Abstract
Disruption of epithelial polarity is a key event in the acquisition of neoplastic growth. JNK signalling is known to play an important part in driving the malignant progression of many epithelial tumours, although the link between loss of polarity and JNK signalling remains elusive. In a Drosophila genome-wide genetic screen designed to identify molecules implicated in neoplastic growth1, we identified grindelwald (grnd), a gene encoding a transmembrane protein with homology to members of the tumour necrosis factor receptor (TNFR) superfamily. Here we show that Grnd mediates the pro-apoptotic functions of Eiger (Egr), the unique Drosophila TNF, and that overexpression of an active form of Grnd lacking the extracellular domain is sufficient to activate JNK signalling in vivo. Grnd also promotes the invasiveness of RasV12/scrib−/− tumours through Egr-dependent Matrix metalloprotease-1 (Mmp1) expression. Grnd localizes to the subapical membrane domain with the cell polarity determinant Crumbs (Crb) and couples Crb-induced loss of polarity with JNK activation and neoplastic growth through physical interaction with Veli (also known as Lin-7). Therefore, Grnd represents the first example of a TNFR that integrates signals from both Egr and apical polarity determinants to induce JNK-dependent cell death or tumour growth.
Similar content being viewed by others
Main
We recently carried out a genome-wide screen to identify molecules that are required for neoplastic growth1. The condition used for this screen was the disc-specific knockdown of avalanche (rotund (rn)>avl-RNAi; avalanche also known as syntaxin 7), a gene encoding a syntaxin that functions in the early step of endocytosis2. rn>avl-RNAi results in ectopic Wingless (Wg) expression, neoplastic disc overgrowth2 (Fig. 1a, b), and a 2-day delay in larva-to-pupa transition1. We screened a collection of 10,100 transgenic RNA interference (RNAi) lines for their ability to rescue the pupariation delay and identified 121 candidate genes1. Interestingly, only eight candidate genes also rescued ectopic Wg expression and neoplastic overgrowth (Extended Data Fig. 1a). These included five lines targeting core components of the JNK pathway (Bendless, Tab2, Tak1, Hemipterous and Basket; Extended Data Fig. 1b). Using a puckered enhancer trap (puc-lacZ) as a readout for JNK activity, we confirmed that JNK signalling is highly upregulated in rn>avl-RNAi discs (Fig. 1a, b, right). One of the remaining lines targets CG10176, a gene encoding a transmembrane protein. Reducing expression of CG10176 by using two different RNAi lines was as efficient as tak1 silencing to restore normal Wg pattern and suppresses JNK signalling and neoplastic growth in the rn>avl-RNAi background (Fig. 1c, d and Extended Data Fig. 1c–e). Sequence analysis of GC10176 identified a cysteine-rich domain (CRD) in the extracellular part with homology to vertebrate TNFRs (Fig. 1e and Extended Data Fig. 2) harbouring a glycosphingolipid-binding motif (GBM) characteristic of many TNFRs including Fas3 (Fig. 1e and Extended Data Fig. 2). We named CG10176 grindelwald (grnd), after a village at the foot of Eiger, a Swiss mountain that lent its name to the unique Drosophila TNF, Egr. Immunostaining and subcellular fractionation of disc extracts confirmed that Grnd localizes to the membrane (Extended Data Fig. 1f–h). Moreover, co-immunoprecipitation experiments showed that both Grnd full-length and Grnd-intra, a form lacking its extracellular domain, directly associate with Traf2, the most upstream component of the JNK pathway (Fig. 1f–h). This interaction is disrupted by a single amino acid substitution within a conserved Traf6-binding motif4 (human TRAF6 is the closest homologue to Traf2; Fig. 1f–h and Extended Data Fig. 3). Overexpression of Grnd-intra, but not full-length Grnd, is sufficient to induce JNK signalling, ectopic Wg expression and apoptosis (Fig. 1i–j and Extended Data Fig. 4a–c), and Grnd-intra-induced apoptosis is efficiently suppressed in a hep75 (JNKK) mutant background (Extended Data Fig. 4d, e), confirming that Grnd acts upstream of the JNK signalling cascade.
a–d, Wing discs stained for Wg (left) or LacZ (right). e, Sequence alignment of Grnd and human TNFRs. Blue, identity; yellow, strong similarity to Grnd; grey, weak similarity to Grnd. f, g, Co-immunoprecipitation experiments from S2R+ cells co-expressing haemagglutinin (HA)–Traf2 or Ihog–HA (control), and either Grnd–Flag, Grnd-intra–Flag or Grnd(E202A)–Flag (f), or HA–Egr and either Grnd–Flag, Grnd(E202A)–Flag or Ihog–Flag (control) (g). Input loading is quantified using β-tubulin. h, Pull-down experiments of Traf2 with various truncated forms of glutathione S-transferase (GST)–Grnd. Commassie blue staining is used to visualize the bound species. WT, wild type. i–j, Wing discs stained for Wg (left), LacZ (middle) and Grnd (right).
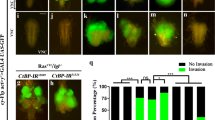
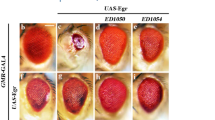
The Drosophila TNF Egr activates JNK signalling and triggers cell death or proliferation, depending on the cellular context5. We therefore tested whether Grnd is required for the small-eye phenotype generated by Egr-induced apoptosis in the retinal epithelium6,7 (GMR>egr; Fig. 2a, b). As previously shown, inhibition of JNK signalling by reducing tak1 (ref. 6) or traf2 (ref. 8) expression, or by overexpressing puckered7, blocks Egr-induced apoptosis and rescues the small-eye phenotype (Extended Data Fig. 5a–d). In contrast to a previous report9, RNAi silencing of wengen (wgn), a gene encoding a presumptive receptor for Egr, does not rescue the small-eye phenotype (Extended Data Fig. 5e). Furthermore, the small-eye phenotype is not modified in a wgn-null mutant background (Fig. 2c and Extended Data Fig. 5h, m, n), confirming that Wgn is not required for Egr-induced apoptosis in the eye. By contrast, reducing grnd levels partially rescues the Egr-induced small-eye phenotype, producing a ‘hanging-eye’ phenotype (Fig. 2d) that is not further rescued in a wgn-knockout (wgnKO ) mutant background (Extended Data Fig. 5e–i). A similar phenotype was previously reported as a result of non-autonomous cell death induced by a diffusible form of Egr10 (Extended Data Fig. 5k, l). This suggests that Grnd prevents Egr from diffusing outside of its expression domain. Co-immunoprecipitation experiments show that both full-length Grnd and Grnd-extra, a truncated form of Grnd lacking the cytoplasmic domain, associate with Egr through its TNF-homology domain (Fig. 2j, k). Although Grnd-extra can bind Egr, it cannot activate JNK signalling. Therefore, we reasoned that Grnd-extra expression might prevent both cell-autonomous and non-autonomous apoptosis by trapping Egr and preventing its diffusion and binding to endogenous Grnd. Indeed, GMR-Gal4-mediated expression of grnd-extra fully rescues the Egr small-eye phenotype (Fig. 2e and Extended Data Fig. 5j). To confirm that the removal of Grnd induces Egr-mediated non-autonomous cell death, we generated wing disc clones expressing egr alone, egr + tak1 RNAi, or egr + grnd RNAi (Fig. 2f–i). As expected, reducing tak1 levels in egr-expressing clones prevents their elimination by apoptosis (Fig. 2f, g). Similarly, reducing grnd levels prevents autonomous cell death, but also induces non-autonomous apoptosis (Fig. 2h, i). This suggests that Egr, like its mammalian counterpart TNF-α, can be processed into a diffusible form in vivo whose interaction with Grnd limits the potential to act at a distance. Flies carrying homozygous (grndMinos/Minos ) or transheterozygous (grndMinos/Df ) combinations of a transposon inserted in the grnd locus express no detectable levels of Grnd protein (Extended Data Fig. 6a, b) and are equally resistant to Egr-induced cell death (Extended Data Fig. 6e–j). In addition, grndMinos/Minos mutant flies are viable and display no obvious phenotype (Extended Data Fig. 6c, d), suggesting that Grnd, like Egr, participates in a stress response to limit organismal damage. Collectively, our data demonstrate that Grnd is a new Drosophila TNF receptor that mediates most, if not all, Egr-induced apoptosis.
a–e, Light micrographs of Drosophila adult eyes of the indicated genotypes (all animals carry the GMR-Gal4 driver). Original magnification, ×30. f–i, Wing discs carrying green fluorescent protein (GFP)-labelled clones (green) expressing egr (f), egr + tak1 RNAi (g) or egr + grnd RNAi (h, i) and stained for Caspase 3 (Cas3; red). A close-up shows non-autonomous cell death around the egr + grnd RNAi clones. j, k, Co-immunoprecipitation (IP) experiments from S2R+ cells expressing HA–Egr or HA–EgrΔTNF (Egr lacking its TNF-homology domain; amino acids 278–415) and either Grnd–Flag, Grnd-extra–Flag or Ihog–Flag (control). Input loading is quantified using β-tubulin.
TNFs probably represent a danger signal produced in response to tissue damage to rid the organism of premalignant tissue or to facilitate wound healing. Disc clones mutant for the polarity gene scribbled (scrib) induce an Egr-dependent response resulting in the elimination of scrib mutant cells by JNK-mediated apoptosis11,12. To test the requirement for Grnd in this process, we compared scrib-RNAi and scrib-RNAi + grnd-RNAi clones obtained 72 h after heat shock induction. As expected, scrib-RNAi cells undergo apoptosis and detach from the epithelium (Fig. 3a–d). By contrast, scrib-RNAi clones with reduced grnd expression survive (Fig. 3e, f), indicating that Grnd is required for Egr-dependent elimination of scrib-RNAi cells. Similar results were obtained by generating scrib−/− mutant clones in the eye disc (Extended Data Fig. 7a–d).
a–f, Wing discs dissected 5 days after egg deposition (AED) stained for Cas3 (red) and carrying GFP-labelled clones (green) of the indicated genotypes induced 2 days AED. a, b, e, Wild-type clones (a) and clones expressing the control Rh3 RNAi (b) to exclude a Gal4 titration effect (e). g–i, Eye–brain complexes dissected 7 days AED carrying RasV12/scrib−/− (g) or RasV12/scrib−/− + grnd RNAi (h) clones (labelled by GFP), or RasV12/scrib−/− GFP-labelled clones in an egr mutant background (i) stained for Mmp1 (g–i, middle, red) and 4′,6-diamidino-2-phenylindole (DAPI; blue). RasV12/scrib−/− clones invade the ventral nerve cord (g, white arrow). c.c., cephalic complex.
In both mammals and flies, TNFs are double-edged swords that also have the capacity to promote tumorigenesis in specific cellular contexts5. Indeed, scrib−/− eye disc cells expressing an activated form of Ras (RasV12 ) exhibit a dramatic tumour-like overgrowth and metastatic behaviour, a process that critically relies on Egr13. RasV12 /scrib−/− metastatic cells show a strong accumulation of Grnd and Mmp1, and invade the ventral nerve cord11,14,15 (Fig. 3g and Extended Data Figs 7e, 8a). Primary tumour cells reach peripheral tissues such as the fat body and the gut, where they form micro-metastases expressing high levels of Grnd (Extended Data Fig. 7f, g). Reducing grnd levels in RasV12 /scrib−/− clones is sufficient to restore normal levels of Mmp1 and abolish invasiveness in a way similar to that observed in an egr−/− background13 (Fig. 3h, i and Extended Data Fig. 8a–c). Therefore, Grnd is required for the Egr-induced metastatic behaviour of RasV12 /scrib−/− tumorous cells. Similarly, reducing grnd, but not wgn levels, strongly suppresses Mmp1 expression in RasV12 /dlg-RNAi cells and limits tumour invasion (Extended Data Fig. 8d–g), indicating that Wgn does not have a major role in the progression of these tumours.
Perturbation of cell polarity is an early hallmark of tumour progression in epithelial cells. In contrast to small patches of polarity-deficient cells, for example, scrib−/− clones, organ compartments or animals fully composed of polarity-deficient cells become refractory to Egr-induced cell death and develop epithelial tumours. The formation of these tumours requires JNK/MAPK signalling, but not Egr13,16, suggesting Egr-independent coupling between loss of polarity and JNK/MAPK-dependent tumour growth. In line with these observations, we noticed that, in contrast to Grnd, Egr is not required to drive neoplastic growth in rn>avl-RNAi conditions (Extended Data Fig. 9a, b). This suggests that, in addition to its role in promoting Egr-dependent functions, Grnd couples loss of polarity with JNK-dependent growth independently of Egr. Disc immunostainings revealed that Grnd co-localizes with the apical determinant Crb in the marginal zone, apical to the adherens junction protein E-cadherin (E-cad) and the atypical protein kinase C (aPKC; Fig. 4a and Extended Data Fig. 9c, d). In avl-RNAi discs, Grnd and Crb accumulate in a wider apical domain2 (Extended Data Fig. 9e, g). Apical accumulation of Crb is proposed to be partly responsible for the neoplastic growth induced by avl knockdown, since overexpression of Crb or a membrane-bound cytoplasmic tail of Crb (Crb-intra) mimics the avl-RNAi phenotype2 (Extended Data Fig. 9f). We therefore examined whether Grnd might couple the activity of the Crb complex with JNK-mediated neoplastic growth. Indeed, reducing grnd levels, but not wgn, in rn>crb-intra discs suppresses neoplastic growth as efficiently as inhibiting the activity of the JNK pathway (Fig. 4c–f and Extended Data Fig. 9h, i). Notably, Yki activation is not rescued in these conditions (Extended Data Fig. 9j–l), illustrating the ability of Crb-intra to promote growth independently of Grnd by inhibiting Hippo signalling through its FERM-binding motif (FBM)17,18. Indeed, neoplastic growth and polarity defects induced by a form of Crb-intra lacking its FBM (CrbΔFBM-intra) are both rescued by Grnd silencing (Fig. 4i, j). As expected, the size of rn>crbΔFBM-intra;grnd-RNAi discs is reduced compared to the size of rn>crb-intra; grnd-RNAi discs (compare Fig. 4e, f with Fig. 4j).
a, b, Transverse sections of wild-type discs stained for Grnd (red), Crb (a, green) or Veli (b, green). c–h, Discs expressing crb-intra alone (c), or combined with the indicated genotypes (d–h), dissected 6 days AED, stained for Wg (green). i, j, Discs expressing Crb-intra lacking its FBM (crbΔFBM-intra) alone (i) or with grnd RNAi (j) stained for Wg (in green). k, l, Discs from puc-LacZ animals expressing activated aPKC (aPKCΔN)26 in the posterior domain alone (k) or together with grnd RNAi (l). m, A model for Grnd function.
Crb, Stardust (Sdt; PALS1 in humans), and Pals1-associated tight junction protein (Patj) make up the core Crb complex19, which recruits the adaptor protein Veli (MALS1–3 in humans)20,21,22. In agreement with previous yeast two-hybrid data23, we find that Grnd binds directly and specifically to the PDZ domain of Veli through a membrane-proximal stretch of 28 amino acids in its intracellular domain (Extended Data Fig. 10a–e). Grnd localization is unaffected in crb−/− and veli RNAi mutant clones (Extended Data Fig. 10f–h). However, reducing veli expression rescues the patterning defects and disc morphology of rn>crb-intra mutant cells (Fig. 4g, h), suggesting that Grnd couples Crb activity with JNK signalling through its interaction with Veli. Interestingly, aPKC-dependent activation of JNK signalling16 also depends on Grnd (Fig. 4k, l). aPKC is capable of directly binding and phosphorylating Crb, which is important for Crb function24. This suggests that aPKC, either directly or through Crb phosphorylation, activates Grnd-dependent JNK signalling in response to perturbation of apico-basal polarity.
Our data are consistent with a model whereby Grnd integrates signals from Egr, the unique fly TNF, and apical polarity determinants to induce JNK-dependent neoplastic growth or apoptosis in a context-dependent manner (Fig. 4m). Recent work reveals a correlation between mammalian Crb3 expression and tumorigenic potential in mouse kidney epithelial cells25. The conserved nature of the Grnd receptor suggests that specific TNFRs might carry out similar functions in vertebrates, in which the link between apical cell polarity and tumour progression remains elusive.
Methods
Fly strains and food
Animals were reared at 25 °C on fly food containing, per litre: 17 g inactivated yeast powder, 83 g corn flour, 10 g agar, 60 g white sugar and 4.6 g Nipagin M (in ethanol). The rn-Gal4, UAS-egr, tubGAL80ts animals were raised at 18 °C, shifted to 30 °C for 40 h during early third instar larval development (7 days AEL), and subjected to dissection immediately after (Extended Data Fig. 6).
Cell culture
Drosophila S2 and S2R+ cells were grown in complete Schneiders medium (CSM; Schneiders medium (Invitrogen) supplemented with 10% heat-inactivated FBS (BioWhittaker), 50 U ml−1 penicillin and 50 μg ml−1 streptomycin (Invitrogen)) at 25 °C. Transfections were done using Effectene reagent (Qiagen).
Plasmids
grnd, grnd-extra and grnd-intra coding sequences were PCR amplified from BDGP EST complementary DNA clones RE28509, and cloned into the pENTR/D-TOPO vector using the following gene-specific primers: sense primers CACCATGTCGGTCAGGAAGTTGAG (grnd and grnd-extra) and CACCATGCTGATCCTCTTGCTGCTGACTATCC (grnd-intra); and antisense primers GAAAGCGACGGGAATCGTCGC (grnd and grnd-intra) and CCATCGAAGGAACTGAAACGCG (grnd-extra). The mutations giving rise to the single E202A and double P200A/E202A amino acid changes in Grnd and Grnd125–241 were introduced using the QuickChange site-directed mutagenesis kit (Stratagene), grnd-pENTR or grnd125–241-pGEX-6PI templates, and the following primers: sense, CGGTGAGCACTCGCTATCCGGCGGCAAACAGTACCACGCCCAC; and antisense, GTGGGCGTGGTACTGTTTGCCGCCGGATAGCGAGTGCTCACCG for grndE202A ; sense, CGGTGAGCACTCGCTATGCGGCGGCAAACAGTACCACGCCCAC; and antisense, GTGGGCGTGGTACTGTTTGCCGCCGCATAGCGAGTGCTCACCG for grndP200A/E202A . To generate grnd-Flag, grndE202A-Flag, grnd-extra-Flag and grnd-intra-Flag, grnd, grndE202A , grnd-extra and grnd-intra coding sequences were cloned into the Gateway Destination vector pActin-3×Flag (Drosophila Gateway Vector Collection) for expression in S2 and S2R+ cells. In addition, grnd, grnd-intra and grnd-extra coding sequences were cloned into pUASg.attB for generation of UAS-grnd, UAS-grnd-intra and UAS-grnd-extra transgenic lines. The HA-traf2 plasmid was from K.B.27. The HA-egr and HA-egrΔTNF plasmids were a gift from M. Miura. pActin-iHog-HA and pActin-iHog-Flag were gifts from L. Ruel.
Transgenic flies
To generate the UAS-grnd UAS-grnd-intra and UAS-grnd-extra lines, the pUASattB-grnd, pUASattB-grnd-intra and pUASattB-grnd-extra constructs were introduced into the germ line by injections in the presence of the PhiC31 integrase and inserted in the 86F8-landing site on the 3R chromosome (Bloomington Drosophila Stock Center, BL24749, BestGene). The following RNAi lines were from the GD or KK collections of the Vienna Drosophila RNAi Center (VDRC): grnd RNAi (GD43454, sense primer CGCGAATTCCCTCTTGCTGCTGACTATCCTTCTCGT, and antisense primer GCGTCTAGAGTTGTGGGCGTGGTACTGTTTTCC; and KK104538, sense primer TGGCGCCCCTAGATGAACGGTTATGACGGAGATCG, and antisense primer CGCATGTAGCCTGCCGTTTGTGGTGATTGCGAATG), tak1 RNAi (KK101357, sense primer TGGCGCCCCTAGATGGCTTGTCCTTGAAGTCCTGC, and antisense primer CGACGCCCGCTGATAGCATGAGCAGGAGATTGTCA), bendless RNAi (KK109638, sense primer TGGCGCCCCTAGATGCGCATCATCAAGGAGACTCA, and antisense primer CGCATGTAGCCTGCCTTCTGGAAAAACGCCGATAC), tab2 RNAi (KK100326, sense primer TGGCGCCCCTAGATGGCACTTTCAGCGGAAGAAAC, and antisense primer TAGCCTCCCTAGCGCAGCTCTGCACTGGGATAGGA), traf2 RNAi (GD16125, sense primer CGCGAATTCAGCACCCACTCCGCCCAAGA, and antisense primer GCGTCTAGAACCGAACAGCCCAGCGAGGA), hep RNAi (KK109277, sense primer TGGCGCCCCTAGATGGGCTTCTTGGACAGCTTGAG, and antisense primer CGCATGTAGCCTGCCAGGAGAACAAACGCATCCTG), bsk RNAi (KK104569, sense primer TGGCGCCCCTAGATGAGCATCCACTTCCTCAGCAT, and antisense primer TAGGTCTAGCCCCGCGGTACACCATCACCCTCGTT), scrib RNAi (GD 45556, sense primer CGCGAATTCCGAAGAGGACAACGCACAGCA, and antisense primer GCGTCTAGACTGGGAGCGGGCAAAGTGTAAG), avl (GD5413, sense primer CGCGAATTCCAACAGTAGCGCCAGCCAGCA, and antisense primer CGCTCTAGACCGTGCCCTGCGAGACAAAA), dilp8 (KK102604, sense primer TGGCGCCCCTAGATGACTCGATGGACCTTCTGTCG, and antisense primer CGCATGTAGCCTGCCGTCATCGGAGTCTGTTGCCT), and rh3 RNAi (KK100853, sense primer TGGCGCCCCTAGATGATCTGGGATGACTTATGGCG, and antisense primer TAGCCTCCCTAGCGCTCGATACGCGACTCTTTGTG). The veli-RNAi lines were from U. Thomas. The wengen-RNAi line (wengen-IR) was provided by M. Miura. The GMR-Gal4 (on chromosome II), UAS-egr (on chromosome III), GMR-Gal4, UAS-ecto-egr60 (on chromosome II), GMR-Gal4, UAS-egr (on chromosome II), UAS-CrbΔFBM-intra, DIAP-GFP, aPKCΔN, FRT, Crb11A22 (null) and UAS-wgn transgenic lines were previously published7,10,27,28,29, the UAS-puc2A line was described previously30, the UAS-crb-intra transgenic lines and veli66 mutant flies were from E. Knust, the egr1 and egr3 mutant lines and the UAS-rasV12; FRT82B, scrib1 /TM6b and UAS-rasV12, egr3 ; FRT82B, scrib1/TM6b and yw, eyFlp1; act>y+>Gal4, UAS-GFP; FRT82B, tubgal80 and yw, eyFlp1; act>y+>Gal4, UAS-GFP, egr1 ; FRT82B, tubGal80 lines were from M. Vidal, the w1118 ; +; rnGal4, UAS-eiger, tubGal80ts /TM6B line was provided by I. Hariharan, the yw, hep75/FM7, GFP; SalPE/cyo line was provided by F. Serras. The rn-Gal4, nub-Gal4, puc-LacZ, grndMinos (Mi{MIC}CG10176MI05292), and grndDf(BSC149) lines were provided by the Bloomington Drosophila Stock Center.
Flies’ eye pictures are acquired on a Leica Fluorescence Stereomicroscope MZ16 FA with a Leica digital camera DFC 490.
Generation of wgnKO/KO mutant flies
The wgn knockout allele was generated by ends-in gene targeting as described previously31. The pTV2 targeting construct carried frameshift mutations in exon 1 and 2, an I-SceI site between these two exons, a white+ marker gene, an I-CreI site, up- and downstream homology sequences and two flanking FRT sites. The frameshift mutations were introduced by cutting the unique intrinsic restriction sites AclI and NotI followed by T4 DNA polymerase-mediated fill-in. F3 Larvae carrying the targeting construct together with an hsp70-flp and a hsp70-I-SceI transgene were heat shocked (1 h at 38 °C on day 2 and 3 after egg laying) to generate a linearized targeting construct that aligns and recombines with the endogenous wild-type locus. After crossing the F3 flies to eye-FLP-expressing flies, the F4 was screened for non-mosaic white+ eyed flies carrying an insertion of a processed targeting construct. After processing with FLP and I-SceI, only one FRT site remained next to the white+ marker gene in these flies. Flies still carrying the original donor insertion lost the white+ marker gene, since it was still flanked by two FRT sites. Female F4 ‘w+ wgnKO ’ flies, carrying two wgn copies on the X chromosome were crossed with males of the genotype ‘v/Y ; +/+ ; [v+, 70I-CreI] 1A/TM6’. The F5 offspring were heat shocked (1 h at 38 °C on day 2 after egg laying). F5 offspring males were selected against TM6 (‘w+/−74/Y ; +/+ ; [v+, 70I-CreI] 1A/+’) and typically displayed white+/− mosaics. When they were individually crossed to the X-chromosomal balancer stock ‘bin/#852’, more than 90% of the males were sterile. The F6 flies that hatched were either white+ or white− . The white− flies were tested for the presence of the mutations. The stocks ‘y w (v); P[ry+, 70FLP]4 P[v+, 70I-SceI]2B Sco/S2 Cyo’ and ‘v; +/+; [v+, 70I-CreI] 1A/ TM6, Ubx, e’ were provided by K. Golic and Y. Rong.
Genome-wide screen
As an unbiased approach to identify molecules that are required for neoplastic growth, we used the elavGal80; rn>avl-RNAi/ TM6b, tub>Gal80 testerline, which is characterized by neoplastic growth in the discs and a delay of about 2 days in pupariation. We crossed this line to the Vienna Drosophila RNAi Center (VDRC) collection of phiC31 (KK) RNAi lines (10,100 lines) and screened for candidates that were able to rescue the delay at pupariation as described previously1. One-hundred and twenty-one candidates significantly rescued the pupariation delay in rn>avl-RNAi larvae. Of these, only 8 lines rescued the neoplastic disc growth (Extended Data Fig. 1)
Immunostainings of larval tissues
Tissues dissected from larvae in 1× PBS (137 mM NaCl, 2.7 mM KCl, 4.3 mM Na2HPO4, 1.47 mM KH2PO4, pH 8) at the indicated hour AED, were fixed in 4% formaldehyde (Sigma) in PBS for 20 min at room temperature washed in PBS containing 0.1% Triton X-100 (PBT), blocked for 2 h in PBT containing 10% FBS (PBS-TF), and incubated overnight with primary antibodies at 4 °C. The next day, cells and tissues were washed, blocked in PBS-TF, and incubated with secondary antibodies at 1/500 dilution (Rhodamine Red-X-conjugated donkey anti-rat, Cy3 conjugated donkey anti-rabbit, Cy3 conjugated donkey anti-chicken, Cy3 conjugated donkey anti-mouse and Cy3 conjugated mouse anti-guinea pig, Cy5 conjugated donkey anti-rat, and Cy5 conjugated donkey anti-guinea pig from Jackson ImmunoResearch, goat anti-mouse Alexa Fluor 488, donkey anti-rat Alexa Fluor 488 and goat anti-rabbit Alexa Fluor 488 from Invitrogen) for 2 h at room temperature. After washing, tissues were mounted in Vectashield containing DAPI for staining of DNA (Vector Labs). Fluorescence images were acquired using a Leica SP5 DS (×20 and ×40 objectives) and processed using Adobe Photoshop CS5.
Lysis and immunoprecipitations
Cells were lysed in 100 μl buffer A (50 mM Tris-HCl (pH 8), 150 mM NaCl, 1% NP-50, 1 mM EGTA, 0.5 M NaF, phosphatase inhibitor cocktail 2 (Sigma), complete protease inhibitor cocktail (Roche)). Immunoprecipitations were performed from about 1 × 107 S2 cells. Cells were lysed in 200 μl buffer A, and cell extracts were cleared of membranous material by centrifugation at 10,000 r.p.m. for 15 min. The cleared extracts were incubated with protein G-Sepharose beads (Sigma) for 1 h to reduce unspecific binding of proteins to the beads in the subsequent purifications. Next, the precleared extracts were incubated with 80 μl of protein G-Sepharose beads and 1 μl of the relevant antibody for 3 h. Subsequently, beads were washed three times in buffer A, boiled in sample and reducing buffers (Invitrogen).
Protein preparation
Drosophila Veli full length (Veli1–195) and PDZ domain (Veli85–195) were expressed with an amino-terminal His6-tag from pET43 (Novagen) in Escherichia coli strain BL21 Rosetta, and purified by affinity on Ni-NTA beads (Qiagen). The Veli L27 domain (Veli1–81) was cloned in a modified pET43 plasmid as a His6–MBP-tagged construct. The fusion protein was expressed in bacteria similarly to the other constructs. After a first step of affinity purification on amylose beads (NEB), the protein was incubated with TEV protease (Invitrogen Life Technologies) to cleave the MBP tag, and loaded on a Resource-Q anion-exchange column (GE-Healthcare). The TRAF domain of Drosophila Traf2 (Traf2319–475) was expressed similarly, and purified by affinity followed by ion-exchange chromatography on a Resource-S column (GE-Healthcare). All Grnd constructs were expressed as GST-fusion proteins using a modified version of the pGEX-6PI vector (GE-Healthcare), and purified by affinity on glutathione sepharose (GSH) beads. The purifications were carried out at 4 °C. The PDZ domains of human Par3 (Par3-PDZ2 and Par3-PDZ3 encompassing residues 465–548 and 582–685, respectively) and Drosophila Bazooka (Baz-PDZ1 corresponding to residues 331–419) were expressed as GST-fusion proteins. After the first step of affinity purification on GSH-beads, the GST tag was removed from the constructs by cleavage with PreScission protease, and the PDZ domains were further purified by ion-exchange chromatography.
In vitro binding assay
To test the interaction properties of intracellular Grnd constructs, GST–Grnd fusion proteins (5 μΜ) adsorbed on GSH-beads were incubated for 40 min at 4 °C with 10 μΜ VeliFL,, VeliL27 or VeliPDZ in a buffer containing 10 mM HEPES pH 7.4, 30 mM NaCl, 5% glycerol, 0.5 mM EDTA and 1 mM dithiothreitol (DTT). After two washing steps, the bound species were resolved on SDS–PAGE and visualized by Coomassie staining. To test the specificity of binding, analogous pull-down assays were performed with the PDZ domains of Par3 and Bazooka under the same conditions. To monitor the ability of Traf2 to bind Grnd, 2 μΜ GST–Grnd125–241 wild type or mutated on P200 and/or E202 were absorbed on GSH beads and incubated with 10 μΜ Traf2319–475 in a buffer consisting of 10 mM HEPES pH 7.4, 0.15 M NaCl, 5% glycerol, 0.5 mM EDTA, 1 mM DTT and 0.05% Tween. After washing, complexes immobilized on beads were analysed by SDS–PAGE.
Subcellular fractioning
Subcellular fractioning of 300 wing imaginal discs dissected from wild-type animals was performed using the Qproteome Cell Compartment Kit (Qiagen) according to the manufacturer’s protocol.
Western blotting
Proteins were resolved by SDS–PAGE using 4–12% gradient gels (NuPAGE Novex gel, Invitrogen) and transferred electrophoretically to polyvinylidene difluoride membranes (Milipore). The membranes were incubated for 1 h in blocking buffer (PBS, 5% milk) and incubated overnight at 4 °C in the same buffer containing primary antibodies at 1:1,000 dilutions: rabbit anti-Smoothened32, mouse anti-Lamin (Developmental Studies Hybridoma Bank (DSHB)), mouse anti-Dlg (4F3) from DSHB, mouse anti-Myc 9E10 (sc-40, Santa Cruz Biotechnology), rabbit anti-Myc A-14 (sc-789, Santa Cruz Biotechnology), mouse anti-Flag F3165 (Sigma), rat anti-HA 3F10 (Roche)), guinea pig anti-Grnd at 1:1,000, anti-tubulin (Sigma) at 1:2,000. The anti-Wgn antibody was used as described previously27. Membranes were washed three times in PBS-T, blocked for 1 h, and probed with secondary antibodies in blocking buffer for 1 h at room temperature. After three washes in PBS-T, chemiluminescence was observed using the ECL-Plus western blotting detection system (Amersham Biosciences). Images were generated using the Fujifilm Multi Gauge software.
Antibodies
To generate purified antisera specific for the Grnd protein, two peptides, CAYPHPATANGKDLNA and NAINHPGSDLERAQSQ, corresponding to amino acids 140–154 and 161–176, respectively, were used as immunogens in guinea pigs (Eurogentec). The following primary antibodies were used for immunoflourescence in this study: mouse anti-Wg (4D4) from DSHB 1/50, mouse anti-Dlg (4F3) from DSHB 1/500, rat anti-E-Cad (DCAD2) from DSHB 1/500, mouse anti-MMP1 (3A6B4/5H7B11/3B8D12; the three different antibodies were mixed in amounts (1:1:1) from DSHB (1:25)), chicken anti-β-galactosidase from GeneTek 1/1,000, rabbit anti-cleaved caspase-3 (Asp 175) from Cell Signaling Technology 1/500, rabbit anti-aPKC from Santa Cruz Biotechnology 1/1,000, guinea pig anti-Grnd 1/100, rabbit anti-Veli 1/1,000 (gift from G. Pietrini), and rat anti-Crb 1/1,000 (gift from E. Knust).
Sequence alignment and homology modelling
Multiple sequence alignment, per cent identity matrix, and phylogenic data were generated using Clustal Omega (http://www.ebi.ac.uk/Tools/services/web/toolform.ebi?tool=clustalo). Data of five human death receptors of the TNFR superfamily with the highest identity with Grnd are shown. Slight manual adjustment of the alignments was performed. Amino acids are highlighted where Grnd shares identity or similarity with death receptor(s) (Fig. 1) or Grnd-like proteins from other insect species (Extended Data Fig. 2d), or where Fas shares identity or similarity with Grnd and/or Wgn (Extended Data Fig. 2b). Amino acid similarity is presented based on scores in Gonnet PAM 250 matrix (>0.5, highly similar; ≤0.5, weakly similar). Amino acid positions are indicated in parentheses or at the end of the shown sequences. The phylogram was drawn using interactive Tree of Life33 (http://itol.embl.de/). Number on each branch indicates branch length (Extended Data Fig. 2a). UniProt protein identifiers are: Fas (P25445), DR4 (O00220), DR5 (O14763), DcR1 (O14798), DcR2 (Q9UBN6), and Wengen (Q9VWS4). Information of protein domains in TNFR superfamily members used to construct protein domain diagram (Extended Data Fig. 2d) was obtained from UniProt (http://www.uniprot.org/).
Genotypes
Fig. 1a: w−; +; rn-Gal4/ puc-lacZ. Fig. 1b: w−; elav-Gal80/+; rn-Gal4, UAS avl RNAi GD/ puc-lacZ. Fig. 1c: w−; elav-Gal80/ UAS grnd RNAi KK; rn-Gal4, UAS avl RNAi GD/puc-lacZ. Fig. 1d: w−; elav-Gal80/ UAS tak1 RNAi KK; rn-Gal4, UAS avl RNAi GD/puc-lacZ. Fig. 1i: w−; nub-Gal4/+; UAS grnd / puc-lacZ. Fig. 1j: w−; nub-Gal4/+; UAS grnd-intra / puc-lacZ.
Fig. 2a: w− ; GMR-Gal4/+; +. Fig. 2b: w− ; GMR-Gal4, UAS egr/+; MKRS/+. Fig. 2c: wgnKO ; GMR-Gal4, UAS egr/+; MKRS/+. Fig. 2d: w− ; GMR-Gal4, UAS egr/ UAS grnd RNAi KK; MKRS/+. Fig. 2e: w− ; GMR-Gal4, UAS egr/ +; MKRS/ UAS grnd-extra. Fig. 2f: yw, hs-FLP; Act>CD2>Gal4, UAS GFP/+; UAS egr/+. Fig. 2g: yw, hs-FLP; Act>CD2>Gal4, UAS GFP/ UAS tak1 RNAi KK; UAS egr/+. Fig. 2h–i: yw, hs-FLP; Act>CD2>Gal4, UAS GFP/ UAS grnd RNAi KK; UAS egr/+.
Fig. 3a: yw, hsFLP; Act>CD2>Gal4, UAS GFP/+; +. Fig. 3b: yw, hs-FLP; Act>CD2>Gal4, UAS GFP/ UAS rh3 RNAi KK; +. Fig. 3c: yw, hs-FLP; Act>CD2>Gal4, UAS GFP/ UAS grnd RNAi KK; +. Fig. 3d: yw, hs-FLP; Act>CD2>Gal4, UAS GFP/+; UAS scrib RNAi GD/+. Fig. 3e: yw, hs-FLP; Act>CD2>Gal4, UAS GFP/ UAS rh3 RNAi KK; UAS scrib RNAi GD/+. Fig. 3f: yw, hs-FLP; Act>CD2>Gal4, UAS GFP/ UAS grnd RNAi KK; UAS scrib RNAi GD/+. Fig. 3g: yw, ey-FLP; Act>y+ >Gal4, UAS GFP; UAS rasV12/+; FRT82B, scrib1 / FRT82B, tubGal80. Fig. 3h: yw, ey-FLP; Act>y+ >Gal4, UAS GFP; UAS rasV12 , UAS grnd RNAi KK /+; FRT82B, scrib1 / FRT82B, tubGal80. Fig. 3i: yw, ey-FLP; Act> y+ >Gal4, UAS GFP; egr1 / UAS rasV12 , egr3 ; FRT82B, tubGal80/ FRT82B, scrib1 .
Fig. 4a, b: w− . Fig. 4c: w−; UAS crb-intra/+; rn-Gal4 /+. Fig. 4d: w−; UAS crb-intra/ UAS puc2A; rn-Gal4 /+. Fig. 4e: w−; UAS crb-intra/ UAS grnd RNAi KK; rn-Gal4 /+. Fig. 4f: w−; grndMinos/ grndMinos; rn-Gal4 / UAS crb-intra. Fig. 4g: w−; UAS crb-intra/ UAS veli RNAi ; rn-Gal4/ UAS veli RNAi. Fig. 4h: w−; UAS crb-intra/ nub-Gal4 ; veli66/ veli66. Fig. 4i: w−; +; UAS crbΔFBM-intra/ rn-Gal4. Fig. 4j: w−; UAS grnd RNAi KK/+ ; UAS crbΔFBM-intra/ rn-Gal4. Fig. 4k: w−; en-Gal4, UAS GFP/+; UAS aPKCΔN/ puc-lacZ. Fig. 4l: w−; UAS grnd RNAi KK/ en-Gal4, UAS GFP; UAS aPKCΔN/ puc-lacZ.
Extended Data Fig. 1c: w−; elav-Gal80/+; rn-Gal4, UAS avl RNAi GD /+. Extended Data Fig. 1d: w−; elav-Gal80/ UAS grnd RNAi GD; rn-Gal4, UAS avl RNAi GD /+. Extended Data Fig. 1e: w−; elav-Gal80/ Rh3 RNAi KK; rn-Gal4, UAS avl RNAi GD /+. Extended Data Fig. 1f, g: w− ; ptc-Gal4, UAS-GFP/ UAS grnd RNAi KK; +.
Extended Data Fig. 4a: w−; +; rn-Gal4 /+. Extended Data Fig. 4b: w−; +; rn-Gal4 / UAS grnd. Extended Data Fig. 4c, d: w−; +; rn-Gal4 / UAS grnd-intra. Extended Data Fig. 4e: yw−; sal-Gal4 /+; UAS grnd-intra /+. Extended Data Fig. 4f: yw−, hep75; sal-Gal4 /+; UAS grnd-intra /+.
Extended Data Fig. 5a: w− ; GMR-Gal4, UAS egr/+; MKRS/+. Extended Data Fig. 5b: w− ; GMR-Gal4, UAS egr/ UAS-traf2 RNAi GD; MKRS/+. Extended Data Fig. 5c: w− ; GMR-Gal4, UAS egr/ UAS puc2A ; MKRS/+. Extended Data Fig. 5d: w− ; GMR-Gal4, UAS egr/ UAS tak1 RNAi KK; MKRS/+. Extended Data Fig. 5e: w− ; GMR-Gal4, UAS egr/ UAS-wengen RNAi IR; MKRS/+. Extended Data Fig. 5f: w− ; GMR-Gal4, UAS egr/ UAS-grnd RNAi GD; MKRS/+. Extended Data Fig. 5g: wgnKO ; GMR-Gal4, UAS egr/ UAS-grnd RNAi GD; MKRS/+. Extended Data Fig. 5h: wgnKO ; +; +. Extended Data Fig. 5i: w− ; GMR-Gal4/ UAS-grnd RNAi KK; +. Extended Data Fig. 5j: w− ; GMR-Gal4/+; UAS-grnd-extra/+. Extended Data Fig. 5k: w− ; GMR-Gal4, UAS-ecto-egr60/+; MKRS/+. Extended Data Fig. 5l: w− ; GMR-Gal4, UAS-ecto-egr60/ UAS puc2A ; MKRS/+.
Extended Data Fig. 6c: w−. Extended Data Fig. 6d: w−; grndMinos/ grndMinos; +. Extended Data Fig. 6e: w−; +; rn-Gal4, UAS egr, tubGal80ts /+. Extended Data Fig. 6f: w−; UAS-grnd RNAi KK/+; rn-Gal4, UAS egr, tubGal80ts/+. Extended Data Fig. 6g: w−; UAS-traf2 RNAi GD /+; rn-Gal4, UAS egr, tubGal80ts/+. Extended Data Fig. 6h: w−; UAS-tak1 RNAi KK/+; rn-Gal4, UAS egr, tubGal80ts/+. Extended Data Fig. 6i: w−; grndMinos/ grndMinos; rn-Gal4, UAS egr, tubGal80ts/+. Extended Data Fig. 6j: w−; grndMinos/ grndDf; rn-Gal4, UAS egr, tubGal80ts/+.
Extended Data Fig. 7a: yw, ey-FLP; Act>y+ >Gal4, UAS GFP; +; FRT82B/ FRT82B, tubGal80. Extended Data Fig. 7b: yw, ey-FLP; Act>y+ >Gal4, UAS GFP; +; FRT82B, scrib1 / FRT82B, tubGal80. Extended Data Fig. 7c: yw, ey-FLP; Act>y+ >Gal4, UAS GFP; UAS grnd RNAi KK /+; FRT82B, scrib1 / FRT82B, tubGal80. Extended Data Fig. 7d: ey-FLP; Act>CD2>Gal4, UAS GFP/ UAS grnd RNAi KK ; +. Extended Data Fig. 7e–g: yw, ey-FLP; Act>y+ >Gal4, UAS GFP; UAS rasV12 /+; FRT82B, scrib1 / FRT82B, tubGal80.
Extended Data Fig. 8a: yw, ey-FLP; Act>y+ >Gal4, UAS GFP; UAS rasV12/+; FRT82B, scrib1 / FRT82B, tubGal80. Extended Data Fig. 8b: yw, ey-FLP; Act>y+ >Gal4, UAS GFP; UAS rasV12 , UAS grnd RNAi KK /+; FRT82B, scrib1 / FRT82B, tubGal80. Extended Data Fig. 8c: yw, ey-FLP; Act>y+ >Gal4, UAS GFP; UAS rasV12 , UAS dilp8 RNAi KK /+; FRT82B, scrib1 / FRT82B, tubGal80. Extended Data Fig. 8d: w; UAS rasV12, UAS-dlg RNAi GD /+; ey-FLP; Act≫Gal4, UAS GFP /+. Extended Data Fig. 8e: w; UAS rasV12, UAS-dlg RNAi GD / UAS-grnd RNAi GD ; ey-FLP; Act≫Gal4, UAS GFP /+. Extended Data Fig. 8f: w; UAS rasV12, UAS-dlg RNAi GD / UAS-grnd RNAi KK ; ey-FLP; Act≫Gal4, UAS GFP /+. Extended Data Fig. 8g: wgnKO; UAS rasV12, UAS-dlg RNAi GD /+; ey-FLP; Act≫Gal4, UAS GFP /+.
Extended Data Fig. 9a: w−; +; rn-Gal4, UAS avl RNAi GD/+. Extended Data Fig. 9b: w−; egr1 / egr3; rn-Gal4, UAS avl RNAi GD/+. Extended Data Fig. 9c, d: w−; +; +. Extended Data Fig. 9e: w−; elav-Gal80/+; rn-Gal4, UAS avl RNAi GD/+. Extended Data Fig. 9f: w−; UAS crb-intra/+; rn-Gal4 /+. Extended Data Fig. 9g: w−; elav-Gal80/+; rn-Gal4, UAS avl RNAi GD/+. Extended Data Fig. 9h: w−; UAS crb-intra/+; rn-Gal4 /+. Extended Data Fig. 9i: wgnKO; UAS crb-intra/+; rn-Gal4 /+. Extended Data Fig. 9j: w−; +; rn-Gal4 / diap-GFP. Extended Data Fig. 9k: w−; UAS crb-intra/+; rn-Gal4 / diap-GFP. Extended Data Fig. 9l: w−; UAS crb-intra/ UAS-grnd RNAi KK ; rn-Gal4 / diap-GFP.
Extended Data Fig. 10f, g: yw, hsFLP; +; FRT82B, Ubi-GFP/ FRT82B, crb11A22 . Extended Data Fig. 10h: yw, hs-FLP; Act>CD2>Gal4, UAS GFP / UAS veli RNAi ; UAS veli RNAi /+.
References
Colombani, J., Andersen, D. S. & Leopold, P. Secreted peptide Dilp8 coordinates Drosophila tissue growth with developmental timing. Science 336, 582–585 (2012)
Lu, H. & Bilder, D. Endocytic control of epithelial polarity and proliferation in Drosophila . Nature Cell Biol. 7, 1232–1239 (2005)
Chakrabandhu, K. et al. The extracellular glycosphingolipid-binding motif of Fas defines its internalization route, mode and outcome of signals upon activation by ligand. Cell Death Differ. 15, 1824–1837 (2008)
Ye, H. et al. Distinct molecular mechanism for initiating TRAF6 signalling. Nature 418, 443–447 (2002)
Vidal, M. The dark side of fly TNF: an ancient developmental proof reading mechanism turned into tumor promoter. Cell Cycle 9, 3851–3856 (2010)
Igaki, T. et al. Eiger, a TNF superfamily ligand that triggers the Drosophila JNK pathway. EMBO J. 21, 3009–3018 (2002)
Moreno, E., Yan, M. & Basler, K. Evolution of TNF signaling mechanisms: JNK-dependent apoptosis triggered by Eiger, the Drosophila homolog of the TNF superfamily. Curr. Biol. 12, 1263–1268 (2002)
Xue, L. et al. Tumor suppressor CYLD regulates JNK-induced cell death in Drosophila . Dev. Cell 13, 446–454 (2007)
Kanda, H., Igaki, T., Kanuka, H., Yagi, T. & Miura, M. Wengen, a member of the Drosophila tumor necrosis factor receptor superfamily, is required for Eiger signaling. J. Biol. Chem. 277, 28372–28375 (2002)
Narasimamurthy, R. et al. Structure–function analysis of Eiger, the Drosophila TNF homolog. Cell Res. 19, 392–394 (2009)
Brumby, A. M. & Richardson, H. E. scribble mutants cooperate with oncogenic Ras or Notch to cause neoplastic overgrowth in Drosophila . EMBO J. 22, 5769–5779 (2003)
Igaki, T., Pastor-Pareja, J. C., Aonuma, H., Miura, M. & Xu, T. Intrinsic tumor suppression and epithelial maintenance by endocytic activation of Eiger/TNF signaling in Drosophila . Dev. Cell 16, 458–465 (2009)
Cordero, J. B. et al. Oncogenic Ras diverts a host TNF tumor suppressor activity into tumor promoter. Dev. Cell 18, 999–1011 (2010)
Pagliarini, R. A. & Xu, T. A genetic screen in Drosophila for metastatic behavior. Science 302, 1227–1231 (2003)
Uhlirova, M. & Bohmann, D. JNK- and Fos-regulated Mmp1 expression cooperates with Ras to induce invasive tumors in Drosophila . EMBO J. 25, 5294–5304 (2006)
Sun, G. & Irvine, K. D. Regulation of Hippo signaling by Jun kinase signaling during compensatory cell proliferation and regeneration, and in neoplastic tumors. Dev. Biol. 350, 139–151 (2011)
Ling, C. et al. The apical transmembrane protein Crumbs functions as a tumor suppressor that regulates Hippo signaling by binding to Expanded. Proc. Natl Acad. Sci. USA 107, 10532–10537 (2010)
Robinson, B. S., Huang, J., Hong, Y. & Moberg, K. H. Crumbs regulates Salvador/Warts/Hippo signaling in Drosophila via the FERM-domain protein Expanded. Curr. Biol. 20, 582–590 (2010)
Bulgakova, N. A. & Knust, E. The Crumbs complex: from epithelial-cell polarity to retinal degeneration. J. Cell Sci. 122, 2587–2596 (2009)
Bachmann, A., Grawe, F., Johnson, K. & Knust, E. Drosophila Lin-7 is a component of the Crumbs complex in epithelia and photoreceptor cells and prevents light-induced retinal degeneration. Eur. J. Cell Biol. 87, 123–136 (2008)
Bachmann, A. et al. Cell type-specific recruitment of Drosophila Lin-7 to distinct MAGUK-based protein complexes defines novel roles for Sdt and Dlg-S97. J. Cell Sci. 117, 1899–1909 (2004)
Olsen, O. et al. Renal defects associated with improper polarization of the CRB and DLG polarity complexes in MALS-3 knockout mice. J. Cell Biol. 179, 151–164 (2007)
Giot, L. et al. A protein interaction map of Drosophila melanogaster . Science 302, 1727–1736 (2003)
Sotillos, S., Diaz-Meco, M. T., Caminero, E., Moscat, J. & Campuzano, S. DaPKC-dependent phosphorylation of Crumbs is required for epithelial cell polarity in Drosophila . J. Cell Biol. 166, 549–557 (2004)
Karp, C. M. et al. Role of the polarity determinant crumbs in suppressing mammalian epithelial tumor progression. Cancer Res. 68, 4105–4115 (2008)
Betschinger, J., Mechtler, K. & Knoblich, J. A. The Par complex directs asymmetric cell division by phosphorylating the cytoskeletal protein Lgl. Nature 422, 326–330 (2003)
Geuking, P., Narasimamurthy, R. & Basler, K. A genetic screen targeting the tumor necrosis factor/Eiger signaling pathway: identification of Drosophila TAB2 as a functionally conserved component. Genetics 171, 1683–1694 (2005)
Izaddoost, S., Nam, S. C., Bhat, M. A., Bellen, H. J. & Choi, K. W. Drosophila Crumbs is a positional cue in photoreceptor adherens junctions and rhabdomeres. Nature 416, 178–183 (2002)
Zhang, L. et al. The TEAD/TEF family of transcription factor Scalloped mediates Hippo signaling in organ size control. Dev. Cell 14, 377–387 (2008)
Martín-Blanco, E. et al. puckered encodes a phosphatase that mediates a feedback loop regulating JNK activity during dorsal closure in Drosophila . Genes Dev. 12, 557–570 (1998)
Rong, Y. S. et al. Targeted mutagenesis by homologous recombination in D. melanogaster . Genes Dev. 16, 1568–1581 (2002)
Ruel, L., Rodriguez, R., Gallet, A., Lavenant-Staccini, L. & Therond, P. P. Stability and association of Smoothened, Costal2 and Fused with Cubitus interruptus are regulated by Hedgehog. Nature Cell Biol. 5, 907–913 (2003)
Ciccarelli, F. D. et al. Toward automatic reconstruction of a highly resolved tree of life. Science 311, 1283–1287 (2006)
Acknowledgements
We thank G. Jarretou for technical assistance, the Vienna Drosophila RNAi Centers, the Drosophila Genetics Resource Center, the Bloomington Stock Center, N. Caridi and S. Pasqualato for technical assistance and P.L. laboratory members for comments on the manuscript. This work was supported by the CNRS, INSERM, Agence Nationale de la Recherche, Fondation pour la Recherche Médicale, Association pour la Recherche contre le Cancer (grant no. PJA20131200042 to J.C.), European Research Council (Advanced grant no. 268813 to P.L.), Marie Curie Life Long Training (grant no. 252373 to D.S.A.), the Labex Signalife program (grant ANR-11-LABX-0028-01 to P.L.), the Italian Association for Cancer Research (AIRC IG-12877) and the Italian Ministry of Health (GR-2008-1134103) to M.M.
Author information
Authors and Affiliations
Contributions
D.S.A., J.C., K.C., A.-O.H., K.B., M.M. and P.L. designed the research; D.S.A., J.C., K.C., V.P., E.B., M.R. and J.T. performed experiments; D.S.A., J.C., K.C., E.B., A.-O.H., K.B., V.P., M.M. and P.L. analysed the data; D.S.A., J.C. and P.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Identification of Grnd in a genetic screen for molecules implicated in neoplastic growth.
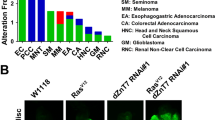
a, Schematic of the genome-wide screen. Ten-thousand one-hundred RNAi lines were screened for their abilities to rescue the delay of the rn>avl-RNAi condition. Of the 121 candidates able to rescue the delay of the rn>avl condition, only 8 also rescued the neoplastic growth. b, Schematic of the five components of the JNK pathway shown to be required for the neoplastic overgrowth in the avl loss of function condition. Although no RNAi line targeting Traf2 was included in the original screen, Traf2 was subsequently shown to be required for neoplastic overgrowth in the avl-RNAi condition using an alternative RNAi line. c–e, Wing discs of the indicated genotypes dissected 5 days (left) or 6 days (right) AED and stained for Wg (green) and Caspase3 (red). f, g, Transverse (f) or xy (g) sections of dissected wing discs expressing grnd RNAi in the patch domain (GFP, green) stained for Grnd (red) and DAPI (g, right, blue) to visualize the nuclei. h, Subcellular fractionation of cleared extract from dissected wing discs shows that Grnd, like the transmembrane receptor Smoothened, localizes to the membrane fraction (PM), whereas Disc-large remains in the cytoplasmic fraction (Cyto). Laminin serves as a control for nuclear fractions.
Extended Data Figure 2 Sequence homology of the extracellular domain of Grnd to CRDs of human TNFRs.
a, Per cent identity matrix and phylogenic data from multiple local sequence alignment of the entire extracellular domains of Grnd and Wgn with extracellular domains (corresponding to a part of CRD2 and all of CRD3) of various human TNFR superfamily members. The extracellular domain of Grnd shares the highest identity with the GBM-harbouring CRD of Fas (26%). A similar result is observed for Wgn, although it exhibits lower identity (22%) and a higher degree of divergence to Fas. b, Sequence alignment of Fas, Grnd and Wgn demonstrates that Grnd shares a higher degree of similarity with the GBM-harbouring CRD of Fas than Wgn. This is particularly apparent in the immediate region encompassing the functionally critical F134 (marked with an asterisk) of the GBM of Fas. Blue indicates residue identity, yellow strong similarity, and grey weak similarity of Fas with Grnd and/or Wgn. c, Schematic representation of Grnd, Wgn, and various members of the human TNFR superfamily showing a high level of diversity in the number and arrangement of CRDs. The colour codes used are: red for signal peptide, green for extracellular domains, orange for transmembrane domains, and blue for intracellular domain. Numbers in each box indicate corresponding amino acid positions. Pro, prodomain; T1–5, TAPE1–5 (threonine-, alanine-, proline- and glutamine-rich repeats). d, Multiple sequence alignment showing the conservation of the extracellular domain of Grnd in other insect species. Whereas global similarity of the extracellular domain of Grnd degenerates outside the melanogaster group, the cysteine residues of the CRD (highlighted in red) and the aromatic residue (F/Y, marked with an asterisk) corresponding to F134 of Fas CRD3 remain highly conserved throughout the Hymenoptera order. Blue indicates residue identity; yellow, strong similarity; and grey, weak similarity to Grnd.
Extended Data Figure 3 Identification of a conserved TRAF6-like binding motif in the intracellular domain of Grnd proteins.
Multiple sequence alignment of the intracellular domain of Grnd and Grnd-like proteins in other species. A binding motif for TRAF6 (PxExx(aromatic/acidic residue)), the closest homologue of Drosophila Traf2, was previously identified in human TNFRs4. The position of a related putative Traf2-binding motif in the intracellular domain of Grnd is indicated. Note that, although sequence similarity of the intracellular domain of Grnd degenerates outside the family Drosophilidae, the core residues of this motif (highlighted in red) remain highly conserved throughout the order Hymenoptera. Blue indicates residue identity; yellow, strong similarity to Grnd.
Extended Data Figure 4 Grnd acts upstream of JNK signalling.
a–d, Wild-type wing discs (a) or discs expressing grnd (b) or grnd-intra (c, d) in the rn domain were dissected 5 days AED and stained for Wg (left; green), Cas3 (middle; red), and Grnd (right; grey). d, e, Wing discs expressing grnd-intra in the sal domain (marked by a truncated line) in a wild-type (d) or hep75 mutant (e) background were dissected 5 days AED and stained for Wg (left; green), Cas3 (middle; red), and Grnd (right; grey).
Extended Data Figure 5 Characterization of the wgnKO mutant allele.
a–e, Light micrographs of Drosophila adult eyes are shown. Original magnification, ×30. The small-eye phenotype caused by GMR-driven expression of egr (a) is rescued by co-expressing puc (c) or reducing the activity of Traf2 (b), but not by reducing Wgn activity (d; also serves as a control to exclude a Gal4 titration effect). f, g, Reducing Grnd activity partially rescues the small-eye phenotype, giving rise to the hanging-eye phenotype (f) and is not further rescued in a wgnKO mutant background (g). h, wgnKO mutant flies do not display any eye phenotype. i, j, Reducing Grnd levels or expressing Grnd-extra in the GMR domain does not affect eye morphology. k, GMR-Gal4-induced expression of a soluble form of Egr lacking the transmembrane and cytoplasmic domain (Ecto-Egr-60) gives rise to a small-eye phenotype10. l, Co-expression of puc2A in a GMR>ecto-egr-60 background partially rescues the small-eye phenotype, giving rise to a hanging-eye phenotype10. m, Sequence analysis of DNA extracted from flies homozygous for the wgnKO mutation showing the two frameshift mutations introduced by cutting the unique intrinsic restriction sites AclI (exon 1) and NotI (exon 2) followed by T4 DNA polymerase-mediated fill-in. n, Western blot of fly extracts from the indicated genotypes probed with an anti-Wgn antibody27. KO, knockout; OE, overexpressing; WT, wild type.
Extended Data Figure 6 Characterization of the grndMinos mutant.
a, Schematic representation of the Minos transposon insertion in the grnd gene region corresponding to the 5′ untranslated region of the transcript. b, Western blot of wing disc extracts from the indicated genotypes probed with anti-Grnd (top) and anti-β-tubulin (bottom) antibodies. Grnd protein migrates at the predicted size of 27 kDa (lane 1, control). Grnd protein levels are decreased below detection level in both grndMinos/Minos and grndMinos/Df mutant flies, indicating that grndMinos/Minos is indistinguishable from a null mutant. c, d, Polarity is not affected in grndMinos/Minos and grndMinos/Df wing discs. Transverse sections of dissected control (c) or grndMinos/Minos (d) wing discs stained for Grnd (top; red), Crb (middle top; blue) and Dlg (middle bottom; green). e–j, Wing discs dissected from third instar larvae after 40 h of Egr expression in the rn domain. Pouch ablation, ubiquitous Wg expression and apoptosis observed in rn>egr, Tub-Gal80ts discs (e) are suppressed to various extents upon co-expression of grnd-RNAi (f) or reduction of JNK activity (g, h) and fully suppressed in grndMinos/Minos (i) and grndMinos/Df (j) mutant backgrounds.
Extended Data Figure 7 Grnd is highly expressed in RasV12/scrib−/− micro-metastases.
a–c, Reducing grnd levels in scrib−/− mutant clones prevent their elimination. Eye discs dissected 5 days AED carrying GFP-labelled MARCM clones of the indicated genotypes stained for Grnd (middle; red) and DAPI (right; grey). d, Reducing Grnd levels does not affect cell viability. Eye discs dissected 5 days AED carrying GFP-labelled grnd RNAi Flip-out clones stained for Grnd (middle; red) and DAPI (right; grey). e–g, Invasive brain tumours and micro-metastases display high levels of Grnd protein. Dissected eye–brain complex (e), gut (f) and fat body (g) displaying GFP-labelled invasive RasV12/scrib−/− clones (e; green) or micro-metastases (f, g; green) stained for Grnd (red) and DAPI (blue).
Extended Data Figure 8 Grnd, but not Wgn, is required for Mmp1 expression both in RasV12/scrib1 and RasV12/Dlg RNAi tumours.
a–c, Eye–brain complexes dissected 7 days AED carrying RasV12/scrib−/− (a), RasV12/scrib−/− + grnd-RNAi (b), or RasV12/scrib−/− + dilp8-RNAi (c; control RNAi to exclude a Gal4 titration effect) clones labelled by GFP and stained for Mmp1 (middle; red). RasV12/scrib−/− and RasV12/scrib−/− + dilp8-RNAi clones invade the ventral nerve cord (a, c; see white arrow). d–g, Eye–brain complexes dissected 7 days AED carrying RasV12/Dlg-RNAi (d) or RasV12/Dlg-RNAi + grnd-RNAi (e, f) clones (labelled by GFP), or RasV12/Dlg-RNAi GFP-labelled clones in a wgn-null mutant background (g) stained for Mmp1 (middle; red) and DAPI (right; blue). RasV12/Dlg RNAi clones invade the ventral nerve cord in wgn-null mutant animals (g; see white arrow).
Extended Data Figure 9 Egr is not required for the rn>avl RNAi-induced neoplastic growth.
a, b, Wing discs of the indicated genotypes dissected 6 days AED and stained for Wg (green). c, d, Transverse sections of dissected wild-type discs stained for Grnd (red) and either E-cad (c; in green), or aPKC (d; in green). e, g, Transverse sections of rn>avl RNAi wing discs dissected 5 days AED and stained for Grnd (top, bottom; red) and either E-cad (e, middle, bottom; green) or Crb (g, middle, bottom; green). The rn expression domain is indicated (e, top). f, Wing discs expressing crb-intra in the rn domain (rn>crb-intra) dissected 6 days AED and stained for Disc-large (Dlg, green) to visualize neoplastic structures and Grnd (red). h, i, Wing discs expressing crb-intra in the rn domain of wild-type (h) or wgn-knockout mutant (i) animals dissected 6 days AED and stained for Wg (green). j–l, Reducing grnd levels does not affect crb-intra-mediated inhibition of Hippo signalling, which causes an upregulation of the Yki transcriptional target Diap1. Control discs (j) or wing discs expressing Crb-intra alone (k) or together with grnd RNAi (l) in the rn domain were dissected from animals carrying a reporter gene for Hippo pathway activity (Diap-GFP, green) and stained for Wg (red).
Extended Data Figure 10 Grnd directly interacts with the PDZ domain of Veli and neither depends on Veli nor Crb for its proper localization.
a, b, Schematic representation of Grnd and Veli (also known as Lin-7) domains and the various truncations used to map the interface of the Grnd–Veli interaction. L27, Lin-2/Lin-7 domain; PDZ, PSD-95/Dlg/ZO-1 domain; TM, transmembrane domain. c, GST–Grnd125–241 immobilized on GSH beads was incubated with full-length Veli (VeliFL) or the complementary Veli fragments L27 (VeliL27) and PDZ domain (VeliPDZ). The PDZ1 domain of the polarity protein Bazooka was included in the pull-down as a specificity control (BazPDZ1). Coomassie-blue stained SDS–polyacrylamide gel electrophoresis (SDS–PAGE) was used to visualize species retained on beads. VeliFL and VeliPDZ bind to Grnd (lanes 10 and 12), but not VeliL27 nor BazPDZ1 (lanes 11 and 9), indicating that Grnd specifically recognizes the PDZ domain of Veli. d, To map the interface between Grnd and VeliPDZ, the same binding assay was performed using a battery of Grnd deletion mutants adsorbed on GSH beads. VeliPDZ forms a complex with Grnd125–241 and Grnd125–168 (lanes 1 and 3), but not with the carboxy-terminal region of Grnd (lanes 2 and 4) or GST alone (lane 5, control). e, To address further the specificity of the binding of Grnd125–241 to the PDZ domain of Veli, we repeated the pull-down assay with the PDZ domains of Baz and its human orthologue PAR3. VeliPDZ is retained on GST–Grnd125–241 beads (lane 9), while PAR3PDZ2, PAR3PDZ3 and BazPDZ1 are not (lanes 10, 11 and 12). f–g, Reducing Crb levels affects the localization of Veli, but not Grnd. f–g, xy (f) or transverse (g) sections of dissected wing discs bearing crb−/− mutant clones (labelled by absence of GFP) stained for Grnd (f, bottom left, g, bottom; white) or Veli (f, bottom right; red). h, Reducing Veli levels does not affect Crb20 or Grnd localization. Transverse sections of dissected wing discs bearing clones expressing veli RNAi (h, bottom; labelled by the presence of GFP) stained for Grnd (top; red) and Crb (middle; blue).
Rights and permissions
About this article
Cite this article
Andersen, D., Colombani, J., Palmerini, V. et al. The Drosophila TNF receptor Grindelwald couples loss of cell polarity and neoplastic growth. Nature 522, 482–486 (2015). https://doi.org/10.1038/nature14298
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14298
This article is cited by
-
Polarity protein AF6 functions as a modulator of necroptosis by regulating ubiquitination of RIPK1 in liver diseases
Cell Death & Disease (2023)
-
Cholinergic neurons trigger epithelial Ca2+ currents to heal the gut
Nature (2023)
-
The TNFR Wengen regulates the FGF pathway by an unconventional mechanism
Nature Communications (2023)
-
A TNFR1–UBCH10 axis drives lung squamous cell carcinoma dedifferentiation and metastasis through a cell-autonomous signaling loop
Cell Death & Disease (2022)
-
A role for Flower and cell death in controlling morphogen gradient scaling
Nature Cell Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.