Abstract
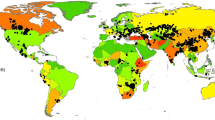
Anthrax is a globally important animal disease and zoonosis. Despite this, our current knowledge of anthrax ecology is largely limited to arid ecosystems, where outbreaks are most commonly reported1,2,3. Here we show that the dynamics of an anthrax-causing agent, Bacillus cereus biovar anthracis, in a tropical rainforest have severe consequences for local wildlife communities. Using data and samples collected over three decades, we show that rainforest anthrax is a persistent and widespread cause of death for a broad range of mammalian hosts. We predict that this pathogen will accelerate the decline and possibly result in the extirpation of local chimpanzee (Pan troglodytes verus) populations. We present the epidemiology of a cryptic pathogen and show that its presence has important implications for conservation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
European Nucleotide Archive
References
Hampson, K. et al. Predictability of anthrax infection in the Serengeti, Tanzania. J. Appl. Ecol. 48, 1333–1344 (2011)
Hugh-Jones, M. E. & de Vos, V. Anthrax and wildlife. Rev. Sci. Tech. 21, 359–383 (2002)
Lindeque, P. M. & Turnbull, P. C. Ecology and epidemiology of anthrax in the Etosha National Park, Namibia. Onderstepoort J. Vet. Res. 61, 71–83 (1994)
Beyer, W. & Turnbull, P. C. Anthrax in animals. Mol. Aspects Med. 30, 481–489 (2009)
Turnbull, P. C. B. WHO Guidelines Approved by the Guidelines Review Committee (World Health Organization, Department of Communicable Diseases Surveillance and Response, 2008)
Good, K. M., Houser, A., Arntzen, L. & Turnbull, P. C. Naturally acquired anthrax antibodies in a cheetah (Acinonyx jubatus) in Botswana. J. Wildl. Dis. 44, 721–723 (2008)
Wafula, M. M., Patrick, A. & Charles, T. Managing the 2004/05 anthrax outbreak in Queen Elizabeth and Lake Mburo National Parks, Uganda. Afr. J. Ecol. 46, 24–31 (2008)
Clegg, S. B., Turnbull, P. C., Foggin, C. M. & Lindeque, P. M. Massive outbreak of anthrax in wildlife in the Malilangwe Wildlife Reserve, Zimbabwe. Vet. Rec. 160, 113–118 (2007)
Muoria, P. K. et al. Anthrax outbreak among Grevy’s zebra (Equus grevyi) in Samburu, Kenya. Afr. J. Ecol. 45, 483–489 (2007)
Turnbull, P. C. et al. Anthrax in wildlife in the Luangwa Valley, Zambia. Vet. Rec. 128, 399–403 (1991)
de Vos, V. The ecology of anthrax in the Kruger National Park, South Africa. Salisbury Med. Bull. 68 (Suppl), 19–23 (1990)
Lembo, T. et al. Serologic surveillance of anthrax in the Serengeti ecosystem, Tanzania, 1996–2009. Emerg. Infect. Dis. 17, 387–394 (2011)
Beyer, W. et al. Distribution and molecular evolution of Bacillus anthracis genotypes in Namibia. PLoS Negl. Trop. Dis. 6, e1534 (2012)
Leendertz, F. H. et al. Anthrax kills wild chimpanzees in a tropical rainforest. Nature 430, 451–452 (2004)
Klee, S. R. et al. The genome of a Bacillus isolate causing anthrax in chimpanzees combines chromosomal properties of B. cereus with B. anthracis virulence plasmids. PLoS ONE 5, e10986 (2010)
Brézillon, C. et al. Capsules, toxins and AtxA as virulence factors of emerging Bacillus cereus biovar anthracis. PLoS Negl. Trop. Dis. 9, e0003455 (2015)
Antonation, K. S. et al. Bacillus cereus biovar anthracis causing anthrax in sub-Saharan Africa—chromosomal monophyly and broad geographic distribution. PLoS Negl. Trop. Dis. 10, e0004923 (2016)
Leendertz, F. H. et al. Anthrax in western and central African great apes. Am. J. Primatol. 68, 928–933 (2006)
Calvignac-Spencer, S. et al. Carrion fly-derived DNA as a tool for comprehensive and cost-effective assessment of mammalian biodiversity. Mol. Ecol. 22, 915–924 (2013)
Berry, H. H. Surveillance and control of anthrax and rabies in wild herbivores and carnivores in Namibia. Rev. Sci. Tech. 12, 137–146 (1993)
Vergnaud, G. et al. Comparison of French and worldwide Bacillus anthracis strains favors a recent, post-Columbian origin of the predominant North-American clade. PLoS ONE 11, e0146216 (2016)
Smith, K. L. et al. Bacillus anthracis diversity in Kruger National Park. J. Clin. Microbiol. 38, 3780–3784 (2000)
Campbell, G., Kuehl, H., Diarrassouba, A., N’Goran, P. K. & Boesch, C. Long-term research sites as refugia for threatened and over-harvested species. Biol. Lett. 7, 723–726 (2011)
Blackburn, J. K., Van Ert, M., Mullins, J. C., Hadfield, T. L. & Hugh-Jones, M. E. The necrophagous fly anthrax transmission pathway: empirical and genetic evidence from wildlife epizootics. Vector Borne Zoonotic Dis. 14, 576–583 (2014)
Hill, K. et al. Mortality rates among wild chimpanzees. J. Hum. Evol. 40, 437–450 (2001)
Köndgen, S. et al. Pandemic human viruses cause decline of endangered great apes. Curr. Biol. 18, 260–264 (2008)
Boesch, C & Boesch-Achermann, H. The Chimpanzees of the Taï Forest: Behavioural Ecology and Evolution (Oxford Univ. Press, 2000)
Turnbull, P. C. B. Guidelines for the Surveillance and Control of Anthrax in Humans and Animals 3rd edn (World Health Organization, Department of Communicable Diseases Surveillance and Response, 1998)
Ellerbrok, H. et al. Rapid and sensitive identification of pathogenic and apathogenic Bacillus anthracis by real-time PCR. FEMS Microbiol. Lett. 214, 51–59 (2002)
Klee, S. R. et al. Characterization of Bacillus anthracis-like bacteria isolated from wild great apes from Cote d’Ivoire and Cameroon. J. Bacteriol. 188, 5333–5344 (2006)
Panning, M. et al. Diagnostic reverse-transcription polymerase chain reaction kit for filoviruses based on the strain collections of all European biosafety level 4 laboratories. J. Infect. Dis. 196 (Suppl 2), S199–S204 (2007)
Leonard, J. A. et al. Animal DNA in PCR reagents plagues ancient DNA research. J. Archaeol. Sci. 34, 1361–1366 (2007)
Gamba, C. et al. Comparing the performance of three ancient DNA extraction methods for high-throughput sequencing. Mol. Ecol. Resour. 16, 459–469 (2016)
Rohland, N. & Hofreiter, M. Ancient DNA extraction from bones and teeth. Nat. Protoc. 2, 1756–1762 (2007)
Buffalo, V. Scythe: a 3′-end adapter contaminant trimmer. https://github.com/vsbuffalo/scythe (2014)
Joshi, N. A. & Fass, J. N. Sickle: a sliding-window, adaptive, quality-based trimming tool for FastQ files. https://github.com/najoshi/sickle (2011)
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013)
Auwera, G. A. et al. From FastQ data to high-confidence variant calls: the genome analysis toolkit best practices pipeline. Curr. Protoc. Bioinform. 43, 11.10.1–11.10. 33 (2013)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011)
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012)
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods 9, 772 (2012)
Posada, D. Using MODELTEST and PAUP* to select a model of nucleotide substitution. Curr. Protoc. Bioinform. 00, 6.5.1–6.5.14 (2003)
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010)
Rambaut, A., Lam, T. T., Max Carvalho, L. & Pybus, O. G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2, vew007 (2016)
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012)
Drummond, A. J., Ho, S. Y., Phillips, M. J. & Rambaut, A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 4, e88 (2006)
Kimura, M. Estimation of evolutionary distances between homologous nucleotide sequences. Proc. Natl Acad. Sci. USA 78, 454–458 (1981)
Baayen, R. H. Analyzing Linguistic Data: A Practical Introduction to Statistics using R (Cambridge Univ. Press, 2008)
McCullagh, P. & Nelder, J. A. Generalized Linear Models Vol. 37 (CRC, 1989)
Barr, D. J., Levy, R., Scheepers, C. & Tily, H. J. Random effects structure for confirmatory hypothesis testing: keep it maximal. J. Mem. Lang. 68, 225–278 (2013)
Schielzeth, H. & Forstmeier, W. Conclusions beyond support: overconfident estimates in mixed models. Behav. Ecol. 20, 416–420 (2009)
Forstmeier, W. & Schielzeth, H. Cryptic multiple hypotheses testing in linear models: overestimated effect sizes and the winner’s curse. Behav. Ecol. Sociobiol. 65, 47–55 (2011)
Dobson, A. J & Barnett, A. An Introduction to Generalized Linear Models (CRC, 2008)
R Core Team. R: A Language and Environment for Statistical Computing http://www.R-project.org/ (R Foundation for Statistical Computing, Vienna, Austria, 2013)
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015)
Ersts, P. Geographic Distance Matrix Generator (version 1.2.3). http://biodiversityinformatics.amnh.org/open_source/gdmg (American Museum of Natural History, 2011)
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004)
Goslee, S. C. & Urban, D. L. The ecodist package for dissimilarity-based analysis of ecological data. J. Stat. Softw. 22, 1–19 (2007)
Kuehl, H. S., Elzner, C., Moebius, Y., Boesch, C. & Walsh, P. D. The price of play: self-organized infant mortality cycles in chimpanzees. PLoS ONE 3, e2440 (2008)
Acknowledgements
We thank the authorities in Côte d’Ivoire for long-term support, especially the Ministry of the Environment and Forests, the Ministry of Research, the directorship of the TNP and the CSRS in Abidjan; and national authorities from all other countries for providing permissions for our research (MINFoF, MINRESI, the Service de la Conservation de la Réserve du Dja, Cameroon, in Central African Republic; the Ministère des Eaux et Fôrets, Chasse et Peche and the Ministère de l’Education Nationale, de l’Alphabetisation, de l’Enseignement Superieur, et de la Recherche, the Agence Nationale des Parcs Nationaux, Gabon; Centre National de la Recherche Scientifique et Technologique, Gabon; Direction des Eaux, Forêts et Chasses, Senegal; Forestry Development Authority, Liberia; Institut Congolais pour la Conservation de la Nature, Democratic Republic of the Congo; Ministère de l’Agriculture de l’Elevage et des Eaux et Forêts, Guinea; Instituto da Biodiversidade e das Áreas Protegidas (IBAP), Guinea-Bissau; Ministère de la Recherche Scientifique, Democratic Republic of the Congo; Ministère de le Recherche Scientifique et Technologique, Democratic Republic of the Congo; Nigeria National Park Service, Nigeria, Uganda National Council for Science and Technology, Ugandan Wildlife Authority, Uganda). We thank the WWF Central African Republic, T. Börding, T. Hicks, Y. Moebius, V. Sommer, K. Zuberbühler and M. Peeters for their logistical support; the field assistants A. Henlin, K. Albrechtova and A. Lang for the collection of samples in TNP; and the field assistants from all other sites for their support; S. Becker, T. Franz, S. Howaldt, A. Lander, P. Lochau, H. Nattermann and A. Schneider for the laboratory work; J. Hinzmann, A. Nitsche and J. Tesch for sequencing; P. Wojciech Dabrowski and T. Semmler from RKI, as well as G. Hamilton at Glasgow Polyomics, for bioinformatic support; and M. Kovacev-Wegener for administrative support. We thank the German Research Council DFG KL 2521/1-1 and the Sonnenfeld-Stiftung for funding; and the Max-Planck-Society and Krekeler Foundation for funding of the Pan African Programme.
Author information
Authors and Affiliations
Contributions
C.H., F.Z., A.A., S.A., M.A., G.B., K.C., T.D., P.D., K.D., H.E., P.F., Y.G.Y., A.G., A.-C.G., S.McG., J.H., S.J., J.J., J.K., K.La., J.L., K.Le., F.L., V.L., T.L., S.Ma., A.M., S.Me., M.M., J.v.S., E.T. and D.W. collected flies, bones and associated field data. Necropsies on wildlife that was found dead were performed by F.Z., K.N., A.B., E.C.-H., A.D., P.F., S.A.L., T.L., S.Me., S.N., H.D.N. and F.H.L. and laboratory analyses were performed by C.H., F.Z., K.N., S.D., R.G., K.M.-R., K.M., S.Me., H.D.N., A.S., U.T., S.R.K., L.H.W., S.C.-S. and F.H.L. The data were analysed by C.H., F.Z., R.B., H.K., R.M. and S.C.-S. and the manuscript was prepared by C.H., F.Z., R.B., H.K., R.M., J.F.G., S.C.-S. and F.H.L. The manuscript was revised and approved by all authors. The study was supervised by C.B., R.M.W., S.C.-S. and F.H.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Necropsies performed since 1996.
The total number of necropsies performed per year in TNP from 1996 to 2015. Grey bars indicate the number of Bcbva-positive necropsies and are annotated with the associated proportion. In the years 2003 and 2010 only limited veterinary service was available at TNP owing to political insecurity in the region.
Extended Data Figure 2 Geographic location of Bcbva-positive carcasses in TNP.
Necropsies that tested Bcbva-positive in TNP since 2001. GPS data was available for 70 of all necropsies that tested positive (n = 81).
Extended Data Figure 3 Effect of mammalian DNA content on anthrax positivity in flies.
Shown is the probability of Bcbva positivity (PA) as a function of the amount of mammalian DNA (copies) found in a fly. The amount of mammal DNA was binned (bin width of 0.25) and the area of the points depicts the number of flies (range, 1–206) in the respective bins. The dashed line indicates the fitted model and the dotted lines the 95% confidence interval.
Extended Data Figure 4 Effect of season on anthrax positivity in flies.
The probability of Bcbva positivity (PA) over the course of a year (binned in 10-day periods) is shown. The area of the points depicts the number of flies in the respective 10-day period. The dashed line indicates the fitted model and the dotted lines the 95% confidence interval.
Extended Data Figure 5 Maximum clade credibility tree based on chromosomal sequences of Bcbva isolates from TNP (Côte d’Ivoire, n = 124) and Grebo (Liberia, n = 2).
One sequence per host (mammals or flies, two divergent isolates for fly 600) was included and the final alignment of variant sites measured 298 bp. The size of the nodes represents posterior probability values. The location of the root received a posterior probability of 1.
Extended Data Figure 6 Maximum likelihood tree for sub-Saharan Bcbva strains.
Maximum likelihood tree based on chromosomal sequences of Bcbva strains from Côte d’Ivoire, Cameroon, Central African Republic and Liberia. The alignment of variant sites measured 1,016 bp. Bootstrap values are shown above the branches and the scale bar indicates substitution per chromosomal site. The tree was rooted using TempEst version 1.5.
Extended Data Figure 7 Fly snapshot sampling scheme.
For the fly snapshot, flies were caught following a 2 × 2-km grid system within and outside the research area within 19 days. In total 908 snapshot flies were analysed.
Extended Data Figure 8 Genetic and geographic distances of Bcbva isolates from the fly snapshot.
a, Maximum likelihood tree based on chromosomal sequences of Bcbva isolates from the 19-day fly snapshot. Each dot represents one fly isolate. Colours were chosen to illustrate the distribution of genetically clustering isolates on the map presented in b. The final alignment of variant sites measured 123 bp. Bootstrap values are shown above all internal branches. The tree was rooted using the ‘best-fit’ option in Path-O-Gen version 1.2. The scale bar shows substitutions per site. b, Geographic origin of Bcbva isolates collected during the fly snapshot. Colours correspond to maximum likelihood tree in a. Large circles represent two isolates.
Extended Data Figure 9 Box plot of genetic and mean geographic distances.
Bcbva isolates from TNP were binned by relative genetic distance (bin size = 0.03, approximately 2.5 SNPs).The two most genetically distant isolates received a value of 1 and all other distances were scaled accordingly. Diamonds indicate the geographic distance means of the groups. To examine variation within genetic lineages, we analysed isolates with low genetic distance (maximum relative genetic distance <0.5, marked in blue) and their mean geographic distance. For low genomic distances, the linear regression of geographic distances on genetic distances has an R2 of 0.72 and a slope coefficient that differs significantly from zero (Student’s t-test, P = 4 × 10−5).
Extended Data Figure 10 Fly species composition based on generalized mixed Yule-coalescent model (GMYC) analysis.
a–c, Fly species composition for three sites with known Bcbva occurrence: TNP, Côte d’Ivoire (a); Dja Faunal Reserve, Cameroon (b); Dzanga-Sangha Protected Areas, Central African Republic (c). The proportions of flies per site (%) belonging to a single fly species identified with GMYC models are shown. Different colours indicate different taxonomic fly families.
Supplementary information
Supplementary Information
This file contains a detailed method section as well as additional tables (Tables S1-10) and figures (Fig. S1-8). (PDF 2638 kb)
Supplementary Table 1
This file contains results that were derived from the analyses of flies caught in TNP analyzed in this study. The file includes results from PCR and culture as well as flymeal analysis results for a selection of flies. (XLSX 148 kb)
Supplementary Table 2
This file contains results of fly meal analysis with taxonomic assignment at genus level. The file provides the number of sequences per amplicon assigned at genus level. (XLSX 97 kb)
Supplementary Table 3
This file contains results of fly meal analysis with taxonomic assignment at order level. The file provides the number of sequences per amplicon assigned at order level. (XLSX 24 kb)
Rights and permissions
About this article
Cite this article
Hoffmann, C., Zimmermann, F., Biek, R. et al. Persistent anthrax as a major driver of wildlife mortality in a tropical rainforest. Nature 548, 82–86 (2017). https://doi.org/10.1038/nature23309
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23309
This article is cited by
-
Zootherapy as a potential pathway for zoonotic spillover: a mixed-methods study of the use of animal products in medicinal and cultural practices in Nigeria
One Health Outlook (2022)
-
The cost of living in larger primate groups includes higher fly densities
EcoHealth (2022)
-
The Movement of Pathogen Carrying Flies at the Human–Wildlife Interface
EcoHealth (2022)
-
Development and validation of portable, field-deployable Ebola virus point-of-encounter diagnostic assay for wildlife surveillance
One Health Outlook (2021)
-
Multiple stages of evolutionary change in anthrax toxin receptor expression in humans
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.