Abstract
Hepatocellular carcinoma (HCC) is the second most common cause of cancer-related death. Non-alcoholic fatty liver disease (NAFLD) affects a large proportion of the US population and is considered to be a metabolic predisposition to liver cancer1,2,3,4,5. However, the role of adaptive immune responses in NAFLD-promoted HCC is largely unknown. Here we show, in mouse models and human samples, that dysregulation of lipid metabolism in NAFLD causes a selective loss of intrahepatic CD4+ but not CD8+ T lymphocytes, leading to accelerated hepatocarcinogenesis. We also demonstrate that CD4+ T lymphocytes have greater mitochondrial mass than CD8+ T lymphocytes and generate higher levels of mitochondrially derived reactive oxygen species (ROS). Disruption of mitochondrial function by linoleic acid, a fatty acid accumulated in NAFLD, causes more oxidative damage than other free fatty acids such as palmitic acid, and mediates selective loss of intrahepatic CD4+ T lymphocytes. In vivo blockade of ROS reversed NAFLD-induced hepatic CD4+ T lymphocyte decrease and delayed NAFLD-promoted HCC. Our results provide an unexpected link between lipid dysregulation and impaired anti-tumour surveillance.
Similar content being viewed by others
Main
HCC commonly arises in patients with underlying chronic liver disease, and is considered a typical inflammation-associated tumour1. Recent epidemiology studies indicate an increase in the rate of NAFLD-induced HCC2,3,4,5. Immune evasion mediated by numerous immune suppressor mechanisms involving different immune cell subsets have been shown to contribute to HCC initiation and progression6, and patients with tumours containing lymphocytic infiltrates show longer survival and lower risk of recurrence7. However, the role of adaptive immune responses in NAFLD and HCC have just begun to be understood8.
Here we investigated how metabolic changes observed in NAFLD promoted hepatocarcinogenesis using a series of different mouse NAFLD and HCC models and confirmed our results using human samples. Inducible liver-specific MYC oncogene transgenic mice (MYC-ON)9 were fed with a methionine–choline-deficient diet (MCD) to induce NAFLD10 (Fig. 1a and Extended Data Fig. 1a, b). Earlier microscopic liver tumour lesions were found in MYC-ON MCD mice (Fig. 1b, top). As expected, MYC-ON MCD mice showed more macroscopic liver tumours (Fig. 1b, bottom, and c). Similar results were obtained in MYC-ON mice fed a choline-deficient and amino-acid-defined diet (CDAA), another NAFLD-inducing diet (Extended Data Fig. 1c–e)11. Again, more liver tumours were found in diethylnitrosamine carcinogen-challenged C57BL/6 mice12,13 fed with a CDAA or high-fat (HF) diet (Extended Data Fig. 1f–i). These results clearly demonstrate that diet-induced NAFLD enhances HCC in different mouse hepatocarcinogenesis and NAFLD models.
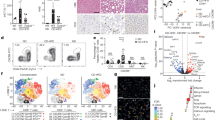
a, Experimental setup. b, Top, representative haematoxylin and eosin (H&E) liver sections. Scale bar, 100 μm. Bottom, representative liver images. Scale bar, 10 mm. c, Liver surface tumour counts. CTR, control diet. n = 10 for CTR, 17 for MCD, P = 0.0067, Student’s t-test. d, e, Intrahepatic CD4+ T lymphocytes (IL CD4+) and intrahepatic CD8+ T lymphocytes (IL CD8+) were measured by flow cytometry. ON, MYC-ON; OFF, MYC-OFF. n = 12 for ON-CTR 4 weeks, 15 for ON-MCD 4 weeks, 6 for OFF-CTR 4weeks, 6 for OFF-MCD 4 weeks, 8 for ON-CTR 8 weeks, 9 for ON-MCD 8 weeks, 6 for OFF-CTR 8 weeks, 6 for OFF-MCD 8 weeks. *P < 0.05, two-way analysis of variance (ANOVA). All data are mean ± standard error of the mean (s.e.m.).
Next, we studied the immune cell subsets in mice with NAFLD and HCC. Consistent with previous reports, dendritic cells, macrophages and CD11b+Gr1+ cells increased (Extended Data Fig. 2a, b)14,15,16. Unexpectedly, significantly fewer CD3hiCD4+ T lymphocytes, which corresponded to conventional intrahepatic CD4+ T lymphocytes, were found in mice with NAFLD (Fig. 1d and Extended Data Fig. 2a–c, e). No significant difference of intrahepatic CD3loCD4+ cells, representing natural killer (NK) T cells (Extended Data Fig. 2d, f), or splenic CD4+ T lymphocytes was observed (Extended Data Fig. 2g). Unlike CD4+ T lymphocytes, intrahepatic CD8+ T lymphocytes remained unchanged (Fig. 1e and Extended Data Fig. 2a, b). The liver-specific reduction of CD4+ but not CD8+ T lymphocytes was also observed in the two other dietary NAFLD models in both tumour-free and tumour-bearing settings (Extended Data Fig. 2h–t), illustrating a tumour-independent but NAFLD-dependent mechanism. In addition, fewer CD4+ but not CD8+ T lymphocytes were found in leptin-deficient (ob/ob) mice (Extended Data Fig. 2u–x).
CD4+ T lymphocytes in NAFLD mice were characterized. Higher levels of CD69 and CD44hiCD62lo subsets were found in MYC-ON MCD mice (Extended Data Fig. 3a–d). Hepatic but not splenic CD4+ T lymphocytes also consistently produced more interferon (IFN)-γ but not interleukin (IL)-4 (Extended Data Fig. 3e–g). Although T-bet, GATA3 and Foxp3 frequency did not change, more ROR-γt was detected in MYC-ON MCD mice (Extended Data Fig. 3h). Accordingly, intrahepatic CD4+ T lymphocytes produced more IL-17 in NAFLD (Extended Data Fig. 3i, j). No change of regulatory T lymphocyte (Treg) frequency was found, and absolute numbers decreased (Extended Data Fig. 3h, k), consistent with a previous report17. In addition, the Treg function remained unchanged in NAFLD (Extended Data Fig. 3l). Together, our results indicate that NAFLD caused activation of hepatic CD4+ T lymphocytes in mice.
CD4+ T lymphocytes have been reported to inhibit HCC initiation and mediate tumour regression18,19. It has also been reported that a considerable fraction of non-synonymous cancer mutations is immunogenic and that the majority of the immunogenic mutanome is recognized by CD4+ T lymphocytes20. Therefore, we studied tumour-specific CD4+ T lymphocytes. α-Fetoprotein (AFP)-specific CD4+ T lymphocytes in MYC-ON MCD mice were detected, suggesting that MYC tumours induced anti-tumour CD4+ T lymphocyte responses (Extended Data Fig. 3m). Next, we depleted CD4+ T lymphocytes to study their relevance to tumour growth. CD4 antibody depletion (Extended Data Fig. 3n) caused more hepatic tumour lesions in MYC-ON MCD mice (Fig. 2a–c). In control-diet-fed MYC mice, CD4 antibody depletion also promoted tumours but at a later time point (Extended Data Fig. 3o, p). These results suggested that loss of CD4+ T lymphocytes strongly contributed to HCC development in MYC-ON mice.
a, Experimental setup. i.p., intraperitoneal. b, c, Representative H&E staining images and microscopic tumour counts. Scale bar, 200 μm. Data are mean ± s.e.m.; n = 5 for IgG, 8 for anti-CD4. *P < 0.05, Student’s t-test.
Next, we studied CD4+ T lymphocyte survival in mice with NAFLD, and a higher annexin V+ level was found in MYC-ON MCD mice than that in MYC-ON mice fed with control diet (Fig. 3a and Extended Data Fig. 4a). We hypothesized that the extensive hepatic lipid accumulation induced the CD4+ T lymphocyte loss. Hepatocytes isolated from MYC-ON MCD mice, which showed accumulation of lipid droplets (Extended Data Fig. 4b), or control mice, were co-cultured with splenocytes. Interestingly, a significant increase in annexin V+7AAD+ cells was seen in CD4+ but not CD8+ T lymphocytes (Extended Data Fig. 4c–e). No cell-to-cell contact was required (Fig. 3b). Higher lipid levels were detected in hepatic CD4+ T lymphocytes in MCD mice (Extended Data Fig. 4f, g). This prompted us to examine whether lipids released from lipid-laden hepatocytes were taken up by CD4+ T lymphocytes and caused cell death. To test this hypothesis, we first measured the hepatic free fatty acid (FFA) composition. Consistent with previous reports, palmitic acid (C16:0), stearic acid (C18:0), linoleic acid (C18:2), arachidonic acid (C20:4) and docosahexaenoic acid (C22:6) are the abundant FFAs (Fig. 3c and Extended Data Table 1). Although the total amount of FFAs did not change significantly (Extended Data Table 1), the levels of C16:0 and C22:6 decreased. C18:2 was the only abundant FFA, which accumulated in the liver after MCD treatment (Fig. 3c). Our data are supported by previous reports of hepatic C18:2 accumulation in HF-diet-induced NAFLD mice and ob/ob mice8,21. Next, we depleted FFAs from conditioned hepatocyte-culture medium. As expected, FFA-depleted conditioned medium no longer caused CD4+ T lymphocyte death (Fig. 3d). FFAs in conditioned medium from lipid-laden hepatocytes were further analysed, and C16:0, C18:0 and C18:2 were identified as predominant FFAs (Extended Data Fig. 4h, i).
a, Ex vivo cell death of intrahepatic CD4+ T lymphocytes from MYC-ON NAFLD mice (n = 7 for CTR, 9 for MCD, Student’s t-test). b, Lymphocyte survival after incubation with hepatocyte conditioned medium (CM) (n = 4, two-way ANOVA). c, Hepatic total FFA composition analysis (n = 6, *P < 0.05, ANOVA). d, FFA depletion (Dep) from conditioned medium (n = 3 for none, 9 for other treatments; two-way ANOVA). e, f, Lymphocyte survival after FFA treatment (n = 4, one-way ANOVA). g, h, CD4+ and CD8+ T lymphocytes in high- or low-C18:2-diet-fed mice (n = 5, Student’s t-test). All data are mean ± s.e.m. *P < 0.05.
Then, isolated CD4+ T lymphocytes were incubated with individual FFAs to study their effect on cell survival. Comparing other FFAs, C18:2 treatment caused a substantially higher level of 7AAD+annexin V+ cells. (Fig. 3e). Unlike CD4+ T lymphocytes, cell death in CD8+ T lymphocytes was not affected at the tested concentration (Fig. 3f). Similar results were found in activated T lymphocytes (Extended Data Fig. 4j). Dose–response and time-course analysis confirmed that CD4+ T lymphocytes were more susceptible to C18:2-induced cell death than CD8+ T lymphocytes (Extended Data Fig. 4k–m). The increase of caspase 3/7 activity confirmed that CD4+ T lymphocytes died through apoptosis (Extended Data Fig. 4n). Similar cell death rates between CD4+ and CD8+ T lymphocytes were observed in an H2O2-induced cell death model, showing that the effect is specific to C18:2 (Extended Data Fig. 4o). Interestingly, mice fed with a high C18:2 diet showed a reduction in CD4+ but not CD8+ T lymphocytes (Fig. 3g, h) suggesting that C18:2 is sufficient to cause CD4+ T lymphocyte death in vivo.
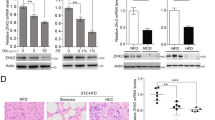
The mechanism of how C18:2 induced CD4+ T lymphocyte death was studied. No difference in cellular C18:2 uptake by CD4+ versus CD8+ T lymphocytes was found (Extended Data Fig. 4p). Direct assessment revealed greater mitochondrial mass in CD4+ lymphocytes (Fig. 4a). Microarray analysis revealed that oxidative phosphorylation and mitochondrial dysfunction pathways were specifically altered in CD4+ but not CD8+ T lymphocytes after C18:2 treatment (Extended Data Fig. 5). CPT1a, the rate-limiting enzyme for importing FFAs into mitochondria, increased in parallel with the decrease of a number of genes coding components for the electron transport complex (Extended Data Fig. 6a). C18:2 was more potent than other FFAs in upregulating CPT1a (Fig. 4b). A similar effect was observed in Jurkat cells, a human CD4+-derived T leukaemia cell line (Extended Data Fig. 6b). Knockdown of CPT1a rescued Jurkat cells from C18:2-induced cell death (Fig. 4c and Extended Data Fig. 6c). All these results pointed towards mitochondria as the critical mediator for CD4+ T lymphocyte death. Inside mitochondria, FFAs are β-oxidized to fuel ATP generation via the electron transport chain (ETC). Fatty acid oxidation (FAO) was measured, and C18:2 showed a greater FAO rate than C16:0 (Fig. 4d). Higher FAO favours more NADH entering the ETC to generate ATP. However, our array data also suggested that C18:2 impaired ETC function (Extended Data Fig. 6a). Indeed, mitochondrial membrane potential, which is maintained by proper ETC activity, was significantly decreased by C18:2 in CD4+ but not CD8+ T lymphocytes (Fig. 4e). A disrupted ETC can become a major site of premature electron leakage to oxygen to generate ROS and lead to cell death22.
a, Mitochondrial mass analysis. b, Cpt1a mRNA levels in FFA-treated CD4+ T lymphocytes (n = 6). c, CPT1a knockdown on C18:2-induced Jurkat cell death (n = 6). shRNA, short hairpin RNA. NT, non-targeting control. d, Oxidation rate of C18:2 or C16:0 in lymphocytes (n = 3). CPM, counts per minutes. e, Mitochondrial membrane potential in C18:2-treated lymphocytes. TMRM, tetramethylrhodamine, methyl ester. f, g, OCR assay of activated CD4+ and CD8+ T lymphocytes treated with FFAs (n = 8). CQ, arbitrary unit obtained by CyQUANT cell proliferation assay (see Methods). h, Ex vivo ROS levels of intrahepatic CD4+ T lymphocytes (n = 6 for CTR, 8 for MCD). DCFDA, 2′,7′-dichlorofluorescin diacetate. MFI, mean fluorescence intensity. i, Mitochondrial ROS levels in lymphocytes. j, Effect of NAC or catalase on hepatocyte-caused lymphocyte death (n = 7). Hep, hepatocytes. k, l, In vivo effect of NAC treatment on intrahepatic CD4+ T lymphocytes and tumour development (n = 3 for CTR, 4 for MCD, 10 for MCD+NAC, 5 for MCD+NAC+anti-CD4). m–o, MitoTEMPO treatment, mitochondrial ROS and survival in CD4+ T lymphocytes in vitro and in vivo (n = 4 for CTR, 4 for MCD+PBS, 5 for MCD+MitoTEMPO). p, Human lymphocyte survival after FFA treatment (n = 6). q, CD4/CD8 ratio of intrahepatic T lymphocytes in patient biopsies (n = 6 for normal, 16 for NASH, 8 for ASH, 15 for hepatitis B virus (HBV)/hepatitis C virus (HCV)). All data are mean ± s.e.m. *P < 0.05, one-way or two-way ANOVA analysis was used.
To assess mitochondrial respiration, oxygen consumption analysis was performed. Normalized oxygen consumption rates (OCRs) were significantly higher in CD4+ T lymphocytes compared with CD8+ T lymphocytes, consistent with previous reports23. Treatment with oligomycin, an inhibitor of mitochondrial ATP synthase, revealed substantial levels of ATP-synthase-dependent oxygen consumption in both CD4+ and CD8+ T lymphocytes (Fig. 4f and Extended Data Fig. 6d). C18:2 abrogated the oligomycin-sensitive fraction of the OCR in CD4+ and CD8+ T lymphocytes without reducing total oxygen consumption levels (Fig. 4f and Extended Data Fig. 6d). These data are consistent with a shift in oxygen consumption from ATP-synthase-dependent to ATP-synthase-independent ROS production. In contrast, C16:0 failed to eliminate ATP-synthase-dependent oxygen consumption in CD4+ T lymphocytes (Fig. 4g and Extended Data Fig. 6e). Consistently, increased total ROS production was found in CD4+ T lymphocytes when co-cultured with C18:2 versus C16:0 (Extended Data Fig. 6f). Moreover, elevated ROS levels were detected in hepatic CD4+ T lymphocytes ex vivo under NAFLD conditions (Fig. 4h) Finally, mitochondrial superoxide was confirmed to increase selectively in CD4+ T lymphocytes after C18:2 treatment (Fig. 4i), and CPT1a knockdown blocked C18:2-induced mitochondrial ROS production (Extended Data Fig. 6g). Taken together, these data suggest that greater levels of mitochondrial-derived ROS accumulate in CD4+ T lymphocytes after C18:2 treatment, leading to their depletion.
Therefore, we tested the role of ROS in NAFLD-associated CD4+ T lymphocyte death and HCC development in vivo. Blocking ROS with catalase or N-acetylcysteine (NAC) abrogated cell death in vitro in CD4+ T lymphocytes when incubated with hepatocytes from MCD-ON MYC mice (Fig. 4j). Similarly, catalase and NAC prevented C18:2-induced CD4+ T lymphocyte death in vitro (Extended Data Fig. 6h). Oxidative stress is an important factor in NAFLD progression24. To test whether ROS mediate hepatic CD4+ T lymphocyte loss in vivo, we treated MCD-diet-fed mice with NAC. Although NAC treatment did not influence steatosis (Extended Data Fig. 6i, j), it effectively reversed the loss of hepatic CD4+ T lymphocytes (Fig. 4k). More importantly, NAC treatment significantly delayed NAFLD-promoted tumour development (Fig. 4l and Extended Data Fig. 6j). Tumour lesions occurred despite NAC treatment when CD4+ T lymphocytes were removed, suggesting that prevention of CD4+ T lymphocyte death mediates at least partially the anti-tumour effect of NAC. Similar results were obtained using mitoTEMPO25, a specific mitochondrial antioxidant in both in vitro and in vivo settings (Fig. 4m–o).
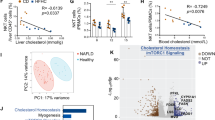
C18:2 has also been identified as an important fatty acid in the context of NAFLD in humans26,27. We tested whether C18:2 also affects human CD4+ T lymphocyte survival. Consistent with our mouse data, C18:2, but no other tested FFAs, caused selective CD4+ but not CD8+ T lymphocyte death (Fig. 4p and Extended Data Fig. 7a). Similarly, C18:2 but not C16:0 increased the ROS level in human CD4+ T lymphocytes (Extended Data Fig. 7b). Finally, intrahepatic CD4+ T lymphocytes in liver biopsies from patients with non-alcoholic steatohepatitis (NASH), alcoholic steatohepatitis (ASH) and viral hepatitis were determined (Extended Data Table 2). While ALT and AST levels did not differ among patients with different liver diseases (Extended Data Fig. 7c, d), fewer CD4+ T lymphocytes were found in NASH and ASH patients than in viral hepatitis patients (Extended Data Figs 7e and 8), and the CD4/CD8 ratio was significantly lower in NASH patients, supporting the idea of selective CD4+ T lymphocyte loss (Fig. 4q). Interestingly, lower CD4 counts were also found in ASH patients; ASH has very similar histological features to NASH.
Dysregulation of lipid metabolism and accumulation of lipids in the liver is part of the aetiology of NAFLD. So far, NASH has been described as causing NF-κB dysregulation, activation of the inflammasome, Toll-like receptor activation and affecting innate immune responses through multiple pathways or directly affecting hepatocytes28,29,30. Our results extend these findings by describing a novel link between obesity-induced lipid accumulation and selective CD4+ T lymphocyte loss, suggest a critical role for CD4+ T lymphocytes in the disease progression from NAFLD to HCC.
Methods
Mouse studies
LAP-tTA and TRE-MYC mice were previously described and MYC expression in the liver was activated by removing doxycycline treatment (100 μg ml−1) from the drinking water of 4-week-old double transgenic mice for both TRE-MYC and LAP-tTA as previously described9,13. C57BL/6 mice were obtained from NCI Frederick. Chemically induced HCC was established by intraperitoneal injection of diethylnitrosoamine (DEN) (Sigma) into 2-week-old male pups at a dose of 20 μg g−1 body weight13. Twelve-week-old male B6.Cg-Lepob/J (ob/ob) mice or wild-type control mice were obtained from Charles River. Foxp3–GFP mice were previously described31. NAFLD was induced by feeding mice with a methionine–choline-deficient (MCD) diet (catalogue number 960439, MP biomedical), a choline-deficient and amino-acid-defined (CDAA) diet (catalogue number 518753, Dyets) or a high-fat diet (catalogue number F3282, Bio Serv) for the indicated time10,11,32. The MCD diet was supplied with corn oil (10%, w/w), and no fish oil was added. Control diet was purchased from MP Biomedical (catalogue number 960441). Custom-made high- or low-linoleic-acid mouse diets were purchased from Research Diets. The modified diets were based on AIN-76A standard mouse diet, and are isocaloric (4.45 kcal g−1) and contained the same high-fat content (23%, w/w). Linoleic-acid-rich safflower oil and saturated fatty-acid-containing coconut oil were supplied at different ratios to yield 2% (w/w) for the low-linoleic-acid diet or 12% (w/w) for the high-linoleic-acid diet. C57BL/6 mice were fed with the high- or low-linoleic-acid diet for 4 weeks. MYC mice were injected i.p. with 50 μg CD4 antibody (clone GK1.5, BioXcell) every week for the indicated time period to deplete CD4+ T cells33. N-acetylcysteine (NAC) was given in drinking water (10 mg ml−1)34 for the indicated time period to prevent excess ROS production. Mitochondrial-specific antioxidant mitoTEMPO was purchased from Sigma. Mice received mitoTEMPO at a dose of 0.7 mg kg−1 per day25 by osmotic minipumps (ALZET). At the experimental end points, mice were killed. For flow cytometry analysis, single-cell suspensions were prepared from spleen, liver and blood as described previously. Red blood cells were lysed by ACK Lysis Buffer (Quality Biologicals). Parts of live tissue were fixed by 10% formaldehyde and subjected to H&E staining. Free fatty acids were purchased from Sigma.
Oil Red O staining
Lipid accumulation was detected by Oil Red O staining in frozen liver sections using the custom service of Histo Serv.
Flow cytometry
Cells were surface-labelled with the indicated antibodies for 15 min at 4 °C. Flow cytometry was performed on BD FACSCalibur or BD LSRII platforms and results were analysed using FlowJo software version 9.3.1.2 (TreeStar). The following antibodies were used for flow cytometry analysis: anti-CD3-FITC (clone 17A2, BD Pharmingen), anti-CD4-PE (clone RM4–4, Biolegend), anti-CD4-APC (clone RM4–5, eBioscience), anti-CD8-Alexa Fluor 700 (clone 53–6.7 Biolegend), anti-CD45, anti-CD44-PE (clone IM7, eBioscience), anti-CD62L-PerCP/Cy5.5 (MEL-14, Biolegend), anti-CD69-Pacific blue (clone H1.2F3, Biolegend), PBS57/CD1d-tetramer-APC (NIH core facility). To determine cytokine production, cells were stimulated with PMA and ionomycine for 30 min, and then were fixed and permeabilized using cytofix/cytoperm kit (BD Pharmingen) followed by anti-IFN-γ-PE (clone XMG1.2, BD Pharmingen), anti-IL-17-PerCP/Cy5.5 (clone TC11-18H10.1, Biolegend) staining. Cell death and apoptosis were detected with annexin V-PE (BD Pharmingen) and 7-AAD (BD Pharmingen) staining according to the manufacturer’s instructions. Intrahepatic CD4+ lymphocytes were gated on the CD3hiCD4+ population from total live hepatic infiltrating mononuclear cells. Absolute numbers were calculated by multiplying frequencies obtained from flow by total live mononuclear cell count, then divided by liver weight. The antibodies used for human peripheral blood mononuclear cell (PBMC) staining are the following: anti-CD3-PE (clone SK7, BD Pharmingen), anti-CD4-FITC (clone RPA-T4, BD Pharmingen), anti-CD8-APC (clone RPA-T8, BD Pharmingen).
Treg suppressive function assay
Murine Treg assays were performed as described31. Briefly, liver Treg cells were isolated as CD4+GFP+ by flow-cytometry-assisted cell sorting from Foxp3–GFP mice kept on an MCD or control diet for 4 weeks. CD4+GFP− T effector (Teff) cells (5 × 104) were stimulated for 72 h in the presence of irradiated T-depleted splenocytes (5 × 104) plus CD3ε monoclonal antibody (1 μg ml−1), with or without Treg cells added at different ratios. 3H-Thymidine was added to the culture for the last 6 h and incorporated radioactivity was measured.
AFP-specific T-cell response
Freshly isolated splenocytes from MYC-ON MCD mice were incubated with 5 μg ml−1 of mouse α-fetoprotein protein (MyBioSource) for 24 h. Golgiplug was added for the last 6 h. Then, cells were fixed and permeabilized using cytofix/cytoperm kit (BD Pharmingen) followed by anti-IFN-γ-PE (clone XMG1.2, BD Pharmingen) staining.
Hepatocyte isolation
Primary mouse hepatocytes were isolated from MYC mice and cultured according to a previous report35. Briefly, mice were anaesthetized and the portal vein was cannulated under aseptic conditions. The livers were perfused with EGTA solution (5.4 mM KCl, 0.44 mM KH2PO4, 140 mM NaCl, 0.34 mM Na2HPO4, 0.5 mM EGTA, 25 mM Tricine, pH 7.2) and Gey’s balanced salt solution (Sigma), and digested with 0.075% collagenase solution. The isolated mouse hepatocytes were then cultured with complete RPMI media in collagen-I-coated plates.
Hepatic fatty acid profiling
Hepatic fatty acid composition was measured at LIPID MAPS lipidomics core at the University of California (San Diego) using an esterified and non-esterified (total) fatty acid panel. Briefly, liver tissues were homogenized and lipid fraction was extracted using a modified Bligh Dyer liquid/liquid extraction method. The lipids were saponified and the hydrolysed fatty acids were extracted using a liquid/liquid method. The extracted fatty acids were derivatized using pentaflourylbenzylbromine (PFBB) and analysed by gas chromatography (GC) using an Agilent GC/mass spectrometry (MS) ChemStation. Individual analytes were monitored using selective ion monitoring (SIM). Analytes were monitored by peak area and quantified using the isotope dilution method using a deuterated internal standard and a standard curve.
Free fatty acid identification
Isolated primary hepatocytes from MYC mice fed with MCD or control diet were cultured in complete RPMI for 24 h. Supernatant were harvested and FFAs were identified by GC/MS.
Microarray analysis
Splenocytes from MYC mice were cultured with or without 50 μM C18:2 for 24 h. CD4+ and CD8+ T lymphocytes were sorted and total RNA was extracted using miRNeasy mini kit (Qiagen). Array analysis was performed in the Department of Transfusion Medicine, clinical centre at NIH. Mouse gene 2.0 ST array (Affymetrix) was used and performed according to the manufacturer’s instruction. Data were log-transformed (base 2) for subsequent statistical analysis. The Partek Genomic Suite 6.4 was used for the identification of differentially expressed transcripts. The Ingenuity Pathway Analysis tool (http://www.ingenuity.com) was used for analysis of functional pathways.
RNA isolation and real-time PCR
RNA was extracted from frozen tissues with RNeasyMini Kit (Qiagen). Complementary DNA was synthesized by iScriptcDNA synthesis kit (BioRad). Sequence of primers used for quantitative RT–PCR can be obtained from the authors. The reactions were run in triplicates using iQSYBR green supermix kit (BioRad). The results were normalized to endogenous GAPDH expression levels.
CD4+ T-cell isolation and co-culture with fatty acids
CD4+ T lymphocytes were isolated from the spleen of MYC mice by negative autoMACS selection using a CD4+ T lymphocytes isolation kit (Miltenyi Biotec) or flow cytometry cell sorting. Human CD4+ T lymphocytes were prepared from PBMCs by autoMACS using a CD4+ T lymphocytes isolation kit (Miltenyi Biotec). The purity of CD4+ T lymphocytes was above 90% after autoMACS separation and above 95% after flow cytometry cell sorting. C16:0, C18:0, C18:1,and C18:2 were purchased from Sigma. Fatty acids were dissolved in DMEM with 2% fatty-acid-free bovine serum albumin (BSA; Sigma, catalogue number A8806) after solvent was evaporated, then followed by two rounds of vortexing and 30 s of sonication. Isolated CD4+ T lymphocytes or splenocytes were incubated with different fatty acids or conditioned medium from hepatocyte culture for 3 days. Unless specifically described, fatty acids were used at 50 μM concentration. For fatty acid depletion, active charcoal (catalogue number C-170, Fisher) was used as described before36. Briefly, 0.5 g of active charcoal was added into every 10 ml of conditioned medium. Then pH was lowered to 3.0 by addition of 0.2 N HCl. The solution was rotated at 4 °C for 2 h. Charcoal was then removed by centrifugation, and the clarified solution was brought back to pH 7.0 by addition of 0.2 N NaOH. NAC (10 mM), catalase (1,000 U ml−1) or mitoTEMPO (10 μM) was used to inhibit ROS production, mitochondrial ROS levels were determined by mitoSOX staining 24 h after treatment, cell death and apoptosis were measured by annexin V and 7-AAD staining 3 days after treatment.
Caspase activity assay
Caspase activity assay was measured by caspase-Glo 3/7 assay kit (Promega) according to the manufacturer’s protocol.
BODIPY staining
Fresh prepared liver-infiltrating mononuclear cells were washed and resuspended in 500 μl of BODIPY 493/503 at 0.5 μg ml−1 in PBS. Cells were stained for 15 min at room temperature. Then cells were subjected to flow cytometry analysis.
RNA interference assay
Two pZIP lentiviral shRNA vectors targeting human CPT1a and a control vector (NT#4) were purchased from TransOMIC Technologies. Lentivirus was packed in 293T cells. Jurkat cells were purchased from the German Collection of Microorganisms and Cell Cultures (DSMZ), and no authentication test was performed by us. Cells were cultured in complete RPMI medium and were tested to be mycoplasma free. Jurkat cells were infected with shRNA lentivirus. Puromycin was added to eliminate non-transduced cells. Doxycycline (100 ng ml−1) was added to induce shRNA and GFP expression for 3 days. Efficiency of shRNAs was confirmed by western blot. Jurkat cells with CPT1a knockdown were treated with 200 μM C18:2 for 24 h. Mitochondrial ROS production and cell survival were measured in GFP+-transduced cells.
Fatty acid oxidation assay
Fatty acid oxidation was measured according to a previous publication37. 1-14C-C18:2 and 1-14C-C16:0 were purchased from PerkinElmer. Briefly, isolated CD4+ or CD8+ T lymphocytes were pretreated with C18:2 or kept in regular media. After 24 h, cell media was changed to media containing 50 μM cold C18:2 plus 1 μCi 1-14C-C18:2 per ml or 50 μM cold C16:0 plus 1 μCi 1-14C-C16:0 per ml. After 2 h, medium was removed and mixed with concentrated perchloric acid (final concentration 0.3 M) plus BSA (final concentration 2%) to precipitate the radiolabelled fatty acids. Samples were vortexed and centrifuged (10,000g for 10 min). Radioactivity was determined in the supernatant to measure water-soluble β-oxidation products.
Mitochondrial membrane potential and ROS staining
Mitochondrial membrane potential was measured by TMRM (ImmunoChemistry Technologies) staining according to the manufacturer’s protocol. Briefly, cells were kept in culture medium with 100 nM of TMRM for 20 min in a CO2 incubator at 37 °C. After washing twice, cells were processed to flow cytometry analysis. Mitochondria-associated superoxide was detected by mitoSOX (Life Technologies) staining according to the manufacturer’s protocol. Briefly, cells were first subjected to surface marker staining. Then cells were stained with 2.5 μM mitoSOX for 30 min in a CO2 incubator at 37 °C. After washing twice, cells were processed for flow cytometry analysis.
Oxygen consumption assay
OCR was measured using an XFe96 Extracellular Flux Analyzer (Seahorse Bioscience) as previously described38. AutoMACS-sorted mouse CD4+ and CD8+ T lymphocytes were attached to XFe96 cell culture plates using Cell-Tak (BD Bioscience) in RPMI media with 11 mM glucose. Cells were activated with 1:1 CD3:CD28 beads (Miltenyi BioTech) and vehicle or 50 μM C18:2 was added. Twenty-four hours after activation, cells were incubated in serum-free XF Base Media (Seahorse Bioscience) supplemented with 10 mM glucose, 2 mM pyruvate and 2 μM glutamine, pH 7.4, along with 50 μM C18:2 if previously present, for 30 min at 37 °C in a CO2-free cell culture incubator before beginning the assay. Five consecutive measurements, each representing the mean of 8 wells, were obtained at baseline and after sequential addition of 1.25 μM oligomycin, 0.25 μM trifluorocarbonylcyanide phenylhydrazone (FCCP), and 1 μM each of rotenone and antimycin A (all drugs from Seahorse Bioscience). OCR values were normalized to cell number as measured by the CyQUANT Cell Proliferation Assay Kit (Life Technologies).
Human studies
Human liver samples were stained as previously described8. For immunostaining, formalin-fixed, paraffin-embedded human liver tissue samples were retrieved from the archives of the Institute of Surgical Pathology, University Hospital Zurich. Fibrosis grade was analysed for NASH according to NAFLD activity score (NAS)39 and for others according to METAVIR score40. The study was approved by the local ethics committee (Kantonale Ethikkommission Zürich, application number KEK-ZH-Nr. 2013-0382). Human PBMCs from healthy donors were obtained on an NIH-approved protocol and prepared as described previously41. Informed consent was obtained from all subjects.
Statistical analysis
The sample sizes for animal studies were guided by a previous study in our laboratory in which the same MYC transgenic mouse stain was used. No animals were excluded. Neither randomization nor blinding were done during the in vivo study. However, mice from the same littermates were evenly distributed into control or treatment groups whenever possible. The sample size for the patient studies was guided by a recent publication also studying NASH-induced HCC, but focused on different aspects8. Statistical analysis was performed with GraphPad Prism 6 (GraphPad Software). Significance of the difference between groups was calculated by Student’s unpaired t-test, one-way or two-way ANOVA (Tukey’s and Bonferroni’s multiple comparison test). Welch’s corrections were used when variances between groups were unequal. P < 0.05 was considered as statistically significant.
References
European Association For The Study Of The Liver & European Organisation For Research And Treatment Of Cancer. EASL-EORTC clinical practice guidelines: management of hepatocellular carcinoma. J. Hepatol. 56, 908–943 (2012)
Sun, B. & Karin, M. Obesity, inflammation, and liver cancer. J. Hepatol. 56, 704–713 (2012)
Michelotti, G. A., Machado, M. V. & Diehl, A. M. NAFLD, NASH and liver cancer. Nature Rev. Gastroenterol. Hepatol. 10, 656–665 (2013)
Schuppan, D. & Schattenberg, J. M. Non-alcoholic steatohepatitis: pathogenesis and novel therapeutic approaches. J. Gastroenterol. Hepatol. 28 (suppl. 1), 68–76 (2013)
Wree, A., Broderick, L., Canbay, A., Hoffman, H. M. & Feldstein, A. E. From NAFLD to NASH to cirrhosis—new insights into disease mechanisms. Nature Rev. Gastroenterol. Hepatol. 10, 627–636 (2013)
Greten, T. F., Wang, X. W. & Korangy, F. Current concepts of immune based treatments for patients with HCC: from basic science to novel treatment approaches. Gut 64, 842–848 (2015)
Greten, T. F., Duffy, A. G. & Korangy, F. Hepatocellular carcinoma from an immunologic perspective. Clin. Cancer Res. 19, 6678–6685 (2013)
Wolf, M. J. et al. Metabolic activation of intrahepatic CD8+ T cells and NKT cells causes nonalcoholic steatohepatitis and liver cancer via cross-talk with hepatocytes. Cancer Cell 26, 549–564 (2014)
Shachaf, C. M. et al. MYC inactivation uncovers pluripotent differentiation and tumour dormancy in hepatocellular cancer. Nature 431, 1112–1117 (2004)
Rinella, M. E. et al. Mechanisms of hepatic steatosis in mice fed a lipogenic methionine choline-deficient diet. J. Lipid Res. 49, 1068–1076 (2008)
Yang, L. et al. Transforming growth factor β signaling in hepatocytes participates in steatohepatitis through regulation of cell death and lipid metabolism in mice. Hepatology 59, 483–495 (2014)
Park, E. J. et al. Dietary and genetic obesity promote liver inflammation and tumorigenesis by enhancing IL-6 and TNF expression. Cell 140, 197–208 (2010)
Kapanadze, T. et al. Regulation of accumulation and function of myeloid derived suppressor cells in different murine models of hepatocellular carcinoma. J. Hepatol. 59, 1007–1013 (2013)
Xia, S. et al. Gr-1+ CD11b+ myeloid-derived suppressor cells suppress inflammation and promote insulin sensitivity in obesity. J. Biol. Chem. 286, 23591–23599 (2011)
Baeck, C. et al. Pharmacological inhibition of the chemokine CCL2 (MCP-1) diminishes liver macrophage infiltration and steatohepatitis in chronic hepatic injury. Gut 61, 416–426 (2012)
Henning, J. R. et al. Dendritic cells limit fibroinflammatory injury in nonalcoholic steatohepatitis in mice. Hepatology 58, 589–602 (2013)
Ma, X. et al. A high-fat diet and regulatory T cells influence susceptibility to endotoxin-induced liver injury. Hepatology 46, 1519–1529 (2007)
Rakhra, K. et al. CD4+ T cells contribute to the remodeling of the microenvironment required for sustained tumor regression upon oncogene inactivation. Cancer Cell 18, 485–498 (2010)
Kang, T. W. et al. Senescence surveillance of pre-malignant hepatocytes limits liver cancer development. Nature 479, 547–551 (2011)
Kreiter, S. et al. Mutant MHC class II epitopes drive therapeutic immune responses to cancer. Nature 520, 692–696 (2015)
Giesbertz, P. et al. Metabolite profiling in plasma and tissues of ob/ob and db/db mice identifies novel markers of obesity and type 2 diabetes. Diabetologia 58, 2133–2143 (2015)
Schönfeld, P. & Wojtczak, L. Fatty acids as modulators of the cellular production of reactive oxygen species. Free Radic. Biol. Med. 45, 231–241 (2008)
Cao, Y., Rathmell, J. C. & Macintyre, A. N. Metabolic reprogramming towards aerobic glycolysis correlates with greater proliferative ability and resistance to metabolic inhibition in CD8 versus CD4 T cells. PLoS ONE 9, e104104 (2014)
Sumida, Y., Niki, E., Naito, Y. & Yoshikawa, T. Involvement of free radicals and oxidative stress in NAFLD/NASH. Free Radic. Res. 47, 869–880 (2013)
Porporato, P. E. et al. A mitochondrial switch promotes tumor metastasis. Cell Reports 8, 754–766 (2014)
Feldstein, A. E. et al. Mass spectrometric profiling of oxidized lipid products in human nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. J. Lipid Res. 51, 3046–3054 (2010)
Muir, K. et al. Proteomic and lipidomic signatures of lipid metabolism in NASH-associated hepatocellular carcinoma. Cancer Res. 73, 4722–4731 (2013)
Beraza, N. et al. Pharmacological IKK2 inhibition blocks liver steatosis and initiation of non-alcoholic steatohepatitis. Gut 57, 655–663 (2008)
Henao-Mejia, J. et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 482, 179–185 (2012)
Miura, K. et al. Toll-like receptor 2 and palmitic acid cooperatively contribute to the development of nonalcoholic steatohepatitis through inflammasome activation in mice. Hepatology 57, 577–589 (2013)
Andersson, J. et al. CD4+ FoxP3+ regulatory T cells confer infectious tolerance in a TGF-β-dependent manner. J. Exp. Med. 205, 1975–1981 (2008)
Nakagawa, H. et al. ER stress cooperates with hypernutrition to trigger TNF-dependent spontaneous HCC development. Cancer Cell 26, 331–343 (2014)
Ikehara, Y. et al. CD4+ Vα14 natural killer T cells are essential for acceptance of rat islet xenografts in mice. J. Clin. Invest. 105, 1761–1767 (2000)
Hanczko, R. et al. Prevention of hepatocarcinogenesis and increased susceptibility to acetaminophen-induced liver failure in transaldolase-deficient mice by N-acetylcysteine. J. Clin. Invest. 119, 1546–1557 (2009)
Radaeva, S. et al. Interferon-α activates multiple STAT signals and down-regulates c-Met in primary human hepatocytes. Gastroenterology 122, 1020–1034 (2002)
Chen, R. F. Removal of fatty acids from serum albumin by charcoal treatment. J. Biol. Chem. 242, 173–181 (1967)
Huynh, F. K., Green, M. F., Koves, T. R. & Hirschey, M. D. Measurement of fatty acid oxidation rates in animal tissues and cell lines. Methods Enzymol. 542, 391–405 (2014)
Pelletier, M., Billingham, L. K., Ramaswamy, M. & Siegel, R. M. Extracellular flux analysis to monitor glycolytic rates and mitochondrial oxygen consumption. Methods Enzymol. 542, 125–149 (2014)
Kleiner, D. E. et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005)
Bedossa, P. & Poynard, T. An algorithm for the grading of activity in chronic hepatitis C. The METAVIR Cooperative Study Group. Hepatology 24, 289–293 (1996)
Hoechst, B. et al. A new population of myeloid-derived suppressor cells in hepatocellular carcinoma patients induces CD4+CD25+Foxp3+ T cells. Gastroenterology 135, 234–243 (2008)
Acknowledgements
We would like to thank J. Berzofsky, W. Stoffel, E. M. Shevach and S. Thorgeirsson for helpful discussion. A.M.T. was supported by the Intramural Research Program of the National Institutes of Health (NIH), National Institute of Allergy and Infectious Diseases. J.L. was supported by the Intramural Program Grant ZIABC011303, NIH, National Cancer Institute (NCI). M.H. was supported by a European Research Council starting grant (LiverCancerMechanism), the Stiftung Experimentelle Biomedizin (Hofschneider Stiftung), the Pre-clinical Comprehensive Center (PCCC) and the Helmholtz foundation. A.W. was supported by grants from the Krebsliga Schweiz (Oncosuisse) and the Promedica Stiftung, Switzerland. D.E.K., A.H.K., D.W.M. and T.F.G. were supported by the Intramural Research Program of the NIH, NCI.
Author information
Authors and Affiliations
Contributions
C.M. and A.H.K. performed experiments. C.M., A.H.K., D.W.M. and T.F.G. analysed data. T.E., J.M.-E., D.E.K., P.J., D.F.S., M.T., V.K., M.E., M.H., A.M.T., H.Z., J.L. and D.F.W. assisted with experiments and analysis or provided valuable reagents. A. H.K. and D.W.M. performed and analysed seahorse studies. M.E., A.W. and M.H. collected/provided human specimens and performed/analysed immunohistochemistries of human samples. C.M. and T.F.G. conceived of and designed the project. C.M. and T.F.G. wrote the manuscript and all authors contributed to writing and provided feedback.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 MCD, CDAA and HF diets induce NAFLD and promote HCC.
a, Representative imagines of Oil Red O staining of MYC-ON mice fed MCD or CTR. Scale bar, 100 μm. b, Serum ALT levels analysis. Data are mean ± s.e.m.; n = 4, *P < 0.05, one-way ANOVA. c–e, The effect of the CDAA diet on tumour development in MYC transgenic mice. Experimental setup, representative liver images and liver surface tumour counts are shown. Scale bar, 10 mm. Data are mean ± s.e.m.; n = 6 for CDAA and n = 5 mice for CTR, P = 0.0345, Student’s t-test. f–i, The effect of the CDAA and HF diet on liver carcinogenesis in diethylnitrosoamine (DEN)-injected C57BL/6 mice. Experimental setup, representative tumour-free H&E stainings, macroscopic liver images and surface tumour counts are shown. Scale bar, 100 μm. Data are mean ± s.e.m.; n = 13 for CTR, n = 9 for HF, n = 10 for CDAA, *P < 0.05, one-way ANOVA.
Extended Data Figure 2 Immune cell monitoring in NAFLD-HCC.
a–j, MYC mice were fed with an MCD diet or CTR diet. a, b, Intrahepatic immune cells were determined by flow cytometry. Composition (a) and absolute numbers (b) of different intrahepatic immune cell subsets in MYC-ON mice, which were kept for 4 weeks on an MCD diet or CTR diet. Data are mean ± s.e.m.; n ≥ 6, *P < 0.05, one-way ANOVA. c, Representative contour plots of intrahepatic CD4+ T lymphocytes. d, Representative dot plots of CD1d-tetramer staining in CD3loCD4+ population. e–g, Absolute number of intrahepatic CD4+ T lymphocytes, frequencies of NK T cells and splenic CD4+ T lymphocytes were measured by flow cytometry. Data are mean ± s.e.m.; n = 4, *P < 0.05, two-way ANOVA. h–j, Intrahepatic CD4+ and CD8+ T lymphocyte levels in MYC-ON mice fed with a CDAA diet for 16 weeks. Data are mean ± s.e.m.; n = 6 for CDAA and n = 5 for CTR, *P < 0.05, Student’s t-test. k, l, Intrahepatic CD4+ T lymphocyte levels in DEN-injected BL/6 male mice treated with a CDAA diet, HF diet or CTR for 7 months. Data are mean ± s.e.m.; n = 13 for CTR, n = 9 for HF, n = 10 for CDAA, *P < 0.05, one-way ANOVA. m–p, Intrahepatic CD4+ and CD8+ T lymphocytes in tumour-free C57BL/6 mice treated with a CDAA diet for 16 weeks. TF, tumour free. Data are mean ± s.e.m.; n = 3 for CTR, n = 5 for CDAA, *P < 0.05, Student’s t-test. q–t, Intrahepatic CD4+ and CD8+ T lymphocytes in tumour-free C57BL/6 mice treated with an HF or low-fat (LF) diet for 6 months. Data are mean ± s.e.m.; n = 2 for CTR, n = 5 for LF, n = 5 for HF, *P < 0.05, one-way ANOVA. u–x, CD4+ and CD8+ T lymphocytes in 12-week-old male ob/ob or wild-type lean mice. Data are mean ± s.e.m., n = 5, *P < 0.05, Student’s t-test w, x, MYC mice were fed with MCD or CTR. Macrophage and CD11b+Gr1+ populations were measured. Data are mean ± s.e.m.; n ≥ 4, *P < 0.05, two-way ANOVA.
Extended Data Figure 3 Intrahepatic CD4+ lymphocytes are activated in NAFLD, and CD4 depletion enhances HCC.
a–k, MYC-ON mice were fed with MCD or CTR for 4 weeks. a–d, CD69 and CD44hiCD62Llo subsets in intrahepatic CD4+ T lymphocytes were measured. Data are mean ± s.e.m.; n = 8 for MCD and n = 6 for CTR, *P < 0.05, Student’s t-test. e–g. Ex vivo IFN-γ, IL-4 production in intrahepatic CD4+ T lymphocytes were determined. Data are mean ± s.e.m.; n = 8, *P < 0.05, Student’s t-test. h, Ex vivo staining of T-bet, GATA3, ROR-γt and Foxp3 levels in intrahepatic and splenic CD4+ T lymphocytes. Data are mean ± s.e.m.; n = 3, *P < 0.05, two-way ANOVA. i, Ex vivo IL-17 production by intrahepatic CD4+ T lymphocytes. Data are mean ± s.e.m.; n = 5, *P < 0.05, Student’s t-test. j, Representative dot plots of ROR-γt/IL-17 staining in intrahepatic CD4+ T lymphocytes. k, Absolute number of intrahepatic CD4+ lymphocyte subsets. Data are mean ± s.e.m.; n = 3, *P < 0.05, two-way ANOVA. l, Suppressive function assay of isolated hepatic Treg cells from Foxp3–GFP mice kept on MCD or CTR for 4 weeks. m, Detection of AFP-specific CD4+ T lymphocytes in spleen from MYC-MCD mice. n, Selective depletion of intrahepatic CD4+ T lymphocytes but not NK T cells by i.p. injection of 50 μg anti-CD4 antibody (clone GK1.5). o, p, MYC-ON mice on CTR received 50 μg of GK1.5 antibody or isotype control i.p. once per week for 8 weeks. Representative liver imagines and surface tumour counts are shown. Scale bar, 10 mm. Data are mean ± s.e.m., n = 3.
Extended Data Figure 4 Lipid-laden hepatocytes release C18:2 and induce CD4+ T lymphocyte death via apoptosis.
a, Representative contour plots of ex vivo 7AAD/annexin V staining of intrahepatic CD4+ T lymphocytes from MYC-ON mice fed with MCD or CTR. b, Representative phase-contrast images of primary hepatocytes from MYC-ON mice after MCD or CTR treatment. c–e, Isolated primary hepatocytes from MYC-ON mice on MCD or CTR were cocultured with isolated CD4+ T lymphocytes or splenocytes. Cell death levels were measured by flow cytometry. Data are mean ± s.e.m.; n = 4, one-way or two-way ANOVA. f, g, BODIPY 493/503 staining of CD4+ T lymphocytes in liver, spleen or blood from MYC-ON mice with MCD or CTR. Data are mean ± s.e.m.; n = 4, *P < 0.05, two-way ANOVA. h, i, Identification of FFAs in hepatocyte conditioned medium by gas chromatography/mass spectrometry (GC/MS). Data are mean ± s.e.m.; n = 3, *P < 0.05, two-way ANOVA. j, Anti-CD3/28 bead-activated splenocytes were treated with different FFAs, and cell death level in CD4+ or CD8+ T lymphocytes was determined. Data are mean ± s.e.m.; n = 4, *P < 0.05, two-way ANOVA. k–m, Dose–response curve and time course of C18:2-induced cell death in CD4+ or CD8+ T lymphocytes. n, Caspase3/7 activity in CD4+ lymphocytes after C18:2 treatment. Data are mean ± s.e.m.; n = 9, *P < 0.05, Student’s t-test. o, Dose–response curve of H2O2-induced cell death in CD4+ or CD8+ T lymphocytes. p, Uptake of C18:2 by CD4+ and CD8+ T lymphocytes after incubation with 50 μM C18:2 for 2 h. Data are mean ± s.e.m.; n = 6, *P < 0.05, two-way ANOVA.
Extended Data Figure 5 Ingenuity pathway analysis of microarray data.
CD4+ and CD8+ T lymphocytes sorted from C18:2-treated splenocytes were subjected to microarray analysis. Pathway analysis was done by ingenuity pathway analysis (IPA). n = 3. Ratio is the number of changed genes divided by total genes in the pathway.
Extended Data Figure 6 Mitochondrial ROS mediates C18:2-induced CD4+ T lymphocyte death in vitro and in vivo.
a, Real-time polymerase chain reaction (PCR) confirmed the gene changes from microarray. Data are mean ± s.e.m.; n = 3, *P < 0.05, two-way ANOVA. b, Cpt1a mRNA level in Jurkat cells after FFA treatment. Data are mean ± SEM; n = 6, *P < 0.05, one-way ANOVA. c, Expression of CPT1a in wild-type and two knockdown Jurkat cells. NT, none-targeting control. d, e, OCR analysis of activated CD4+ and CD8+ T lymphocytes upon C18:2 or C16:0 incubation. f, ROS levels of CD4+ or CD8+ T lymphocytes in splenocytes treated with C18:2 or C16:0. Data are mean ± s.e.m.; n = 8, *P < 0.05, two-way ANOVA. g, Mitochondrial ROS in wild-type and two CPT1 knockdown Jurkat cells. i, Cell death of CD4+ or CD8+ T lymphocytes in splenocytes treated with C18:2 in the presence of NAC or catalase. Data are mean ± s.e.m.; n = 4, *P < 0.05, two-way ANOVA. i, j, In vivo blocking ROS with NAC in MYC-ON mice treated with MCD. Some mice also received CD4 antibody depletion. Experimental setup and representative H&E liver sections are shown. Scale bar, 200 μm.
Extended Data Figure 7 C18:2 induces cell death in human CD4+ T lymphocytes, and NASH patients have lower intrahepatic CD4+ T lymphocytes.
a, Cell death levels of sorted human CD4+ T lymphocytes treated with different FFAs. Data are mean ± s.e.m.; n = 4, *P < 0.05, one-way ANOVA. b, ROS level of CD4+ or CD8+ T lymphocyte in peripheral blood mononuclear cells (PBMCs) treated with C18:2 or C16:0. Data are mean ± s.e.m.; n = 6, *P < 0.05, two-way ANOVA. c, d, Serum ALT and AST concentration in different patients. e, Intrahepatic CD4+ T lymphocyte count in biopsies. CD4+ T lymphocytes were identified by immunohistochemistry. Data are mean ± s.e.m.; normal = 6, NASH = 16, ASH = 8, HBV/HCV = 16, *P < 0.05, one-way ANOVA.
Extended Data Figure 8 Immunohistochemistry staining of intrahepatic CD4+ or CD8+ T lymphocytes in patient biopsies.
Representative CD4 or CD8 immunohistochemistry images of liver biopsies from healthy individuals, NASH, ASH patients or patients with HBV or HCV. For each condition, two different magnifications are shown. Scale bar, 100 μm.
Rights and permissions
About this article
Cite this article
Ma, C., Kesarwala, A., Eggert, T. et al. NAFLD causes selective CD4+ T lymphocyte loss and promotes hepatocarcinogenesis. Nature 531, 253–257 (2016). https://doi.org/10.1038/nature16969
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16969
This article is cited by
-
A hepatic network of dendritic cells mediates CD4 T cell help outside lymphoid organs
Nature Communications (2024)
-
Blocking methionine catabolism induces senescence and confers vulnerability to GSK3 inhibition in liver cancer
Nature Cancer (2024)
-
The convergent application of metabolites from Avena sativa and gut microbiota to ameliorate non-alcoholic fatty liver disease: a network pharmacology study
Journal of Translational Medicine (2023)
-
Non-alcoholic fatty liver disease risk prediction model and health management strategies for older Chinese adults: a cross-sectional study
Lipids in Health and Disease (2023)
-
Visualization of hydrocarbon chain length and degree of saturation of fatty acids in mouse livers by combining near-infrared hyperspectral imaging and machine learning
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.