Abstract
Mitochondrial electron transport chain complexes are organized into supercomplexes responsible for carrying out cellular respiration. Here we present three architectures of mammalian (ovine) supercomplexes determined by cryo-electron microscopy. We identify two distinct arrangements of supercomplex CICIII2CIV (the respirasome)—a major ‘tight’ form and a minor ‘loose’ form (resolved at the resolution of 5.8 Å and 6.7 Å, respectively), which may represent different stages in supercomplex assembly or disassembly. We have also determined an architecture of supercomplex CICIII2 at 7.8 Å resolution. All observed density can be attributed to the known 80 subunits of the individual complexes, including 132 transmembrane helices. The individual complexes form tight interactions that vary between the architectures, with complex IV subunit COX7a switching contact from complex III to complex I. The arrangement of active sites within the supercomplex may help control reactive oxygen species production. To our knowledge, these are the first complete architectures of the dominant, physiologically relevant state of the electron transport chain.
Similar content being viewed by others
Main
Cellular respiration occurs through the mitochondrial electron transport chain and is responsible for the majority of ATP synthesis in the cell. This process, oxidative phosphorylation, is carried out by five large membrane protein complexes (CI, CII, CIII, CIV and CV) of the inner mitochondrial membrane and two main electron carriers: membrane-bound ubiquinone (Q) and soluble cytochrome c. The proton-pumping complexes (CI, CIII and CIV) convert energy from a series of electron transfers into a proton electrochemical gradient across the inner mitochondria membrane1, harvested by the ATPase (CV) for synthesis of ATP.
Early on, a solid-state model of the electron transport chain, in which each complex existed in a larger particle called the oxysome was proposed2. However, this model was weakened by the isolation of the individual functional complexes3 and the demonstration that mitochondrial electron transport is a diffusion-coupled process4,5, resulting in the proposal of a fluid model, in which all electron transport chain components are in constant and independent diffusional motion5. More recently, higher-order organization of the electron transport chain into supercomplexes of defined stoichiometry has been demonstrated6. The majority of CI is found bound with a CIII dimer and CIV (CICIII2CIV), a structure that contains all complexes required to pass electrons from NADH to O2 and hence termed the ‘respirasome’ (distinct from the oxysome due to its lack of CV), or with a CIII dimer alone (CICIII2)7. Additionally, a CIII dimer forms a supercomplex with CIV (CIII2CIV1) independent of CI (ref. 6). Supercomplex formation is important for the stability of the electron transport chain complexes and for reducing the production of reactive oxygen species (ROS)8,9.
A possible role of supercomplexes in substrate channelling was first suggested from flux control analysis10, but this has remained controversial11,12,13. A proposal for the dynamic regulation of two weakly exchanging pools of ubiquinone in the inner mitochondrial membrane, one associated with supercomplexes and one freely diffusing in the membrane14, was supported by the discovery of the supercomplex assembly factor SCAF1 (also known as COX7a2l)14,15. It has been shown that SCAF1 promotes the interaction between CIII and CIV in both CICIII2CIV and CIII2CIV1 (refs 15, 16, 17).
To date, only low-resolution (~20 Å) electron microscopy maps of supercomplexes have been reported18,19,20. To understand their role better, we sought to obtain higher-resolution cryo-electron microscopy (cryo-EM) structures of supercomplexes.
The respirasome adopts two distinct architectures
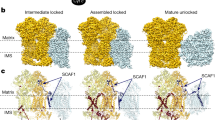
We purified mammalian supercomplexes from Ovis aries (ovine) heart mitochondrial membranes solubilized in the detergent digitonin (Extended Data Fig. 1a). Structures calculated from cryo-EM images revealed two distinct architectures for the respirasome, which we refer to as ‘tight’ and ‘loose’, as well as the architecture of CICIII2 (Extended Data Fig. 1). In all cases, the membrane domains of the complexes are aligned, forming a flat membrane-exposed area, visualized by the bound detergent belt (Extended Data Fig. 2). In both respirasome structures, the density for CIV is weaker relative to CI and CIII, indicating the greater conformational flexibility of CIV. The clearest density for CIV can be seen in the tight respirasome in which CIV contacts both CI and CIII. In the loose respirasome, CIV has well-defined contacts only with CI, and in CICIII2 density for CIV is completely absent (Fig. 1).
a–c, Density for the tight respirasome (a), the loose respirasome (b) and CICIII2 (c) is shown viewed from the CI side (top), the CIII side (middle) and as a slice within the membrane looking from the matrix (bottom). CI density is blue, CIII density is green and CIV density is pink.
This series of architectures may represent independent structural entities or may interconvert, indicating different stages of assembly or disassembly. A subset of data collected after overnight incubation indicated that the loose conformation accumulates slowly in the extracted supercomplexes and so interconversion between the tight and loose particles is possible. This interconversion may result from the loss of cardiolipin, which has been shown to stabilize interactions between CIII and CIV (ref. 21).
In the final maps of the supercomplexes (at 5.8 Å, 6.7 Å and 7.8 Å resolution for the tight, loose and CICIII2 supercomplexes, respectively) α-helical densities are clearly resolved (Fig. 1 and Extended Data Figs 3, 4, 5). For the tight respirasome, β-sheet planes and isolated protein loops are also observed. In all maps, protein density can be clearly differentiated from the detergent belt, which is disordered, with weak fragmented density (Extended Data Fig. 6a, b). Strong peaks can be seen for CI Fe–S clusters in all maps and for the central iron atoms of all haem cofactors in CIII and CIV in the tight respirasome maps (Extended Data Fig. 6c). A peak corresponding to the Fe–S cluster of the CIII Rieske protein (UQCRFS1) is not visible due to a higher level of disorder, consistent with crystal structures lacking substrate or inhibitors22. Nonetheless, UQCRFS1 is clearly in the distal (from the Qp site) position. It has been suggested that UQCRFS1 is fixed in the alternative proximal (to the Qp site) position in reduced CIII, but is mobile in the oxidized complex22. Our data suggest that in the oxidized state UQCRFS1 is not fully mobile, but adopts a conformation close to the distal position.
Using tight respirasome maps and the recent low-resolution cryo-EM model of the isolated bovine enzyme23, an improved model for CI was built (Extended Data Fig. 7). We extended many of the previously assigned subunits and assigned three additional supernumerary subunits.
All observed protein density in both respirasome architectures can be attributed to the known 80 subunits of the individual complexes (45 from CI, 22 from the CIII dimer and 13 from CIV), including 132 transmembrane helices (78 from CI, 26 from the CIII dimer and 28 from CIV; Extended Data Figs 3, 4, 5, 6, 7). The structures of the isolated complexes22,23,24 are sufficient to account for all observed protein density. Therefore, no clear density is seen around CIV for proposed subunits NDUFA4 or HIGD1a, predicted to contain transmembrane helices25,26,27,28.
The overall structure of the tight respirasome agrees (with some discrepancy at the position of CIV) with the low resolution (~20 Å) maps of the respirasomes in digitonin19, but not with those from amphipols18 (Extended Data Fig. 8). Whereas previously it was not possible to propose specific interactions between subunits, direct interactions, defined here as a distance of less than 1 nm between the poly-alanine models, are now observed (Fig. 2; in some cases we also note possible interactions at less than 2 nm).
a, b, A matrix view (left) and intermembrane space view (right) of the tight (a) and loose (b) respirasome models with complexes coloured as in Fig. 1. Nine interaction sites are indicated and described in the main text. Interacting subunits are darker. Subunits of CI that have been recently assigned are shown in red32. Arrows in b indicate the motions of CIII and CIV between the tight and loose respirasomes. c, View of the tight respirasome from the matrix side sliced to show the transmembrane helices, overlaid with the loose CIII and CIV models (white, aligned via CI). The interaction sites at the matrix and inner mitochondrial membrane interface are indicated. d, Side view of the tight respirasome with interaction sites between CI and CIII indicated and CIV removed for clarity, overlaid with the loose CIII aligned as in c.
Interactions between the complexes
Interactions between the supercomplexes can be organized into nine sites (Fig. 2). In all architectures, contacts between complexes I and III remain similar. At site 1, the B14.7/NDUFA11 (our study was on the ovine complex, but for reference we indicate the bovine/human nomenclature, respectively) subunit and the B15/NDUFB4 transmembrane helix of CI interact with UQCRQ of CIII at the matrix and inner mitochondrial membrane interface. B14.7 may play a role in stabilizing the CI lateral helix (HL)3,23,29,30,31 and interaction with CIII would further stabilize this region. At site 2, B22/NDUFB9 and the N terminus of B15 (ref. 32) of CI interact with UQCR1 and UQCRFS1 of CIII in the matrix. B22 and the N terminus of B15 protrude towards CIII and so may have been adapted specifically for supercomplex formation, along with two helices seen protruding from CI towards CIII in the intermembrane space. Possible interactions here are at site 3 between the C-terminal helix of PDSW/NDUFB10 and B14.7 of CI and UQCRH of CIII and at site 4 between B18/NDUFB7 of CI and COX7a of CIV. The long helix of B18 is poised at the interface of all three complexes and may contact both CIII and CIV. Both B18 and UQCRH contain disulfide bonds, suggesting that redox regulation may modulate the interactions between the complexes and providing another possible explanation for the interconversion between the tight and loose respirasomes. All interactions with CIII are mediated by CI supernumerary subunits, absent in bacteria. This is consistent with the lack of supercomplexes in bacteria, with the possible exception of P. denitrificans33, in which CI contains three supernumerary subunits34.
Contacts to CIV change between the tight and loose configurations. In the tight respirasomes, site 5 is mediated by the fifteenth transmembrane helix (TM15) of ND5 and B12/NDUFB3 (ref. 32) of CI, and subunits COX7c and COX8b of CIV at the matrix and inner mitochondrial membrane interface (Fig. 2a, d). On the basis of homology modelling of CI, this interaction is probably stabilized by a salt bridge between the conserved Glu503 of ND5 and Arg20 of COX7c. At site 6, the C terminus of CI subunit AGGG/NDUFB2 (ref. 32) runs along the helix of B18 and probably interacts with COX8b of CIV at the intermembrane space and inner mitochondrial membrane interface.
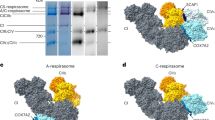
The interaction between CIII and CIV at site 7 is defined by a single CIV subunit, COX7a, which contacts subunit 10 (UQCR11) and UQCR1 of CIII at the matrix and inner mitochondrial membrane interface. After fitting the high-resolution structures of CIII and CIV into the maps, favourable salt-bridge interactions between highly conserved charged residues can be proposed (Fig. 3a).
a, b, Close-up view of site 7 looking from the matrix showing the fitted structures (with ovine sequence) of COX7a, UQCR1 and UQCR11 (a), same view with COX7a replaced by a homology model of SCAF1 (b). c, d, Close-up view of site 8 focusing on the interaction between COX7a and ND5. c, The fitted model of COX7a and a homology model of ND5 TM15. d, The same view with subunit COX7a replaced by a homology model of SCAF1. Possible stabilizing salt-bridge interactions are shown with ovals and interacting residues are labelled and numbered according to the mature subunit or in the case of SCAF1, the full-length protein. Complexes are coloured as in Fig. 1, COX7a is magenta and SCAF1 is orange.
In the loose respirasomes, CIV swings away from CIII (Figs. 1b and 2b) and COX7a contacts CI at site 8 through TM15 of ND5 (Fig. 3c), along with additional possible stabilizing contacts with B12 at the matrix and inner mitochondrial membrane interface (Fig. 2b, d). Otherwise CIV only contacts the rest of the supercomplex through possible interactions at site 9 between COX5b and UQCR1 of CIII (Fig. 2b). A rigid body rotation is observed between CI and CIII resulting in an increased distance between the matrix domains (Fig. 2). In CICIII2, the orientation of CIII is more similar to that in the tight respirasome, indicating that CIV, although only making few contacts, is influencing the architecture of the loose respirasome.
A role for supercomplexes in the stabilization of individual complexes, especially CI (ref. 8), is apparent from the multiple tight contacts between CI and CIII. The organization of the respiratory chain into even larger assemblies, or respiratory ‘strings’, has been suggested35 and the tight respirasome may provide a basis for such higher-order structures (Extended Data Fig. 9).
The absence of scaffold proteins
The transmembrane proteins SCAF1 and HIGD1a have been shown to promote supercomplex formation14,15,17,28, but are not seen in our maps. HIGD1a has only been detected co-migrating with CIV or CIII2CIV1 on blue native polyacrylamide gel electrophoresis (BN–PAGE)28,36 and hence is not expected to be found here. Conversely, SCAF1 co-migrates with most supercomplexes containing CIV on BN–PAGE, but not the individual complexes alone14,15,17. Therefore, the presence of SCAF1 is expected, but the question arises why no density for it is observed.
SCAF1 is a homologue of CIV subunit COX7a37. It is conceivable that SCAF1 is not an additional subunit in supercomplexes but replaces the endogenous COX7a subunit to promote supercomplex formation. Complementation of COX7a1 in the heart by the liver isoform COX7a2 has been observed in COX7a1 knockout mice38. When a homology model of SCAF1 based on the structure of COX7a1 (ref. 39) is placed within the tight respirasome, most amino acid changes have no effect on the interface between the subunit and the rest of CIV (Extended Data Fig. 10), and a similar set of favourable salt-bridging interactions can be seen between conserved basic residues of SCAF1 and the conserved acidic UQCR1 residues of CIII (Fig. 3a, b). The N terminus of mature SCAF1 extends by at least an additional 11 amino acids compared to mature COX7a14. These additional residues, predicted to be unstructured, may be disordered and are not seen in our maps, but would be well-positioned for further interaction with CIII. In the loose respirasome, similar interactions between CIV and CI are maintained between conserved residues with either COX7a1 or SCAF1 (Fig. 3c, d).
The resolution is currently not sufficient to determine the identity of the specific COX7a subunit present in the different respirasome architectures. However, these architectures provide a potential mechanism by which SCAF1 may promote the stability of supercomplexes.
Substrate channelling and ROS production
The structures presented here demonstrate that there is no protein-mediated substrate channel connecting the Q binding sites of CI and CIII, but instead both active sites are open to the membrane and separated by ~10 nm (Fig. 4a). The same is true for the flow of cytochrome c between CIII and CIV. The binding sites for cytochrome c are located ~10 nm apart and no barriers to free diffusion in the intermembrane space are seen (Fig. 4b). It cannot be excluded that full exchange with the Q pool becomes kinetically limiting at high respiration rates, which may explain some of the observations of substrate channelling10,13.
a. Side view of the tight respirasome with complexes coloured as in Fig. 1. The Q binding sites of CIII are shown by Q-2 molecules (red) based on crystal structures43. The entrance to the Q binding tunnel in CI is shown in red. Approximate membrane boundaries are indicated. b. Intermembrane space view of the tight respirasome coloured as in a, the loose position of CIV is also shown in yellow for comparison. Density for the detergent micelle is in grey. Black lines indicate the accessibility of the Q binding cavities in CIII with the thin dashed line indicating the change in accessibility in the loose architecture. Thick dashed lines indicate the shortest paths for cytochrome c diffusion from the two CIII binding sites to CIV in the tight respirasome. The arrows in a and b indicate the most direct path of QH2 from the CI tunnel to the proximal CIII cavity. Distances shown are in nm. IMS, intermembrane space.
The CIII dimer contains two Q binding cavities, each containing two Q binding sites (QP and QN, each provided by the opposite complex of the dimer). Reduced QH2 binds at the QP site and transfers one electron to cytochrome c in the intermembrane space and its second electron to a second Q bound at the QN site, generating the highly reactive intermediate ubisemiquinone (Q•). A second QH2 then binds at the QP site and reduces a second molecule of cytochrome c, in the process passing an electron to Q• at the QN site generating a fully reduced QH2. This is called the Q-cycle. Each CIII Q cavity is open to the membrane on opposite sides of the CIII dimer with a shared wall of protein preventing exchange of Q between them. In the respirasome, one of the Q cavities is adjacent to the CI Q tunnel and the other is adjacent to CIV (Fig. 4); we refer to them as the proximal and distal cavities, respectively, for their proximity to CI, the source of QH2. The proximal Q cavity is open to the membrane, whereas the distal Q cavity is partially occluded by CIV (Fig. 4b).
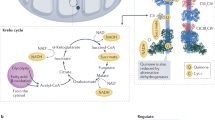
This arrangement may help limit reactive oxygen species (ROS) production by CIII (Fig. 5). The most rapid path for the electrons from QH2 oxidation in the proximal cavity would be to the UQCRFS1 proximal to CI and to the QN site of the same cytochrome b (MT-CYB) subunit, actually located in the distal Q cavity. Therefore, Q• would be generated in the distal cavity, partially capped by CIV. Other electron paths through CIII are possible through tunnelling between the bL haems of the dimer40. However, in the tight respirasome, the close approach between the distal UQCRFS1 and CIV (subunit COX6a2) may hamper the free rotation of this UQCRFS1 (necessary for electron transfer from the Qp site to CYC1 (ref. 41)) and thus prevent oxidation of Q in the distal cavity. This would break the inherent symmetry of CIII, meaning that it would have two distinct Q cavities, one cavity mainly for the oxidation of QH2, adjacent to the major source of QH2 (CI) and fully open to the membrane, and another cavity for the reduction of Q, capped by an oxidase (CIV). Because of the high activity of CIV, limited by the diffusion of O2, the local concentration of O2 would be low near the distal cavity, ideal for shielding a reactive intermediate (Q•). In the second part of the Q cycle, Q• would be reduced to QH2 and could be exchanged with the pool. For each Q reduced in the distal cavity, two QH2 would be oxidized in the proximal cavity, so the exchange rate in the distal cavity could be lower, consistent with its partial occlusion (Fig. 4b). It has been shown that the activities of the isolated complexes are higher than the activities of the complexes in the supercomplexes9,12,42. In addition to the possible role in the stabilization of individual complexes, architectures of supercomplexes may have been optimized during evolution, so that ROS production is limited9 by breaking the symmetry of CIII, at the expense of maximal activity.
Two electrons from NADH (top right) are passed through CI (blue), reducing Q to QH2. Diffusing from the Q-tunnel, QH2 may enter the proximal Q cavity of CIII (the oxidation cavity) or may diffuse into the membrane pool. QH2 passes one electron to cytochrome c in the intermembrane space and one to Q in the distal cavity (the reduction cavity), creating the Q• intermediate. Oxidation of a second QH2 in the proximal cavity will lead to reduction of Q• in the distal cavity to QH2, which can then leave and join the membrane pool. Reduced cytochrome c can diffuse from CIII to CIV, probably through the cytochrome c pool. The distal (from CI) cytochrome c reduction site in CIII is shown in a pale colour, as its operation may be suboptimal due to the restricted movement of nearby UQCRFS1.
Methods
Data reporting
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Sample preparation
Mitochondria were isolated from ovine heart tissue according to procedure 3 of ref. 44 and stored at −80 °C. Before supercomplex extraction with digitonin, frozen mitochondria were thawed on ice and washed by resuspension to a final concentration of ~6 mg protein per ml by manual homogenization in milliQ (18 MΩ) water to which KCl was added to a final concentration of 150 mM. Next, the membranes were pelleted by centrifugation at 32,000g for 45 min, followed by a second wash with resuspension in buffer M (20 mM Tris-HCl pH 7.4, 50 mM NaCl, 1 mM EDTA, 10% v/v glycerol, 2 mM DTT and 0.005% PMSF) by manual homogenization to a concentration of ~4 mg protein per ml and centrifugation at 32,000g for 45 min. Finally, membranes were resuspended in buffer M at ~10 mg protein per ml and either refrozen for storage at −80 °C or used directly for preparation of supercomplexes.
Supercomplexes were isolated from the washed mitochondrial membranes by digitonin extraction followed by sucrose gradient ultracentrifugation, as described previously18,45 with slight modifications. Briefly, an aliquot of washed mitochondrial membranes, containing ~5 mg of total protein, was resuspended in 2 ml buffer MX (150 mM potassium acetate, 30 mM HEPES pH 7.7, 10% glycerol and 0.002% PMSF) by manual homogenization. To the mitochondrial suspension, 1 ml of a 3% digitonin solution was added giving a final detergent concentration of 1% and a detergent to protein ratio of 6:1 (w:w). The sample was then agitated by rotation at 4 °C for 30 min before centrifugation at ~16,000g for 10 min. The supernatant was then concentrated to 0.5 ml and applied to linear sucrose gradients (10–45% sucrose in 15 mM HEPES pH 7.7, 20 mM KCl) prepared on a BioComp Gradient Station. The gradients were spun at 130,000g for 21 h, then fractionated. The fractions were run on BN–PAGE using linear gradient gels (4–20% polyacrylamide) in order to visualize the protein content (Extended Data Fig. 1). Fractions containing respirasomes were run over a PD-10 desalting column equilibrated in 100 mM NaCl, 20 mM HEPES pH 7.7, 0.1% digitonin and allowed to stand for ~1 h at room temperature until a white precipitate formed from the excess digitonin. The precipitate was pelleted by centrifugation and the supernatant was concentrated to an absorbance of ~2.8 absorbance units (AU) at 480 nm, which corresponds to about ~2.0 mg ml−1.
Electron microscopy
Aliquots of 2.7 μl of the isolated supercomplexes were applied to Quantifoil Cu R0.6/1, 300 mesh holey carbon film grids, which were glow-discharged in air for 120 s at 25 mA. Using FEI Vitrobot MKIII, the grids were blotted for 32 s at 4 °C at 100% humidity, plunged into liquid ethane and stored in liquid nitrogen. The grids were loaded onto a FEI Titan Krios transmission electron microscope (MRC LMB, Cambridge, UK) operated at 300 kV. Images were collected using EPU software on a Falcon-II detector at a calibrated magnification of ×81,395 (pixel size of 1.72 Å) and a dose rate of 17.0 electrons per Å2 per second (Extended Data Fig. 1b). Each image was exposed for a total of 4 s and dose-fractionated into 69 movie frames. A defocus range of 1.0–5.0 μm was used. Three datasets were collected, two datasets of freshly prepared supercomplexes comprising 1,178 micrographs and one dataset of overnight incubated supercomplexes of 430 micrographs for a total of 1,608 micrographs.
The dataset for supercomplex particles prepared in the same fashion but with an overnight incubation at 4 °C before grid preparations was collected in order to determine if the ratio of the tight-to-loose respirasomes changes with time. In the original datasets, the distribution of particles was 51% tight, 26% loose and 22% CICIII2 (of 29,659 total good particles after 3D classification). For the particles incubated overnight, the distribution was 26% tight, 49% loose and 25% CICIII2 (of 12,474 total good particles). Both supercomplex datasets were combined for the calculation of the final structures.
Image processing
All processing steps were performed using RELION v.1.4 (ref. 46) unless otherwise stated. A subset of ~2,000 particles were picked manually from 4-s averaged images, extracted using 2962 pixel box and subjected to reference-free 2D classification. Representative 2D classes were then used as reference images for automatic particle picking of all micrographs. The automatically picked particles were manually screened to remove false positives and pick any additional particles that were missed, resulting in an initial dataset of 67,650. MOTIONCORR47 was used for whole-image drift correction of movie frames 1–32 of each micrograph (the remaining frames were not used subsequently). Contrast transfer function (CTF) parameters of the corrected micrographs were estimated using Gctf and refined locally for each particle48. The particles were extracted using 2962 pixel box and sorted by reference-free 2D classification. We selected 54,484 particles from good 2D classes for the 3D classification (Extended Data Fig. 1c, d), which was run for 15 iterations, using an angular sampling of 1.8°, a regularization parameter T of 8 and a 30 Å low-pass filtered initial model from a previous low-resolution structure of the respirasome19, with a soft mask around the model. 3D classification was then continued as above for 15 iterations with an angular sampling of 0.9° and finally for 20 iterations with an angular sampling of 0.5°. This resulted in three good classes (tight, loose and supercomplex I–III) and a class of bad particles with no clearly resolved features (Extended Data Fig. 1d).
A subset of 42,133 particles including the three good classes was selected for the first 3D auto-refinement. This subset of particles were re-extracted from the motion corrected micrographs with a 4962 pixel box (to allow for high-resolution CTF correction49), and all further refinement was performed using this box size. After initial auto-refinement, a particle-based beam-induced motion correction and radiation-damage weighing (particle polishing) was performed (Extended Data Fig. 2e, f)50. Auto-refinement of all the polished particles together resulted in a reconstruction at 6.1 Å overall resolution with an estimated angular accuracy of 0.8°. The particles were then split into their respective 3D classes (tight, 18,397 particles; loose, 13,910 particles; and CICIII2, 9,844 particles; Extended Data Fig. 1d) and auto-refinement was run for each class independently, resulting in final reconstructions at 5.8, 6.7 and 7.8 Å resolution with angular accuracies of 0.6°, 0.7° and 1.0°, respectively. All resolutions are based on the gold-standard (two halves of data refined independently) Fourier shell correlation (FSC) = 0.143 criterion51 (Extended Data Fig. 2d). FSC curves were calculated using soft masks around the protein and high-resolution noise substitution was used to correct for convolution effects of the masks on the FSC curves52. Prior to visualization, all maps were corrected for the modulation transfer function of the detector.
Local resolution analysis by Resmap53 revealed a range of resolution for each supercomplex reconstruction with the highest resolution in the core of complexes I and III (Extended Data Fig. 2). The tight respirasome architecture shows resolutions ranging from 5.0 Å in the core of CI to ~8–9 Å resolution on the distal end of CIV (Extended Data Fig. 2a). The loose respirasome architecture shows resolutions ranging from 6.0 Å in the core of CI to ~10–12 Å resolution for the distal end of CIV (Extended Data Fig. 2b). Supercomplex CICIII2 shows 7.0 Å resolution at the core of CI ranging to ~15 Å at the peripheries of the CI and CIII matrix domains (Extended Data Fig. 2c). In Fig. 1 and Extended Data Figs 3, 4, 5, the maps have been carved in order to remove the detergent micelle and give a clear view of the transmembrane helices. Due to weaker density for CIV in Fig. 1a and Extended Data Fig. 3, it was contoured at a lower level and in Fig. 1b and Extended Data Fig. 4 it was filtered to 8 Å, whereas the maps for CI and CIII are filtered at 6.7 Å.
Model building
For the CIII and CIV models, available high-resolution structures of the bovine enzymes (PDB accession codes 1BGY (ref. 54) 1NTM (ref. 43) and 1V54 (ref. 39)) were used as starting models. Mutations to the ovine sequences were made in COOT55 manually, as only few changes were needed. Sequences for ovine COX7c and COX8b were not available in the online databases and hence the bovine sequences were used. The amino acid residues in these models were truncated at the beta-carbon using CHAINSAW56 in the CCP4 program suite57 to generate a ‘poly-alanine’ model and fit into the cryo-EM density map for the tight respirasomes as a rigid body.
For CI, the low-resolution poly-alanine model of the bovine enzyme was used as a starting model (PDB accession code 4UQ8 (ref. 23)). This model was fit into the tight respirasome map and manual building was performed in COOT55 using Ramachandran and secondary-structure restraints. Mammalian CI consists of 14 conserved core subunits present throughout the species, including bacteria, and 31 mitochondria-specific supernumerary subunits (29 unique supernumerary subunits and two copies of the acyl carrier protein SDAP). As noted before23, core subunits retain the fold and architecture originally characterized in bacteria29,58,59. Building was guided largely by secondary structure predictions and homology structure predictions generated using the programs PSIPRED60,61 and Phyre2 (ref. 62). We improved the completeness of the poly-alanine model for all currently assigned subunits (with the exception of the B8 and SDAP subunits whose structures are known from homologues63,64) and assigned three additional subunits (Extended Data Fig. 7).
B18 and PDSW are both found on the intermembrane space side of CI and both contain strong secondary structure predictions for helix-turn-helix motifs with B18 containing a double CX9C CHCH domain65. Two regions of density were previously identified as probable candidates for either of these subunits, but a definitive assignment between the two was not made66. Based on secondary structure prediction, which differentiates the subunits by their helical structure at the C terminus, we assigned B18 to the density near the interface of CI and IV that extends a long (~30 amino acid residues) helix into the interface of the three complexes (Fig. 2 and Extended Data Fig. 7). This means that the helical density found underneath ND4 and ND5 corresponds to PDSW, which is predicted to have an additional shorter C-terminal α-helix (15–20 amino acid residues) following its predicted helix-turn-helix motif. Density for an α-helix (which we suggest belongs to the PDSW subunit) can be seen near the end of the PDSW helix-turn-helix motif that protrudes from CI towards CIII in the supercomplex structures.
Additionally, we assigned and built subunit B17.2. B17.2 contains strong secondary structure prediction on its N terminus for two short helices followed by a 3–4 strand β-sheet, then by a long coil (~70 amino acid residues) with no predicted secondary structure. Starting from a model generated by homology structure prediction using Phyre2 (ref. 62), we were able to fit B17.2 into density adjacent to the 49-kDa, TYKY and PSST subunits of the CI Q-modules. After manual adjustment of the secondary structure elements from the predicted homology model, density could be seen extending away from the C terminus, which clearly belonged to the B17.2 C-terminal coil. These C-terminal residues snake along the surface of the peripheral arm and extend towards the N-module. This extended coil connecting the N- and Q-modules of the hydrophilic arm speaks to the role of B17.2 and its homologue B17.2L in CI assembly67.
In subsequent work with isolated ovine CI, we assigned all remaining CI subunits32 and, where relevant, these assignments (B15, B12 and AGGG) are used here.
A single round of real space refinement (morphing plus minimization) was performed using PHENIX real space refine68 for each complex individually in the tight respirasome map. Each of the refined complexes were fit into the lower resolution loose respirasome and CICIII2 maps as rigid bodies. Figures showing the fits of each complex into the density are shown in Extended Data Figs 3, 4, 5 for each map.
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
The EM maps have been deposited in the Electron Microscopy Data Bank under accession codes EMD-8130, EMD-8128 and EMD-8129. The models have been deposited in the Protein Data Bank (PDB) under accession codes 5J4Z, 5J7Y and 5J8K.
References
Sazanov, L. A. A giant molecular proton pump: structure and mechanism of respiratory complex I. Nature Rev. Mol. Cell Biol. 16, 375–388 (2015)
Chance, B., Estabrook, R. W. & Lee, C. P. Electron transport in the oxysome. Science 140, 379–380 (1963)
Hatefi, Y., Haavik, A. G. & Griffiths, D. E. Studies on the electron transfer system. XL. Preparation and properties of mitochondrial DPNH-coenzyme Q reductase. J. Biol. Chem. 237, 1676–1680 (1962)
Chazotte, B. & Hackenbrock, C. R. The multicollisional, obstructed, long-range diffusional nature of mitochondrial electron transport. J. Biol. Chem. 263, 14359–14367 (1988)
Hackenbrock, C. R., Chazotte, B. & Gupte, S. S. The random collision model and a critical assessment of diffusion and collision in mitochondrial electron transport. J. Bioenerg. Biomembr. 18, 331–368 (1986)
Schägger, H. & Pfeiffer, K. Supercomplexes in the respiratory chains of yeast and mammalian mitochondria. EMBO J. 19, 1777–1783 (2000)
Schägger, H. & Pfeiffer, K. The ratio of oxidative phosphorylation complexes I–V in bovine heart mitochondria and the composition of respiratory chain supercomplexes. J. Biol. Chem. 276, 37861–37867 (2001)
Acín-Pérez, R. et al. Respiratory complex III is required to maintain complex I in mammalian mitochondria. Mol. Cell 13, 805–815 (2004)
Maranzana, E., Barbero, G., Falasca, A. I., Lenaz, G. & Genova, M. L. Mitochondrial respiratory supercomplex association limits production of reactive oxygen species from complex I. Antioxid. Redox Signal. 19, 1469–1480 (2013)
Bianchi, C., Genova, M. L., Parenti Castelli, G. & Lenaz, G. The mitochondrial respiratory chain is partially organized in a supercomplex assembly: kinetic evidence using flux control analysis. J. Biol. Chem. 279, 36562–36569 (2004)
Blaza, J. N., Serreli, R., Jones, A. J. Y., Mohammed, K. & Hirst, J. Kinetic evidence against partitioning of the ubiquinone pool and the catalytic relevance of respiratory-chain supercomplexes. Proc. Natl Acad. Sci. USA 111, 15735–15740 (2014)
Enríquez, J. A. Supramolecular organization of respiratory complexes. Annu. Rev. Physiol. 78, 533–561 (2016)
Lenaz, G., Tioli, G., Falasca, A. I. & Genova, M. L. Complex I function in mitochondrial supercomplexes. BBA Bioenergetics 1857, 991–1000 (2016)
Lapuente-Brun, E. et al. Supercomplex assembly determines electron flux in the mitochondrial electron transport chain. Science 340, 1567–1570 (2013)
Ikeda, K., Shiba, S., Horie-Inoue, K., Shimokata, K. & Inoue, S. A stabilizing factor for mitochondrial respiratory supercomplex assembly regulates energy metabolism in muscle. Nature Commun. 4, 2147 (2013)
Mourier, A., Matic, S., Ruzzenente, B., Larsson, N.-G. & Milenkovic, D. The respiratory chain supercomplex organization is independent of COX7a2l isoforms. Cell Metab. 20, 1069–1075 (2014)
Williams, E. G. et al. Systems proteomics of liver mitochondria function. Science 352, aad0189 (2016)
Althoff, T., Mills, D. J., Popot, J. L. & Kühlbrandt, W. Arrangement of electron transport chain components in bovine mitochondrial supercomplex I1III2IV1 . EMBO J. 30, 4652–4664 (2011)
Dudkina, N. V., Kudryashev, M., Stahlberg, H. & Boekema, E. J. Interaction of complexes I, III, and IV within the bovine respirasome by single particle cryoelectron tomography. Proc. Natl Acad. Sci. USA 108, 15196–15200 (2011)
Mileykovskaya, E. et al. Arrangement of the respiratory chain complexes in Saccharomyces cerevisiae supercomplex III2IV2 revealed by single particle cryo-electron microscopy. J. Biol. Chem. 287, 23095–23103 (2012)
Mileykovskaya, E. & Dowhan, W. Cardiolipin-dependent formation of mitochondrial respiratory supercomplexes. Chem. Phys. Lipids 179, 42–48 (2014)
Xia, D. et al. Structural analysis of cytochrome bc1 complexes: implications to the mechanism of function. BBA Bioenergetics 1827, 1278–1294 (2013)
Vinothkumar, K. R., Zhu, J. & Hirst, J. Architecture of mammalian respiratory complex I. Nature 515, 80–84 (2014)
Tsukihara, T. et al. The whole structure of the 13-subunit oxidized cytochrome c oxidase at 2.8 Å. Science 272, 1136–1144 (1996)
Pitceathly, R. D. S. et al. NDUFA4 mutations underlie dysfunction of a cytochrome c oxidase subunit linked to human neurological disease. Cell Reports 3, 1795–1805 (2013)
Balsa, E. et al. NDUFA4 is a subunit of complex IV of the mammalian electron transport chain. Cell Metab. 16, 378–386 (2012)
Hayashi, T. et al. Higd1a is a positive regulator of cytochrome c oxidase. Proc. Natl Acad. Sci. USA 112, 1553–1558 (2015)
Chen, Y.-C. et al. Identification of a protein mediating respiratory supercomplex stability. Cell Metab. 15, 348–360 (2012)
Baradaran, R., Berrisford, J. M., Minhas, G. S. & Sazanov, L. A. Crystal structure of the entire respiratory complex I. Nature 494, 443–448 (2013)
Zickermann, V. et al. Structural biology. Mechanistic insight from the crystal structure of mitochondrial complex I. Science 347, 44–49 (2015)
Letts, J. A. & Sazanov, L. A. Gaining mass: the structure of respiratory complex I — from bacterial towards mitochondrial versions. Curr. Opin. Struct. Biol. 33, 135–145 (2015)
Fiedorczuk, K. et al. Structure of the entire mammalian mitochondrial complex I. Nature http://dx.doi.org/10.1038/nature19794 (2016)
Stroh, A. et al. Assembly of respiratory complexes I, III, and IV into NADH oxidase supercomplex stabilizes complex I in Paracoccus denitrificans. J. Biol. Chem. 279, 5000–5007 (2004)
Yip, C.-Y., Harbour, M. E., Jayawardena, K., Fearnley, I. M. & Sazanov, L. A. Evolution of respiratory complex I: “supernumerary” subunits are present in the alpha-proteobacterial enzyme. J. Biol. Chem. 286, 5023–5033 (2011)
Nübel, E., Wittig, I., Kerscher, S., Brandt, U. & Schägger, H. Two-dimensional native electrophoretic analysis of respiratory supercomplexes from Yarrowia lipolytica. Proteomics 9, 2408–2418 (2009)
Hayashi, H. et al. HIG1, a novel regulator of mitochondrial γ-secretase, maintains normal mitochondrial function. FASEB J. 26, 2306–2317 (2012)
Segade, F., Hurlé, B., Claudio, E., Ramos, S. & Lazo, P. S. Identification of an additional member of the cytochrome c oxidase subunit VIIa family of proteins. J. Biol. Chem. 271, 12343–12349 (1996)
Hüttemann, D. et al. Mice deleted for heart-type cytochrome c oxidase subunit 7a1 develop dilated cardiomyopathy. Mitochondrion 12, 294–301 (2012)
Tsukihara, T. et al. The low-spin heme of cytochrome c oxidase as the driving element of the proton-pumping process. Proc. Natl Acad. Sci. USA 100, 15304–15309 (2003)
Swierczek, M. et al. An electronic bus bar lies in the core of cytochrome bc1 . Science 329, 451–454 (2010)
Zhang, Z. et al. Electron transfer by domain movement in cytochrome bc1 . Nature 392, 677–684 (1998)
Shinzawa-Itoh, K. et al. Purification of active respiratory supercomplex from bovine heart mitochondria enables functional studies. J. Biol. Chem. 291, 4178–4184 (2016)
Gao, X. et al. Structural basis for the quinone reduction in the bc1 complex: a comparative analysis of crystal structures of mitochondrial cytochrome bc1 with bound substrate and inhibitors at the Qi site. Biochemistry 42, 9067–9080 (2003)
Smith, A. L. Preparation, properties, and conditions for assay of mitochondria: slaughterhouse material, small-scale. Methods Enzymol. 10, 81–86 (1967)
Dudkina, N. V., Eubel, H., Keegstra, W., Boekema, E. J. & Braun, H.-P. Structure of a mitochondrial supercomplex formed by respiratory-chain complexes I and III. Proc. Natl Acad. Sci. USA 102, 3225–3229 (2005)
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013)
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Scheres, S. H. W. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3, e03665 (2014)
Scheres, S. H. W. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nature Methods 9, 853–854 (2012)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2013)
Iwata, S. et al. Complete structure of the 11-subunit bovine mitochondrial cytochrome bc1 complex. Science 281, 64–71 (1998)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Stein, N. CHAINSAW: a program for mutating pdb files used as templates in molecular replacement. J. Appl. Cryst. 41, 641–643 (2008)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Sazanov, L. A. & Hinchliffe, P. Structure of the hydrophilic domain of respiratory complex I from Thermus thermophilus. Science 311, 1430–1436 (2006)
Efremov, R. G. & Sazanov, L. A. Structure of the membrane domain of respiratory complex I. Nature 476, 414–420 (2011)
Jones, D. T. Protein secondary structure prediction based on position-specific scoring matrices. J. Mol. Biol. 292, 195–202 (1999)
Buchan, D. W. A., Minneci, F., Nugent, T. C. O., Bryson, K. & Jones, D. T. Scalable web services for the PSIPRED protein analysis workbench. Nucleic Acids Res. 41, W349–W57 (2013)
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nature Protocols 10, 845–858 (2015)
Brockmann, C. et al. The oxidized subunit B8 from human complex I adopts a thioredoxin fold. Structure 12, 1645–1654 (2004)
Parris, K. D. et al. Crystal structures of substrate binding to Bacillus subtilis holo-(acyl carrier protein) synthase reveal a novel trimeric arrangement of molecules resulting in three active sites. Structure 8, 883–895 (2000)
Szklarczyk, R. et al. NDUFB7 and NDUFA8 are located at the intermembrane surface of complex I. FEBS Lett. 585, 737–743 (2011)
Zhu, J. et al. Structure of subcomplex Iβ of mammalian respiratory complex I leads to new supernumerary subunit assignments. Proc. Natl Acad. Sci. USA 112, 12087–12092 (2015)
Ogilvie, I., Kennaway, N. G. & Shoubridge, E. A. A molecular chaperone for mitochondrial complex I assembly is mutated in a progressive encephalopathy. J. Clin. Invest. 115, 2784–2792 (2005)
Afonine, P. V., Headd, J. J., Terwilliger, T. C. & Adams, P. D. New tool: phenix.real_space_refine. Computational Crystallography Newsletter 4, 43–44 (2013)
Van Kuilenburg, A. B., Van Beeumen, J. J., Van der Meer, N. M. & Muijsers, A. O. Subunits VIIa,b,c of human cytochrome c oxidase. Identification of both ‘heart-type’ and ‘liver-type’ isoforms of subunit VIIa in human heart. Eur. J. Biochem. 203, 193–199 (1992)
Acknowledgements
We thank the MRC LMB Cambridge for the use of the Titan Krios microscope. Data processing was performed using the IST high-performance computer cluster. J.A.L. holds a long-term fellowship from FEBS. K.F. is partially funded by a MRC UK PhD fellowship.
Author information
Authors and Affiliations
Contributions
J.A.L. purified supercomplexes, prepared cryo-EM grids, processed and analysed data, built models and co-wrote the manuscript; K.F. prepared cryo-EM grids, collected cryo-EM data and aided with model building; L.A.S. designed and supervised the project, analysed data and co-wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks A. Engel, D. Winge and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Supercomplex preparation and image processing procedures.
a, BN–PAGE of fractions from a typical sucrose gradient of digitonin extracted supercomplexes. Partially purified CI and molecular weight standards (thyroglobulin, 669 kDa; ferritin, 440 kDa; catalase, 240 kDa) were also included in the gel. Fraction eight (marked with an asterisk) was taken and used for subsequent supercomplex grid preparation. See the Supplementary Information for full uncropped scans of the gels. b, Representative micrograph of the 1,608 micrographs collected with slightly higher than average particle count; additionally, each micrograph differed slightly in ice thickness and defocus. Scale bar, 100 nm. c, Representative 2D class averages obtained from reference-free classification. d, Classification and refinement procedures used in this study.
Extended Data Figure 2 Local resolution and overall resolutions of supercomplex maps.
a–c, Local resolution estimation by Resmap53 of the tight respirasome (a), the loose respirasome (b) and CICIII2 (c). The supercomplexes are shown from the CI side (left) and the CIII side (right), with the full map (detergent micelle visible), and with the detergent micelle cut out by carving the maps around the protein model (middle images). d, Gold-standard FSC curves for the overall maps. e, f, Per-frame B-factors (e) and Guinier plot intercepts (Cf) (f) used for particle polishing in RELION46 calculated using all good particles.
Extended Data Figure 3 Tight respirasome models and maps for each complex.
Two views of each respirasome complex model fit into the cryo-EM density, with CI in blue (top), CIII in green (middle) and CIV in pink (bottom). The detergent micelle density has been removed for clarity.
Extended Data Figure 4 Loose respirasome models and maps for each complex.
Two views of each respirasome complex model fit into the cryo-EM density, with CI in blue (top), CIII in green (middle) and CIV in pink (bottom). The detergent micelle density has been removed for clarity. The CIV map has been filtered at 8 Å.
Extended Data Figure 5 Supercomplex CICIII2 models and maps for each complex.
Two views of each CICIII2 complex model fit into the cryo-EM density, with CI in blue (top) and CIII in green (bottom). The detergent micelle density has been removed for clarity.
Extended Data Figure 6 Uncut cryo-EM density for the tight respirasome at different contour levels.
Density within 5 Å of the poly-alanine models was coloured with CI in blue, CIII in green and CIV in pink, excess density is coloured grey. a, Low-contour level showing protein and the detergent micelle which is disordered and so has much weaker, fragmented density with no features. b, Medium contour showing mainly protein density. Note that grey density around complexes III and IV is completely featureless, indicating that the known structures account for all observed protein density. c, High contour showing FeS clusters and haem co-factors. Density for N1a cluster in CI, the FeS clusters of UQCRFS1 in CIII and the binuclear copper centre in CIV are not visible due to disorder. N1b cluster is occluded by N4 at this angle. In the loose respirasome, maps peaks for haem a and a3 are not visible in CIV due to disorder.
Extended Data Figure 7 CI model and density.
a–d, Four views of CI showing the density (top) and model (bottom) from within the membrane plane along the side of the membrane arm (a, b), within the membrane plane from the interface of the hydrophilic and hydrophobic arm (c) and from the intermembrane space side (d). a–c, Images are shown with the matrix up and the intermembrane space down. Subunits and density are coloured as indicated. Subunits within the specified colours are: (1) core subunits 75 kDa, 49 kDa, 30 kDa, PSST, TYKY, 51 kDa, 24 kDa, ND1, ND2, ND3, ND4, ND4L, ND5, ND6; (2) supernumerary subunits 39 kDa, B8, B13, 42 kDa, B16.6, B14.7, SDAP-α/β; (3) core subunits 75 kDa and 30 kDa, extended C termini. Supernumerary subunits rebuilt or extended B14, 18 kDa, 13 kDa, B16.6, PGIV, 15 kDa, B14.5b, 42 kDa, ESSS, B22; (4) B17.2, PDSW, B18; (5 and 6) KFYI, B15; (7 and 8) MWFE, B9, B14.5a, MNLL, AGGG, B12, SGDH, B17, ASHI. These subunits have been assigned in our subsequent work32; (9) the unmodelled density, which represents ~7% of the total volume of the CI density. Unmodelled density is primarily located on the solvent-exposed surfaces and represents mainly N- and C termini of known subunits. Similar unmodelled density is also observed in the 5 Å cryo-EM structure of isolated bovine CI (ref. 23).
Extended Data Figure 8 Comparison to previous low-resolution respirasome structures.
a, Models for complexes determined in this study were fit into the cryo-EM density of ref. 18 (a, left) and ref. 19 (a, right), and are consistent with the depictions in those publications, as well as for structures deposited in the PDB. In the structure of the respirasome prepared in amphipols (a, left), all of the complexes are much more separate without forming any of the tight contacts seen here (Fig. 2). This difference may be attributed to the disruptive effects of amphipols or other differences in the preparation or processing. These best fits are compared to our models for the tight (b) and loose (c) respirasomes (both left and right for direct comparison to the models above). For the previous structures in a, 3D-classification was not performed. Therefore, to determine whether the low-resolution maps correspond to an average of the different states observed here, we also fitted our models into a map generated from auto-refinement of all our particles. d, Tight + loose + CICIII2 combined map (Extended Data Fig. 1). This map is dominated by the tight conformation and the model remains distinct from the fits into the previously published low-resolution maps18,19.
Extended Data Figure 9 Modelling of putative higher-order organization of respiratory chain.
One possibility for respiratory strings model could be based on the dimerization of CIV observed in crystal structures24. CIV from the tight respirasome (cyan) was aligned with one of the monomers of CIV from PDB accession code 2X2Q. The second monomer of the crystallographic CIV dimer was then aligned with another copy of respirasome (magenta). a, View from the mitochondrial matrix. b, Side view in the membrane plane. In such an arrangement, there are no clashes between the two copies of the respirasome, with complexes III and IV forming a flat interaction area. The excessive curvature of the model (contrasting with the aligned in plane transmembrane domains in individual respirasomes) may be explained if the contacts between monomers in the crystallographic CIV dimer deviate somewhat from the contacts in the putative CIV dimer within the membrane. Such a symmetric ‘twin’ respirasome may form a building block for putative respiratory strings.
Extended Data Figure 10 COX7a homologues.
a, Sequence alignment of ovine COX7a1, COX7a2 and SCAF1 (also known as COX7a-2RP, COX7a2l). Amino acid changes relative to the COX7a1 sequence are indicated with an asterisk. The N termini of mature COX7a1 and COX7a2 have been determined experimentally69, however, the exact N terminus of mature SCAF1 is unknown. Mass-spectrometry-identified peptides from BN–PAGE indicate that the N terminus contains at least an additional 11 amino acids14, however, these residues are predicted to be unstructured and hence have not been included in the sequence alignment. The sequence numbering is based on mature COX7a1 and full-length protein for SCAF1. b, c, Amino acid differences between COX7a1 and COX7a2 (b) or SCAF1 (c) are mapped onto the structure of bovine COX7a1 indicating that most differences point away from CIV and may contribute to its interactions with other electron transport chain complexes.
Supplementary information
Supplementary Figures
This file contains the uncropped scans with size marker indications for Extended Data Figure 1a. (PDF 421 kb)
Rights and permissions
About this article
Cite this article
Letts, J., Fiedorczuk, K. & Sazanov, L. The architecture of respiratory supercomplexes. Nature 537, 644–648 (2016). https://doi.org/10.1038/nature19774
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19774
This article is cited by
-
SCAF1 drives the compositional diversity of mammalian respirasomes
Nature Structural & Molecular Biology (2024)
-
Di-(2-ethylhexyl) phthalate exposure induces premature testicular senescence by disrupting mitochondrial respiratory chain through STAT5B-mitoSTAT3 in Leydig cell
GeroScience (2024)
-
Structures of Tetrahymena thermophila respiratory megacomplexes on the tubular mitochondrial cristae
Nature Communications (2023)
-
Noncanonical role of singleminded-2s in mitochondrial respiratory chain formation in breast cancer
Experimental & Molecular Medicine (2023)
-
A FRET-based respirasome assembly screen identifies spleen tyrosine kinase as a target to improve muscle mitochondrial respiration and exercise performance in mice
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.