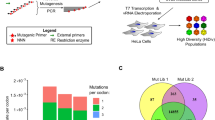
Abstract
RNA viruses exist as genetically diverse populations1. It is thought that diversity and genetic structure of viral populations determine the rapid adaptation observed in RNA viruses2 and hence their pathogenesis3. However, our understanding of the mechanisms underlying virus evolution has been limited by the inability to accurately describe the genetic structure of virus populations. Next-generation sequencing technologies generate data of sufficient depth to characterize virus populations, but are limited in their utility because most variants are present at very low frequencies and are thus indistinguishable from next-generation sequencing errors. Here we present an approach that reduces next-generation sequencing errors and allows the description of virus populations with unprecedented accuracy. Using this approach, we define the mutation rates of poliovirus and uncover the mutation landscape of the population. Furthermore, by monitoring changes in variant frequencies on serially passaged populations, we determined fitness values for thousands of mutations across the viral genome. Mapping of these fitness values onto three-dimensional structures of viral proteins offers a powerful approach for exploring structure–function relationships and potentially uncovering new functions. To our knowledge, our study provides the first single-nucleotide fitness landscape of an evolving RNA virus and establishes a general experimental platform for studying the genetic changes underlying the evolution of virus populations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Accessions
Sequence Read Archive
Data deposits
Sequencing data has been deposited in the NCBI Sequence Read Archive under accession number PRJNA222998. Software complementary to this analysis is available at http://andino.ucsf.edu.
References
Domingo, E., Sabo, D., Taniguchi, T. & Weissmann, C. Nucleotide sequence heterogeneity of an RNA phage population. Cell 13, 735–744 (1978)
Burch, C. L. & Chao, L. Evolvability of an RNA virus is determined by its mutational neighbourhood. Nature 406, 625–628 (2000)
Vignuzzi, M., Stone, J. K., Arnold, J. J., Cameron, C. E. & Andino, R. Quasispecies diversity determines pathogenesis through cooperative interactions in a viral population. Nature 439, 344–348 (2006)
Lou, D. I. et al. High-throughput DNA sequencing errors are reduced by orders of magnitude using circle sequencing. Proc. Natl Acad. Sci. USA (in the press)
Sanjuán, R., Nebot, M. R., Chirico, N., Mansky, L. M. & Belshaw, R. Viral mutation rates. J. Virol. 84, 9733–9748 (2010)
Crotty, S., Cameron, C. E. & Andino, R. RNA virus error catastrophe: direct molecular test by using ribavirin. Proc. Natl Acad. Sci. USA 98, 6895–6900 (2001)
Wakeley, J. The excess of transitions among nucleotide substitutions: new methods of estimating transition bias underscore its significance. Trends Ecol. Evol. 11, 158–162 (1996)
Dohm, J. C., Lottaz, C., Borodina, T. & Himmelbauer, H. Substantial biases in ultra-short read data sets from high-throughput DNA sequencing. Nucleic Acids Res. 36, e105 (2008)
Freistadt, M. S., Vaccaro, J. A. & Eberle, K. E. Biochemical characterization of the fidelity of poliovirus RNA-dependent RNA polymerase. Virol. J. 4, 44 (2007)
Arnold, J. J. & Cameron, C. E. Poliovirus RNA-dependent RNA polymerase (3Dpol): pre-steady-state kinetic analysis of ribonucleotide incorporation in the presence of Mg2+. Biochemistry 43, 5126–5137 (2004)
Radford, A. D. et al. Application of next-generation sequencing technologies in virology. J. Gen. Virol. 93, 1853–1868 (2012)
Orr, H. A. The rate of adaptation in asexuals. Genetics 155, 961–968 (2000)
Kimura, M. The Neutral Theory of Molecular Evolution 55–97 (Cambridge Univ. Press, 1983)
Cuevas, J. M., González-Candelas, F., Moya, A. & Sanjuán, R. Effect of ribavirin on the mutation rate and spectrum of hepatitis C virus in vivo. J. Virol. 83, 5760–5764 (2009)
Hämmerle, T., Hellen, C. U. & Wimmer, E. Site-directed mutagenesis of the putative catalytic triad of poliovirus 3C proteinase. J. Biol. Chem. 266, 5412–5416 (1991)
Hellen, C. U. T., Lee, C.-K. & Wimmer, E. Determinants of substrate recognition by poliovirus 2A proteinase. J. Virol. 66, 3330–3338 (1992)
Gohara, D. W. et al. Poliovirus RNA-dependent RNA polymerase (3Dpol): structural, biochemical, and biological analysis of conserved structural motifs A and B. J. Biol. Chem. 275, 25523–25532 (2000)
Gould, S. J. Dollo on Dollo’s law: irreversibility and the status of evolutionary laws. J. Hist. Biol. 3, 189–212 (1970)
Haldane, J. B. S. A mathematical theory of natural and artificial selection, part V: selection and mutation. Math. Proc. Camb. Philos. Soc. 23, 838–844 (1927)
Cuevas, J. M., Domingo-Calap, P. & Sanjuán, R. The fitness effects of synonymous mutations in DNA and RNA viruses. Mol. Biol. Evol. 29, 17–20 (2012)
Eyre-Walker, A. & Keightley, P. D. The distribution of fitness effects of new mutations. Nature Rev. Genet. 8, 610–618 (2007)
Sanjuán, R., Moya, A. & Elena, S. F. The distribution of fitness effects caused by single-nucleotide substitutions in an RNA virus. Proc. Natl Acad. Sci. USA 101, 8396–8401 (2004)
Chao, L. Fitness of RNA virus decreased by Muller’s ratchet. Nature 348, 454–455 (1990)
Mueller, S., Papamichail, D., Coleman, J. R., Skiena, S. & Wimmer, E. Reduction of the rate of poliovirus protein synthesis through large-scale codon deoptimization causes attenuation of viral virulence by lowering specific infectivity. J. Virol. 80, 9687–9696 (2006)
Coleman, J. R. et al. Virus attenuation by genome-scale changes in codon pair bias. Science 320, 1784–1787 (2008)
Tokuriki, N. & Tawfik, D. Protein dynamism and evolvability. Science 324, 203–207 (2009)
Jäger, S. et al. Global landscape of HIV–human protein complexes. Nature 481, 365–370 (2012)
Gong, P. & Peersen, O. B. Structural basis for active site closure by the poliovirus RNA-dependent RNA polymerase. Proc. Natl Acad. Sci. USA 107, 22505–22510 (2010)
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359 (2012)
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Herold, J. & Andino, R. Poliovirus requires a precise 5′ end for efficient positive-strand RNA synthesis. J. Virol. 74, 6394–6400 (2000)
Draper, N. R. & Smith, H. Applied Regression Analysis (Wiley, 1998)
Gelman, A., Carlin, J. B., Stern, H. S. & Rubin, D. B. Bayesian Data Analysis (Chapman & Hall/CRC Texts in Statistical Science, 2003)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995)
Lama, J., Sanz, M. A. & Rodríguez, P. L. A role for 3AB protein in poliovirus genome replication. J. Biol. Chem. 270, 14430–14438 (1995)
Lama, J., Sanz, M. A. & Carrasco, L. Genetic analysis of poliovirus protein 3A: characterization of a non-cytopathic mutant virus defective in killing Vero cells. J. Gen. Virol. 79, 1911–1921 (1998)
Dewalt, P. G., Blair, W. S. & Semler, B. L. A genetic locus in mutant poliovirus genomes involved in overproduction of RNA polymerase and 3C proteinase. Virology 174, 504–514 (1990)
Blair, W. S., Nguyen, J. H. C., Parsley, T. B. & Semler, B. L. Mutations in the poliovirus 3CD proteinase S1-specificity pocket affect substrate recognition and RNA binding. Virology 218, 1–13 (1996)
Hobson, S. D. et al. Oligomeric structures of poliovirus polymerase are important for function. EMBO J. 20, 1153–1163 (2001)
Acknowledgements
We thank J. Frydman, S. Bianco, H. Dawes, K. Ehmsen and members of the Andino laboratory for critical reading of the manuscript and G. Schroth, M. Harrison, P. Wassam and T. Collins for technical advice. This work was financially supported by a National Science Foundation graduate research fellowship to A.A., NIAID AI091575, AI36178 and AI40085 to R.A., and DARPA Prophecy to R.A. and L.B.
Author information
Authors and Affiliations
Contributions
R.A. and A.A. conceived and designed the experiments. A.A. performed experiments and sequencing. A.A. and L.B. analysed the data and performed statistical analyses. R.A. and A.A. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
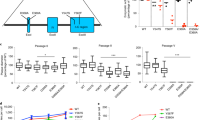
Extended Data Figure 1 CirSeq library preparation scheme.
As described in Methods, purified populations of ssRNA viral RNA genomes are converted by a series of molecular cloning steps to a library compatible with Illumina sequencing. Illumina paired-end Y-adaptors are represented in blue.
Extended Data Figure 2 Mutation frequencies of transitions and transversions.
Because transitions (Ts) and transversion (Tv) occur at different rates, the overall frequencies of these types of mutations stabilize at different levels. The lower the mutation frequency, the longer it takes to stabilize, because smaller quantities of error can more dramatically impact their measured frequency. An important consideration for CirSeq is at what quality score to threshold data in order to minimize the contribution of error in the final output and maximize the total quantity of the data used.
Extended Data Figure 3 Genome coverage per base.
a, Coverage for sequenced passages. The coverage for each base for each library above the minimum quality threshold of average Q20 was mapped. On average, we obtained 204,205-fold coverage for our populations. The coverage profile is extremely consistent between libraries and experiments. b, Effect of RNA fragment size on coverage bias. Use of fragments less than 80–90 bases in length results in over-representation of A-rich sequences. This bias is likely the result of inefficient priming of certain short templates by reverse transcriptase. Fragments should be at least 80–90 bases, which limits coverage bias to within approximately 10×, typical of RNA-seq.
Extended Data Figure 4 Frequency measurement error.
a, b, Error in measurement of mutation frequencies is determined by coverage depth and mutation frequency. A library prepared from 30 base fragments, which increases variability in the level of coverage (see Extended Data Fig. 3b) over different regions of the poliovirus genome, was broken into 10 million read sets (sets 1 and 2). The frequency of each variant for the two sets was mapped against each other to visualize their correlation. a, Measurement error can be estimated as the standard error of a binomial distribution. Per cent error is obtained by dividing this standard error by the variant frequency. Low measurement error corresponds to high correlation between variant frequencies measured in each set. b, Correlation between measured variant frequencies also corresponds to coverage, where greater coverage increases correlation. The amount of coverage required to obtain good correlation between measurements scales with variant frequency. c, Amplification bias. The distribution of frequencies of nonsense mutations generated by C > U mutation are shown for passages 2 and 3. In each case, frequencies are tightly distributed around the mean, ruling out PCR amplification bias in contributing substantially to measurement error of variant frequencies.
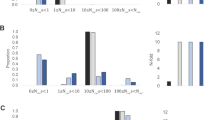
Extended Data Figure 5 Inferred population structure and selection over seven passages.
a, Simulation of population structure from sequencing data. The histograms display the proportion of genomes at each passage containing the given number of mutations (Hamming distance from the reference) after removing genomes containing lethal mutations from the population. The proportion of genomes containing single point mutations is relatively constant throughout the passages whereas the proportions of wild-type and multi-variant genomes decrease and increase, respectively. Theses proportions are based on a simulation where mutations are distributed randomly and all viable mutants have fitness equivalent to wild type. b, Accumulation of mutations by selection. The frequency of mutations accumulated as a result of selection, that is, after removing de novo mutations, is plotted for each passage. Mutations accumulate approximately linearly over the course of the experiment suggesting that selection is constant.
Extended Data Figure 6 Analysis of mutational fitness effects.
a, Spatial distribution of synonymous mutations by fitness effect. Synonymous mutations were binned by the magnitude of their fitness effect and plotted against their respective genome position. Each bin of fitness effects is well distributed across the genome, indicating that synonymous mutations with strong fitness effects map to discrete regions. b, The distributions of mutational fitness effects of synonymous mutations for structural (black) and non-structural (green) genes are similar. c, Summary of mutational fitness effects. Differences in variance are statistically significant between non-synonymous mutations in structural and non-structural genes both including and excluding lethal mutations (P < 0.001, one-sided F-test). Differences in variance are also statistically significant between non-synonymous and synonymous mutations the coding sequence both including and excluding lethal mutations (P < 0.001, one-sided F-test).
Extended Data Figure 7 Number of passages used to calculate fitness affects accuracy.
Fitness for each variant was calculated for varying numbers of serial passages and normalized to the fitness calculated using the full set of seven passages. As the number of passages used to calculate fitness increases, the variation in fitness decreases, indicating that the calculated fitness is more accurate.
Extended Data Figure 8 Simulation of genetic drift and its impact on fitness measurement.
Top row shows one thousand simulations of a mutation-selection-drift process in a population of 106 genomes are shown for mutations initiated at their mutation rate: 10−3 (black), 10−4 (blue), 10−5 (green) and 10−6 (red). Because of the low number of mutations in populations where the mutation rate was set to 10−6, it is common for the population to lose the mutant by drift. As frequency was plotted on a log scale, a frequency of 0 was represented as 10−7. The histograms show fitness calculated using a simple mutation-selection model for each simulation. The standard deviation for each set of calculations is noted in the title of each set of simulations. The stronger drift experienced by low frequency variants reduces the accuracy of fitness measurements. To account for this effect, we have incorporated drift into our fitness model.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Supplementary References. (PDF 330 kb)
Rights and permissions
About this article
Cite this article
Acevedo, A., Brodsky, L. & Andino, R. Mutational and fitness landscapes of an RNA virus revealed through population sequencing. Nature 505, 686–690 (2014). https://doi.org/10.1038/nature12861
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12861
This article is cited by
-
Targeted accurate RNA consensus sequencing (tARC-seq) reveals mechanisms of replication error affecting SARS-CoV-2 divergence
Nature Microbiology (2024)
-
Evolutionary conservation of the fidelity of transcription
Nature Communications (2023)
-
Mutational spectrum of hepatitis C virus in patients with chronic hepatitis C determined by single molecule real-time sequencing
Scientific Reports (2022)
-
Inferring the distribution of fitness effects in patient-sampled and experimental virus populations: two case studies
Heredity (2022)
-
Adaptation to host cell environment during experimental evolution of Zika virus
Communications Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.