01), the University of California San Francisco Program for Breakthrough Biomedical Research (New Technology Award) to Y.C.; the Johnson Cancer Research Center, the National Center for Research Resources (5P20RR017708 and P20 RR016475) and NIH (8 P20 GM103420 and P20 GM103418) to J.R.; and grants from the NIH to P.C. (R01GM045335) and D.F. (R37GM043601). S.P. was supported by the Charles A. King Trust Postdoctoral Research Fellowship Program of the Medical Foundation. Use of IMCA-CAT was supported by the Industrial Macromolecular Crystallography Association though a contract with the Hauptman-Woodward MRI. Use of the Advanced Photon Source was supported by the US Department of Energy (contract no. DE-AC02-06CH11357).
Author information
Authors and Affiliations
Contributions
S.P. performed reconstitution of the base–CP complex and holoenzyme stability. X.L. performed all cryoEM experiments and analysis. H.M.K. and C.R.S. generated yeast strains. H.M.K. purified GST-fused CP, and participated in cryoEM experiments and analysis. C.R.S. performed purifications, and M.Z. performed ultracentrifugation. K.P.B. and S.L. determined crystal structures, J.R. and G.T. performed structural analysis and modelling. M.A.H., H.M.K. and P.C. performed phenotypic and native gel analysis of Rpt6 mutations. J.R. wrote the supplement with contributions from all authors. The manuscript was drafted by D.F. and Y.C., and modified by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-21, Supplementary Tables 1-3, Supplementary Methods and Supplementary References. (PDF 12079 kb)
Rights and permissions
About this article
Cite this article
Park, S., Li, X., Kim, H. et al. Reconfiguration of the proteasome during chaperone-mediated assembly. Nature 497, 512–516 (2013). https://doi.org/10.1038/nature12123
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12123
This article is cited by
-
Proteasome regulation by reversible tyrosine phosphorylation at the membrane
Oncogene (2021)
-
Conformational maps of human 20S proteasomes reveal PA28- and immuno-dependent inter-ring crosstalks
Nature Communications (2020)
-
Regulation of proteasome assembly and activity in health and disease
Nature Reviews Molecular Cell Biology (2018)
-
Delineation of molecular pathway activities of the chronic antidepressant treatment response suggests important roles for glutamatergic and ubiquitin–proteasome systems
Translational Psychiatry (2017)
-
An atomic structure of the human 26S proteasome
Nature Structural & Molecular Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.