Abstract
Modifications on mRNA offer the potential of regulating mRNA fate post-transcriptionally. Recent studies suggested the widespread presence of N1-methyladenosine (m1A), which disrupts Watson–Crick base pairing, at internal sites of mRNAs1,2. These studies lacked the resolution of identifying individual modified bases, and did not identify specific sequence motifs undergoing the modification or an enzymatic machinery catalysing them, rendering it challenging to validate and functionally characterize putative sites. Here we develop an approach that allows the transcriptome-wide mapping of m1A at single-nucleotide resolution. Within the cytosol, m1A is present in a low number of mRNAs, typically at low stoichiometries, and almost invariably in tRNA T-loop-like structures, where it is introduced by the TRMT6/TRMT61A complex. We identify a single m1A site in the mitochondrial ND5 mRNA, catalysed by TRMT10C, with methylation levels that are highly tissue specific and tightly developmentally controlled. m1A leads to translational repression, probably through a mechanism involving ribosomal scanning or translation. Our findings suggest that m1A on mRNA, probably because of its disruptive impact on base pairing, leads to translational repression, and is generally avoided by cells, while revealing one case in mitochondria where tight spatiotemporal control over m1A levels was adopted as a potential means of post-transcriptional regulation.
Similar content being viewed by others
Main
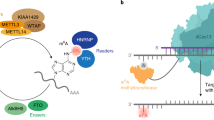
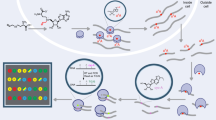
We developed a protocol for mapping m1A at single-nucleotide resolution (m1A-seq), relying on the property of m1A to lead to typical misincorporation and truncation profiles upon reverse transcription3. Key steps in the protocol include (1) antibody-based enrichment of m1A-comprising mRNA fragments1,2, (2, optional step) Dimroth rearrangement, by which m1A sites are converted to N6-methyladenosine (m6A)2,4, which eliminates the m1A misincorporation and truncation patterns, and (3) reverse transcription using either a highly processive reverse transcriptase (TGIRT) with a greater tendency to lead to misincorporations at m1A sites5,6, or a less processive one (SuperScript III (SS)) tending to lead to premature truncations (Fig. 1a and Supplementary Note 1).
a, Scheme depicting the m1A-seq pipeline. Following poly(A) selection, mRNA is fragmented into approximately 100-nucleotide-long fragments. m1A-containing fragments undergo enrichment using an anti-m1A antibody, following which they are subjected to reverse transcription using either TGIRT or SuperScript enzymes, leading predominantly to misincorporation and premature truncation, respectively. As controls, the immunoprecipitation is either omitted (input) or followed by a Dimroth rearrangement which converts m1A to m6A (IP + Dimroth). b, Venn diagram depicting the overlap between sites detected across the four datasets analysed here. The number of putative m1A sites within all classes of RNA are indicated in black; sites within tRNA molecules are indicated in parenthesis in red. AMV, avian myeloblastosis virus reverse transcriptase. c, Pie chart summarizing the classes of RNAs in which the 205 putative m1A containing sites were observed. Within tRNAs, the sites are further classified on the basis of the position within tRNA harbouring the sites. d, Table presenting all the putative m1A sites detected in this study, excluding those in tRNA molecules. For each site, we present the percentage of misincorporation in input and IP experiments (Mis. (input) (%) and Mis. (IP) (%)), which were calculated as the mean respective values across the datasets in which these sites were detected, the percentage truncation (Trunc. (%)) as estimated from m1A-seq-SS upon m1A-IP, the number of samples and experiments (Samp. and Exp.) in which the site was independently detected, and the sequence surrounding the putative m1A site. We highlight two sequence motifs that are reproducibly found within them in red and green. The putative m1A site is highlighted in yellow. e, Misincorporation, truncation, and coverage plots for putative m1A sites identified in this study. The graphical representation, inspired by ref. 3, depicts the truncation rate (black line, left y axis), misincorporation rate (stacked barplot, left y axis), and the overall coverage (grey shade, right y axis) in a sequence window surrounding the putative m1A site.
We applied m1A-seq-SS and m1A-seq-TGIRT to RNA derived from HEK293T cells. Each dataset included samples sequenced either directly (‘input’), or following m1A-immunoprecipitation (‘IP’), or following both m1A-IP and Dimroth rearrangement (‘IP + Dimroth’). We enhanced these two datasets with 8 input and IP sample pairs from ref. 2, and 14 sample pairs from ref. 1. These experimental datasets collectively comprised an unprecedented depth of more than 2 billion reads (Supplementary Table 1). We developed a single, common analytical pipeline to identify m1A-specific misincorporation profiles, relying on statistical tests assessing whether misincorporation rates were significantly higher in (1) IP samples, compared with input, or (2) IP compared with IP + Dimroth. In addition, we required putatively modified sites to have at least two distinct types of misincorporation (for example, A→T and A→G), and used the reverse transcriptase truncation rate as an optional filter (Extended Data Fig. 1a and Methods). Collectively, we identified 205 high-confidence putative m1A sites, and 72 additional lower-ranking ones (Fig. 1b–e and Supplementary Table 2). Quantifications of reverse transcriptase truncation and misincorporation levels were highly reproducible among replicates (Extended Data Fig. 1b–d). Each site was detected, on average, in approximately 11 independent samples (interquartile range 8–16), and 115 sites were independently identified in at least 2 datasets (Fig. 1b, c). As expected, use of TGIRT-based m1A-seq led to higher misincorporation and lower truncation rates at the detected sites compared with the SuperScript-based libraries (Extended Data Fig. 1e, f); IPs highly enriched for misincorporation rates, and Dimroth rearrangement reduced them (Extended Data Fig. 1g, h). We established our approach as highly sensitive and specific, as it allowed de novo discovery of almost all known classes of m1A sites in cytosolic tRNAs and rRNA, with very few false positives (Supplementary Note 2a). We establish misincorporation rates as a quantitative, relative estimate of m1A levels, providing a lower boundary on m1A stoichiometry (Extended Data Fig. 1i and Supplementary Note 2b).
Surprisingly, our dataset comprised predominantly well-established sites in rRNA and tRNA (Fig. 1c), and only 15 sites in mRNA and long non-coding RNAs (lncRNAs), 10 of which were in cytosolic transcripts and 5 in mitochondria (Fig. 1d). Most sites had very low misincorporation rates (less than 2.5%) in the absence of antibody-mediated enrichment (Fig. 1d), suggesting that they are modified at low stoichiometries. Two notable exceptions, both previously shown to harbour RNA:DNA sequencing differences of unknown nature, were in (1) tRNA-like mascRNA (MALAT1-associated small cytoplasmic RNA)7 and (2) mitochondrially encoded ND5 transcript8,9 (Supplementary Note 3). Misincorporation levels at the detected mRNA sites in the cytosol across thousands of samples from more than 50 tissues in the Genotype-Tissue Expression (GTEx) collection10 similarly revealed very low misincorporation levels, but high levels in rRNAs and in mascRNA serving as controls (Extended Data Fig. 2 and Supplementary Note 4), suggesting that the low m1A levels are not specific only to tumorous cell lines. We further exclude the possibility that the low number of sites in mRNAs is due to decreased detection power in mRNAs that are less expressed (Supplementary Note 5 and Extended Data Fig. 3a, b). In addition, we observe the previously described 5′-biased distribution of m1A-seq peaks1,2 (Extended Data Fig. 4a–d and Supplementary Table 3), but find no m1A-specific misincorporation profiles within these regions (Supplementary Note 5). It remains to be established whether these enrichments originate from complex modification patterns at the first transcribed nucleotide, or are experimental artefacts. Thus, with few exceptions, m1A is rarely observed at internal sites on mRNA, and typically at low stoichiometry.
All sites in cytosolic mRNAs and in Malat1 mascRNA comprised a single motif, or slight deviations thereof, consisting of the sequence GUUCNANNC (A = m1A) within a strong hairpin structure typically comprising a 5-base pair (bp) stem and a 7-bp loop (Fig. 2a; see also Supplementary Note 6 and Extended Data Fig. 5a, b). This sequence and structural motif is identical to the T-loop of tRNAs, where m1A at position 58 is catalysed by the TRMT6/TRMT61A complex at precisely the same relative position11,12. Consistently, knockdown of TRMT6/TRMT61A resulted in elimination of m1A from T-loop harbouring mRNAs (Extended Data Fig. 5c). Conversely, overexpression of TRMT6/TRMT61A followed by m1A-seq-TGIRT markedly increased misincorporation rates at the detected sites (Fig. 2b), and led to accumulation of m1A at 384 cytosolic mRNA and lncRNA sites (Fig. 2c and Supplementary Table 4), a massive enrichment with respect to the 10 originally identified sites. The GUUCNANNC motif was highly enriched at these sites (present in 193 out of 384 sites) (Fig. 2d), which were further enriched for a stable T-loop-like structure (Fig. 2e), typically consisting of a 7 bp loop (Extended Data Fig. 5d) and a 6–7 bp stem (Extended Data Fig. 5e). A subset of the peaks could be further validated by seeking m1A sites that had converted to m6A following Dimroth treatment of mRNA extracted from TRMT6/TRMT61A-overexpressing cells (Supplementary Note 7 and Extended Data Fig. 6a–e). To directly explore the determinants of specificity of TRMT6/TRMT61A, we used a plasmid library comprising thousands of T-loop sequences and systematically mutated counterparts13, all cloned as 3′ untranslated region (UTR) elements downstream of a reporter. Co-transfection of this plasmid pool with TRMT6/TRM61A into HEK293T cells allowed reconstitution of the m1A misincorporation signal precisely at the predicted position (Fig. 2f). Systematic point-mutation of each position in the GUUCNANNC motif allowed functionally reconstruction of the consensus required for modification via the TRMT6/TRMT61A complex (Fig. 2g), highlighting requirements for G–C base pairing at positions −5 and +3, and the requirements for a pyrimidine, a cytosine, and a purine at positions −3, −2 and −1, respectively. Systematic structural mutants and compensatory mutations demonstrated direct dependency of misincorporation on stem stability (Fig. 2h). These analyses demonstrate that within the cytosol, m1A is catalysed at T-loop-like elements via TRMT6/TRMT61A.
a, Predicted secondary structures of sequence environment surrounding putative m1A-containing sites in cytoplasmic mRNA and lncRNAs (see also Extended Data Fig. 5a); note the 7 bp loop stabilized by a relatively strong stem with terminal G–C base pair and UUCNANY loop. b, Frequencies of misincorporation rates obtained in input versus IP RNA fractions, in wild type (WT) versus upon overexpression of TRMT6/TRMT61A, shown for three putative m1A sites. Error bars reflect binomial confidence intervals, following read aggregation from two biological replicates. c, Pie chart as in Fig. 1c, depicting the distribution into RNA classes of 495 putative m1A sites detected upon overexpression of TRMT6/TRMT61A, of which 384 are within mRNA and lncRNA molecules. Mito., mitochondrial; Cyto., cytosolic. d, Sequence motif obtained by unbiased sequence analysis of the sequences surrounding the putative sites in mRNA upon overexpression. e, Distributions of predicted free energies among the 384 putative m1A sites in mRNA + lncRNAs, compared with randomly shuffled controls. These values are derived on the basis of parsing of the predicted secondary structures obtained using RNAfold. Note that lower free energies are indicative of more thermodynamically stable structures. P value is based on Mann–Whitney U-test. f, Misincorporation percentages across three regions that were monitored in the massively parallel reporter assay. Positions are numbered with respect to the predicted m1A site, and the ‘0’ position with predicted m1A is highlighted in yellow. g, Functionally reconstructed sequence motif, based on measurement of misincorporation rates at each of the nucleotides in each of the displayed positions into every other nucleotide across each of the 74 T-loops (Methods). h, Misincorporation rates following abolition and gradual restoration of the stem (thick line, median; box boundaries, 25% and 75% percentiles; whiskers, 1.5-fold interquartile range). The numbers of consecutive complementary bases in the stem region (beginning with the T-loop proximal bases) are indicated in the x axis.
We next focused on the site in the mitochondrially encoded ND5 gene, harbouring the highest levels of modification in our dataset, with misincorporation rates of approximately 25% in polyadenylated RNA (poly(A) mRNA) from HEK293T cells (Supplementary Note 8). The absence of misincorporations in reads from the transcribed antisense strand (Extended Data Fig. 7a) and in DNA (Extended Data Fig. 7b) ruled out DNA heteroplasmy as their source. Misincorporation levels in ND5 RNA across thousands of GTEx samples were highly tissue specific: essentially absent in muscle and in heart but approximately 30% (median) in ovary and pituitary gland (Fig. 3a). Targeted sequencing of the ND5 locus in human muscle and ovary samples confirmed these findings (Fig. 3b). The relatively high m1A levels observed in the human ovary samples prompted an exploration of ND5 methylation in development. Strikingly, single-cell RNA-seq (scRNA-seq) from 1,529 individual cells14 revealed misincorporation levels greater than 75% at the eight-cell stage (Fig. 3c), roughly equivalent to those observed in 16S rRNA, which was previously shown to be methylated at nearly 100% stoichiometry15. Levels of misincorporation decreased with developmental progression, and by day 7 reached approximately 12.5% (Fig. 3c). scRNA-seq analysis of 124 single cells, spanning a wider developmental range16, extended these findings, and revealed that from the metaphase II oocyte to the four-cell embryo, methylation levels at ND5 are probably close to 100% (Fig. 3d and Supplementary Note 9) followed by a precipitous decrease (Fig. 3d). As zygotic mitochondrial transcription begins around the eight-cell developmental stage17, we speculated that m1A might mark particularly stable maternal transcripts that persist up to zygotic mitochondrial transcription. Indeed, transcriptional arrest in HEK293T cells using actinomycin D or ethidium bromide led to an approximately fourfold increase in ND5 misincorporation levels (Fig. 3e and Extended Data Fig. 7c, respectively), confirming the association between ND5 stability and methylation. Thus, m1A in ND5 is highly tissue- and developmentally specific, and may serve as a mark of a subset of ND5 transcripts, which are maternally inherited and dominate until zygotic mitochondrial transcription at roughly the eight-cell stage.
a, Highly tissue-specific distribution of misincorporation rates at ND5:1374 across 29 tissues on the basis of more than 9,000 RNA-seq datasets obtained from the GTEx collection; particularly high levels are observed in the pituitary gland and ovary. In comparison, misincorporation levels, pooled across all tissues, are presented for chromosome M (chrM):2617, a position in the mitochondrial 16S rRNA; This position was previously demonstrated to be methylated close to 100% (ref. 15), and thus allows a rough calibration of the readout. Boxplots represent median and interquartile range, as indicated in Fig. 2h. b, Misincorporation levels at ND5:1374 in human muscle and ovary samples, on the basis of targeted, strand-specific sequencing of the ND5 locus in poly(A) RNA. Error bars, binomial confidence interval from a single sample. c, Distributions of misincorporation levels across 1,529 individual cells from 88 human preimplantation embryos ranging from developmental day 3 (corresponding to the eight-cell stage) to day 7 (ref. 14); in this study the authors used SuperScript II for reverse transcriptase. 16S rRNA methylation levels are shown in comparison as in a. d, Distributions of misincorporation levels across 124 cells spanning a developmental range from metaphase II oocytes to late blastocyst16; in this study the authors used SuperScript III for reverse transcriptase. e, Misincorporation levels at ND5:1374 measured at the indicated time-points following actinomycin-D-mediated transcriptional arrest (n = 3). f, Misincorporation levels at ND5:1374 in HEK293T cells after overexpression (OE) of TRMT10C, treatment with short interfering RNAs (siRNAs) directed against TRMT10C (siTRMT10C), or treatment with a control siRNA. Error bars, binomial confidence interval based on a single, representative replicate. g, Misincorporation levels at ND5:1374 across 102 samples from human ovaries, colour-coded on the basis of the presence of a SNP at position 13708 (red, wild type; blue, SNP).
We hypothesized that m1A in ND5 is catalysed by TRMT10C, which catalyses methylation at position 9 of mitochondrial tRNAs18. Indeed, TRMT10C knockdown led to almost complete abolition of methylation at ND5:1374; TRMT10C overexpression resulted in a 50% increase in methylation levels (Fig. 3f). Analysis of the GTEx data revealed that ND5 methylation levels are under genetic control, as they are strongly correlated across different tissues from the same individual (Extended Data Fig. 7d, e). Detailed analysis revealed a relatively common single nucleotide polymorphism (SNP) (G13708A) two bases upstream of the ND5 site (at 13710), severely reducing the ability of ND5 to undergo methylation (Fig. 3g). Targeted sequencing of the ND5 locus in lymphoblastoid cell lines from individuals harbouring this SNP compared with controls confirmed these results (Extended Data Fig. 7f). G13708A is among the defining SNPs of the Eurasian J haplogroup, and is thought to affect the clinical expression of Leber hereditary optic neuropathy in western Eurasians19,20,21,22. Our results indicate that this haplotype has lost the ability to undergo efficient ND5 methylation.
We speculated that the Watson–Crick disruptive nature of m1A would prevent effective translation of modified codons. Polysome fractionation experiments revealed a substantial and highly significant reduction in misincorporation levels in ND5 in the heavier fractions relative to lighter ones (Fig. 4a and Extended Data Fig. 8a), suggesting repressed translation of m1A-harbouring transcripts. Consistently, upon overexpression of TRMT6/TRMT61A, we found a pronounced and highly significant depletion of m1A-modified mRNA in the ribosome-heavy fractions relative to the ribosome-poor fraction (Fig. 4b). This was observed for cytosolic sites that were present either in the 5′ UTR or within the coding DNA sequence (CDS), but not for a site present in the 3′ UTR (Fig. 4b). We next cloned a 60 bp region harbouring the m1A sites in the PRUNE gene in-frame and upstream of a firefly luciferase coding region (Fig. 4c). Co-transfection of this construct with TRMT6/TRM61A led to high m1A levels (Fig. 4d), which were eliminated upon disrupting the sequence or structure, but restored via a compensatory structural mutation (Fig. 4c, d). The point-mutation of the m1A site in PRUNE or structural disruption led to an approximately twofold increased luciferase levels compared with the sequences harbouring intact consensus sequences and T-loop structures (Fig. 4e). Conversely, no decrease—and even a slight increase (P = 0.03)—was observed when introducing the wild-type element into the 3′ UTR upon overexpression of TRMT6/TRMT61A compared with controls (Extended Data Fig. 8b–d). The consistent translational repression associated with m1A sites within the 5′ UTR or CDS, but not in the 3′ UTR, suggests that it may be dependent on ribosomal scanning or translation (Supplementary Note 10 and Extended Data Fig. 9).
a, Misincorporation rates at the ND5 locus across the polysomal fractions, measured using strand-specific targeted sequencing (n = 3). Dots, measurements; red bar, mean. b, Misincorporation rates at four selected loci following overexpression of TRMT6/TRMT61A, measured across four polysomal fractions (n = 3), displayed as in a. c, Depiction of the four designed variants, perturbing either the modified site or the secondary structure, on the basis of the methylated site in the PRUNE gene. The methylated position is plotted in red, the perturbed position in magenta. d, Misincorporation percentages at the designed site, quantified across the four constructs using targeted sequencing (n = 3). Dots, measurements; red bar, mean. Note that for the ‘site mutation’ variant the misincorporation rate reflects the fraction of reads not harbouring a ‘T’, in contrast to all remaining variants in which it reflects the fraction not harbouring an ‘A’. e, Renilla-normalized firefly luciferase levels in TRMT6/TRMT61A-overexpressing cells divided by the corresponding ratio in non-overexpressing control for the four tested constructs. Dots, measurements; red bars, mean. Presented are t-test-based P values.
Collectively, the ability to map and quantify m1A at single-nucleotide resolution allowed redefinition of its genome-wide distribution, diverging substantially from previous reports1,2, and, further, its biogenesis, functions, and potential mechanisms of action to be addressed. Similarly to pseudouridine13,23,24,25 and 5-methylcytosine26, m1A is also catalysed via co-opting of the tRNA/rRNA modifying machineries. The repressive impact of m1A on translation probably underlies its scarcity in cytosolic mRNAs. The marked reduction in ND5 methylation in mitochondria following the eight-cell stage coincides with activation of zygotic transcription at this stage, an increase in oxygen consumption27,28,29, and changes in mitochondrial morphology30, collectively suggestive of a developmental, regulatory role for m1A methylation at this position.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Cell culture for knockdown and overexpression experiments
Human HEK293T cells (American Type Culture Collection (ATCC); passage numbers 5–15; no further verification of cell line identity was performed, but the cells screened negatively for mycoplasma) were plated in six-well plates at 20% confluency. siRNAs targeting TRMT10C (Thermo Fisher s29784), TRMT6 (s28400), and TRMT61A (s41859) were transfected using Lipofectamine RNAiMAX (Life Technologies AB4427037) following the manufacturer’s protocols, with two siRNA boosts with a 48 h interval between them; siRNA targeting TRMT6/TRMT61A was co-transfected with half of the recommended amount of siRNA targeting TRMT6 and the other half targeting TRMT61A. As negative controls, we used Ambion In Vivo Negative Control #2 siRNA (catalogue number 4390846). Cells were harvested at 96 h. For overexpression, plasmids encoding full-length TRMT6/TRMT61A under a cytomegalovirus promoter were obtained from OriGene. The plasmids were transfected into HEK293T cells using PolyJet (SignaGene) with one boost of the plasmid at 24 h. Cells were harvested 48 h after transfection.
Blocking transcription or translation
Transcription was blocked using actinomycin D (Sigma) at a concentration of 10 μg ml−1 or ethidium bromide (Amresco) at a concentration of 0.4 μg ml−1. Translation was blocked using cycloheximide (Sigma) at concentration of 100 μg ml−1.
Human RNA
Total RNA extracted from ovary and muscles from a human donor was obtained from Takara. Lymphoblastoid cell lines were obtained from Coriell.
RNA preparation for m1A-seq
RNA was extracted from cells using NucleoZOL (Macherey-Nagel). Enrichment of poly(A)+ RNA from total RNA was performed using Oligo(dT) Dynabeads (Invitrogen) according to the manufacturer’s protocol. The mRNA was chemically fragmented into approximately 100-nt-long sections using RNA fragmentation reagent (Ambion). The sample was cleaned using Dynabeads (Life Technologies), and resuspended in 20 μl IPP buffer (150 mM NaCl, 0.1% NP-40, 10 mM Tris-HCl, pH 7.5).
m1A-seq and m6A-seq
The protocol we developed for transcriptome-wide mapping m1A is based on our previously published protocol for mapping N6-methyladenosine (m6A)31. Briefly, 40 μl of protein-G magnetic beads were washed and resuspended in 200 μl of IPP buffer, and tumbled with 5 μl of affinity-purified anti-m1A polyclonal antibody (MBL) at room temperature for 30 min. RNA was added to the antibody-bead mixture, and incubated for 2 h at 4 °C. The RNA was then washed twice in 200 μl of IPP buffer, twice in low-salt IPP buffer (50 mM NaCl, 0.1% NP-40, 10 mM Tris-HCl, pH 7.5), and twice in high-salt IPP buffer (500 mM NaCl, 0.1% NP-40, 10 mM Tris-HCl, pH 7.5), and eluted in 30 μl RLT (Qiagen). To purify the RNA, 20 μl of MyOne Silane Dynabeads (Life Technologies) were washed in 100 μl RLT, resuspended in 30 μl RLT, and added to the eluted RNA. Sixty microlitres of 100% ethanol were added to the mixture, the mixture attached to the magnet, and the supernatant discarded. After two washes in 100 μl of 70% ethanol, the RNA was eluted from the beads in 10 μl of H2O. Dimroth rearrangements were performed as described in ref. 2. For mapping m6A following Dimroth conversion of m1A, RNA extracted from TRMT6/TRMT61A-overexpressing cells was first Dimroth rearranged as described in ref. 2, and subjected to m6A-seq as previously published31.
Library preparation
Strand-specific m1A RNA-seq libraries were generated on the basis of the protocol described in refs 32, 33. Briefly, RNA was first subjected to FastAP Thermosensitive Alkaline Phosphatase (Thermo Scientific), followed by a 3′ ligation of an RNA adaptor using T4 ligase (New England Biolabs). Ligated RNA was reverse transcribed either using SuperScript III (Invitrogen) or TGIRT-III (InGex), and the cDNA was subjected to a 3′ ligation with a second adaptor using T4 ligase. The single-stranded cDNA product was then amplified for 9–12 cycles in a PCR reaction. Libraries were sequenced on Illumina Nextseq platforms generating short paired-end reads, ranging from 25 to 55 bp from each end.
Identification of putative m1A sites
A human reference genome was generated on the basis of the hg19 assembly of the human genome, supplemented with tRNA, rRNA, and snRNAs, obtained from the modomics database34. Non-enriched (input), m1A-enriched (IP), and Dimroth-treated m1A-enriched (IP + Dimroth) samples were aligned to the genome, using STAR aligner35 with an increased stringency allowing only up to three mismatches per each read pair (‘–outFilterMismatchNmax 3′). All duplicates were marked using picard tools MarkDuplicates.jar’, and non-primary alignments were removed. The identity of each nucleotide at each genomic position was extracted using ‘samtools mpileup’, with max per-file depth settings of ‘-d 100000’. A custom script was used to parse the pileup format into a tabular format summarizing the abundance of each nucleotide at each position. All positions harbouring an ‘A’ in the annotated sense strand, with at least two mismatches, occurring in at least 10% of the reads overlapping it were recorded.
All sites, recorded across any replicates and any of the samples, were pooled into a single data set. Misincorporation rates at each of these pooled sites were subsequently re-extracted for each of the experiments (allowing analysis of each site in each experiment, even if they did not pass the initial thresholds in that particular experiment). A bona fide m1A site is expected to have higher mismatch rates in IP compared with input, and lower levels in IP + Dimroth compared with IP. Hence, we used a χ2 test on the basis of the aggregated misincorporation counts (across replicates) to test the hypotheses that (1) the number of mismatches in the IP sample was higher than input, and (2) the number of mismatches in IP was higher than in IP + Dimroth. The following criteria were then used for identification of putative m1A sites: (1) at least one of the two calculated P values was significant (P < 0.05); (2) the product of the P values was less than 0.01; (3) a difference in misincorporation rate between the sample with the lowest levels of misincorporation and the highest level of at least 0.2; (4) the sites had to be covered by at least ten reads in at least two samples; (5) at least 1% of all reads mapping to the site (across all replicates and samples) had to be ‘A’, at least 1% of all reads mapping to the site (across all replicates and samples) had to be ‘T’, and at least 1% of all reads mapping to the site (across all replicates and samples) had to be ‘C’ or ‘G’. These last criteria were set to aid in the discrimination of SNPs (where only one alternative to an ‘A’ is expected) from m1A sites (where typically more than one type of misincorporation is observed). Sites harbouring identical sequences in a 24 bp window (12 bp upstream + 11 bp downstream) surrounding the putative site were filtered, to retain only one. Nonetheless, owing to merging of sites from different data sets and the multiple loci from which identical or nearly identical tRNAs were transcribed, a subset of duplication was retained—and are flagged as such—within tRNA entries in Supplementary Table 2.
This pipeline was applied to four batches of samples: (1) input, IP, and IP + Dimroth (two replicates each) to which we applied m1A-seq-SS; (2) input, IP, and IP + Dimroth (two replicates each) to which we applied m1A-seq-TGIRT; (3) 16 samples generated in ref. 2 with RNA-seq readouts from HepG2 cells in m6A-IP or input, with and without Dimroth rearrangements downloaded from Gene Expression Omnibus (GEO; accession number GSE70485); (4) 28 samples generated in ref. 1, with m1A mapped under different genetic perturbations and upon different stimuli downloaded from GEO (accession number GSE73941). The ‘high-confidence’ data set of 205 sites comprised all sites for which at least two significant P values were obtained across any of the comparisons performed across any of the data sets; the ‘low confidence’ sites comprised all sites associated with a single significant P value. Of note, to accommodate the distinct experimental design in the data sets obtained from refs 1, 2, we adapted the precise sets of comparisons that were performed by the analytical pipeline. Specifically, in addition to assessing whether IP differed from input, and from IP + Dimroth, in the ref. 2 data set we further derived χ2-based P values to assess whether the Dimroth rearrangement in the input samples led to reduced mismatch levels compared with in its absence. In the data set produced in ref. 1, the authors did not use a Dimroth rearrangement, but instead relied on treating the RNA with AlkB, an Escherichia coli-derived demethylase that eliminates m1A. Reference 1 further compared measurements upon knockout of AlkBH3, which they found to demethylate m1A. Accordingly, we performed four statistical tests, examining differences in mutation rates between (1) IP versus input in wild-type samples, (2) IP versus input across stress conditions (H2O2, starvation), (2) IP versus IP + AlkB, and (3) IP in wild-type cells versus IP in ALKBH3 knockout cells. Of note, given that any site passing any of the statistical tests was considered a putative site, effectively our criteria for identifying putative m1A sites in the data sets of refs 1, 2 are more lenient than the criteria we applied for the two data sets we generated. Finally, in the data set generated upon overexpression of TRMT6/TRMT61A, we did not perform Dimroth rearrangements; instead we performed two tests, examining differences in misincorporation rates between (1) input samples and IP samples, and (2) input samples following overexpression of TRMT6/TRMT61A compared with controls. All analyses were performed using the identical computational pipeline, into which we fed, as parameters, the precise comparisons to be made.
Although our m1A-seq approach provides strand-specific data, in the initial analyses in Figs 1 and 2 strand specificity was inferred from the genomic annotation rather than the read. This allowed application of an identical pipeline to the data generated in this study and the two previously published data sets. To analyse m1A at the ND5 locus, we subsequently called mutations in a strand-specific manner (separately inferring mutations on the + and − strands), to prevent dilution of misincorporation signal from the antisense strand.
Quantification of reverse transcriptase truncations
For each of the putatively identified m1A sites, we calculated the rate of transcription termination at position +1 with respect to the site. This was performed by first artificially merging all read pairs into a single, artificial read extending from the beginning of one read to the end of its mate, and then using bedtools to count the number of reads beginning and overlapping each position. The ratio between the two was defined as the stop rate at that position, as performed in ref. 23.
Identification of m1A peaks
Peak detection was performed on the basis of our previously published approach for detecting peaks in m6A-seq data31,36. Specifically, an in-house script was first used to project all reads aligning to the genome upon the human transcriptome. Only reads fully matching a transcript structure, as defined by the ‘UCSC Known Genes’ transcriptome annotation, were retained. Such reads were computationally extended in transcriptome space from the beginning of the first read to the end of its mate, and coverage in transcriptome space was calculated for each nucleotide across all transcripts.
Putative m1A sites were identified using a three-step approach, as follows. (1) Peak detection within genes. To search for enriched peaks in the m6A IP samples, we scanned each gene using sliding windows of 100 nt with 50-nt overlaps. Each window was assigned a peak over median (POM) score, defined as mean coverage in the window/median coverage across the gene. Windows with POM scores greater than 4 (that is, greater than fourfold enrichment) and with a mean coverage of more than ten reads were retained. Overlapping windows were merged together, and for each disjoint set of windows in transcriptome space we recorded its start, end, and peak position, corresponding to the position with the maximal coverage across the window. (2) Ensuring that peaks were absent in input. We repeated the procedure in step 1 for the input sample. We eliminated from all subsequent analysis all windows that were detected in both steps 1 and 2. (3) Comparison of multiple samples. To search for consistently occurring peaks across different samples, we first merged the coordinates of all windows from all samples passing steps 1 and 2, to define a set of disjoint windows passing these filters in at least one of the samples. For each such window, we recalculated the peak start, end, peak position, and POM score (as defined above) across each of the samples using the approach in step 1. In addition, for each window we calculated a peak over input (POI) score, corresponding to the fold change of coverage across the window in the IP sample over the coverage in the input sample. To account for differences in sample depth, we estimated the mean difference between IP and input samples across the 500 most highly expressed genes, which we used as an estimate for background. We subtracted this background from the POI score.
On the basis of careful examination of the peaks at the beginning of transcripts, which revealed that in many cases the peaks originated from the first transcribed nucleotide, we used the approach we described in ref. 36. Briefly, this approach relies on calculating the fold change upon m1A-IP, compared with input, in reads beginning at each of the first 50 annotated positions in each transcript. For the analysis displayed in Extended Data Fig. 4d, we first calculated these ratios across the set of SuperScript IP and input samples. We then integrated the quantifications of fold changes by extracting the maximum fold change per position per transcript. We then quantified the proportion of pileups harbouring an ‘A’ as a function of this fold change, revealing that pileups beginning with an ‘A’ were more frequent at the higher-confidence sites, harbouring stronger fold changes.
mRNA expression analysis
To estimate expression levels, reads were aligned against the human genome using RSEM (version 1.2.31) with default parameters37. For robust comparison between different samples, we used trimmed mean of M values (TMM) normalization of the RSEM read counts38 as implemented by the edgeR package39 in R.
Prediction of RNA secondary structure
For predicting secondary structure in the region surrounding putative m1A sites, we extracted a sequence window of 24 bp, including 12 bp upstream of the modified site and 11 bp downstream. Free-energy calculations and predicted secondary structures were calculated using RNAfold version 2.1.5. The secondary structures were subsequently parsed, using an in-house script, to quantify the stem and loop lengths (Fig. 2).
Massively parallel reporter assay
The design and cloning of the massively parallel reporter assay library into a plasmid were described in ref. 13. A 10 cm plate of HEK293T cells was transiently transfected with equal amounts of TRMT6, TRMT61A, and the library plasmid using PolyJet (SignaGene). RNA was purified using NucleoZOL reagent (Macherey-Nagel). Sequence-specific m1A-seq-TGIRT was performed on total RNA essentially as described in ref. 13, except reverse transcription from the constant sequence stemming from the library plasmid (AGCATTAACCCTCACTAAAGGGAAAGG) was done using TGIRT-III (InGex). Adaptor ligation and PCR enrichment with an inner-plasmid-specific primer (GGTCCGATATCGAATGGCGC) were performed as previously described13.
Alignment of the massively parallel reporter assay data was performed as previously described13. To quantify misincorporations we used ‘samtools mpileup’, as described above. For the sequence logo depicted in Fig. 2j, we first extracted the 75th percentile of misincorporation rates following point-mutation of each of the indicated sites across each of the assayed 74 T-loops into each of the 4 nucleotides. For each position, this value was then divided by the sum of this value across all four nucleotides, to yield ‘relative misincorporation rate’ (summing up to 1, at each position). The height of each nucleotide at each position was then plotted in direct proportion to its relative misincorporation rate.
Annotation of mitochondrial and tRNA sites
All reads were aligned to the chrM assembly forming part of the human hg19 assembly, and the Supplementary Tables provide positions with respect to it. For consistency with the mitochondrial community, within the manuscript we refer to positions with respect to the slightly more updated chrM_rCRS assembly. For tRNAs, we refer to all positions in the figures on the basis of the standard tRNA nomenclature (so that the anticodon nucleotides are always numbered 34–36, and the T-loop between positions 54 and 60).
Quantification of misincorporation in GTEx and single-cell RNA-seq data
Raw fastq files were obtained for each of the files in these data sets, and aligned using STAR (as above). Mpileup was applied to quantify misincorporation levels across the positions detected in this study. For Fig. 4c, d, we filtered out all single cells in which an SNP was observed at position 13708, as this SNP severely reduces methylation. An SNP was called in this position, on the basis of the RNA-seq data, if more than 80% of the reads corresponded to the SNP nucleotide.
Polysome fractionation
Polysome fractionation was done as specified in ref. 40, with one exception: we used 10–50% sucrose gradient instead of 5–50%.
Targeted sequencing of m1A amplicons
For targeted measurement of m1A levels at specific loci, reverse transcription was done on 1 μg of RNA using random hexamers and TGIRT-III (InGex) reverse transcriptase. Amplicons were PCR amplified using a nested PCR approach, involving a first amplification step with gene-specific primers and a partial Illumina adaptor tail, and a second amplification leading to the incorporation of the full-length Illumina adapters. For ND5 amplicons, reverse transcription was done using a strand-specific primer instead of random hexamers, to avoid contamination by the ND5 antisense transcript. The resultant amplicon was amplified with primers including the full-length Illumina adapters in a single step. All primers can be found in Supplementary Table 5.
Luciferase assay
For the luciferase experiments, two plasmids were used: (1) pGL4.73, for expression of Renilla luciferase under an SV40 promoter, and (2) a plasmid encoding an ATG start codon followed by 60 bp surrounding the PRUNE m1A site (GCGGAGGCCGATTCGCCGTGTGGCGGGTTCGAGTCCCGCCTCCTGACTCTGGCCTCTAGTC) followed by firefly luciferase, all driven by a cytomegalovirus promoter. We constructed three derivatives based on this plasmid, point mutating the sequence and structure, as described in the text. These plasmids were pooled together and transfected into cells with either control DNA or with TRMT6/61A overexpression plasmids. The luciferase assay was done with a Promega kit according to the manufacturer’s instructions.
Code availability
Code for the analyses described in this paper is available from the corresponding author upon request.
Data availability
All m1A-seq data sets generated in this manuscript have been deposited in the GEO under accession number GSE97419. All other data are available from the corresponding author upon reasonable request.
References
Li, X. et al. Transcriptome-wide mapping reveals reversible and dynamic N1-methyladenosine methylome. Nat. Chem. Biol. 12, 311–316 (2016)
Dominissini, D. et al. The dynamic N1-methyladenosine methylome in eukaryotic messenger RNA. Nature 530, 441–446 (2016)
Hauenschild, R. et al. The reverse transcription signature of N-1-methyladenosine in RNA-seq is sequence dependent. Nucleic Acids Res. 43, 9950–9964 (2015)
Jones, J. W. & Robins, R. K. Purine nucleosides. III. Methylation studies of certain naturally occurring purine nucleosides. J. Am. Chem. Soc. 85, 193–201 (1963)
Mohr, S. et al. Thermostable group II intron reverse transcriptase fusion proteins and their use in cDNA synthesis and next-generation RNA sequencing. RNA 19, 958–970 (2013)
Zubradt, M. et al. DMS-MaPseq for genome-wide or targeted RNA structure probing in vivo. Nat. Methods 14, 75–82 (2017)
Wilusz, J. E., Freier, S. M. & Spector, D. L. 3′ end processing of a long nuclear-retained noncoding RNA yields a tRNA-like cytoplasmic RNA. Cell 135, 919–932 (2008)
Hodgkinson, A. et al. High-resolution genomic analysis of human mitochondrial RNA sequence variation. Science 344, 413–415 (2014)
Bar-Yaacov, D. et al. RNA-DNA differences in human mitochondria restore ancestral form of 16S ribosomal RNA. Genome Res. 23, 1789–1796 (2013)
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015)
Ozanick, S., Krecic, A., Andersland, J. & Anderson, J. T. The bipartite structure of the tRNA m1A58 methyltransferase from S. cerevisiae is conserved in humans. RNA 11, 1281–1290 (2005)
Jühling, F. et al. tRNAdb 2009: compilation of tRNA sequences and tRNA genes. Nucleic Acids Res. 37, D159–D162 (2009)
Safra, M., Nir, R., Farouq, D. & Schwartz, S. TRUB1 is the predominant pseudouridine synthase acting on mammalian mRNA via a predictable and conserved code. Genome Res. http://dx.doi.org/10.1101/gr.207613.116 (2017)
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016)
Bar-Yaacov, D. et al. Mitochondrial 16S rRNA is methylated by tRNA methyltransferase TRMT61B in all vertebrates. PLoS Biol. 14, e1002557 (2016)
Yan, L. et al. Single-cell RNA-seq profiling of human preimplantation embryos and embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1131–1139 (2013)
Pikó, L. & Taylor, K. D. Amounts of mitochondrial DNA and abundance of some mitochondrial gene transcripts in early mouse embryos. Dev. Biol. 123, 364–374 (1987)
Vilardo, E. et al. A subcomplex of human mitochondrial RNase P is a bifunctional methyltransferase—extensive moonlighting in mitochondrial tRNA biogenesis. Nucleic Acids Res. 40, 11583–11593 (2012)
Hudson, G. et al. Clinical expression of Leber hereditary optic neuropathy is affected by the mitochondrial DNA-haplogroup background. Am. J. Hum. Genet. 81, 228–233 (2007)
Torroni, A. et al. Haplotype and phylogenetic analyses suggest that one European-specific mtDNA background plays a role in the expression of Leber hereditary optic neuropathy by increasing the penetrance of the primary mutations 11778 and 14484. Am. J. Hum. Genet. 60, 1107–1121 (1997)
Carelli, V. et al. Haplogroup effects and recombination of mitochondrial DNA: novel clues from the analysis of Leber hereditary optic neuropathy pedigrees. Am. J. Hum. Genet. 78, 564–574 (2006)
Brown, M. D., Sun, F. & Wallace, D. C. Clustering of Caucasian Leber hereditary optic neuropathy patients containing the 11778 or 14484 mutations on an mtDNA lineage. Am. J. Hum. Genet. 60, 381–387 (1997)
Schwartz, S. et al. Transcriptome-wide mapping reveals widespread dynamic-regulated pseudouridylation of ncRNA and mRNA. Cell 159, 148–162 (2014)
Carlile, T. M. et al. Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells. Nature 515, 143–146 (2014)
Lovejoy, A. F., Riordan, D. P. & Brown, P. O. Transcriptome-wide mapping of pseudouridines: pseudouridine synthases modify specific mRNAs in S. cerevisiae. PLoS ONE 9, e110799 (2014)
Edelheit, S., Schwartz, S., Mumbach, M. R., Wurtzel, O. & Sorek, R. Transcriptome-wide mapping of 5-methylcytidine RNA modifications in bacteria, archaea, and yeast reveals m5C within archaeal mRNAs. PLoS Genet. 9, e1003602 (2013)
Mills, R. M. & Brinster, R. L. Oxygen consumption of preimplantation mouse embryos. Exp. Cell Res. 47, 337–344 (1967)
Ginsberg, L. & Hillman, N. ATP metabolism in cleavage-staged mouse embryos. J. Embryol. Exp. Morphol. 30, 267–282 (1973)
Biggers, J. D. & Borland, R. M. Physiological aspects of growth and development of the preimplantation mammalian embryo. Annu. Rev. Physiol. 38, 95–119 (1976)
Wassarman, P. M. & Josefowicz, W. J. Oocyte development in the mouse: an ultrastructural comparison of oocytes isolated at various stages of growth and meiotic competence. J. Morphol. 156, 209–235 (1978)
Schwartz, S. et al. High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell 155, 1409–1421 (2013)
Shishkin, A. A. et al. Simultaneous generation of many RNA-seq libraries in a single reaction. Nat. Methods 12, 323–325 (2015)
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013)
Machnicka, M. A. et al. MODOMICS: a database of RNA modification pathways--2013 update. Nucleic Acids Res. 41, D262–D267 (2013)
Dobin, A . et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Schwartz, S. et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5′ sites. Cell Reports 8, 284–296 (2014)
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011)
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010)
Robinson, M. D., McCarthy, D. J. & Smyth, G. K . edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Gandin, V. et al. Polysome fractionation and analysis of mammalian translatomes on a genome-wide scale. J. Vis. Exp. 87, http://dx.doi.org/10.3791/51455 (2014)
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012)
Acknowledgements
This project was supported by the Israel Science Foundation (543165), the European Research Council under the European Union’s Horizon 2020 research and innovation programme (grant agreement number 714023), by the Abisch-Frenkel-Stiftung, by research grants from The Abramson Family Center for Young Scientists, the David and Fela Shapell Family Foundation INCPM Fund for Preclinical Studies, the Estate of David Turner, and the Berlin Family Foundation New Scientist Fund.
Author information
Authors and Affiliations
Contributions
M.S., A.S.-C., R.N., D.B., M.E., W.R., N.S.-G., and S.S. designed the experiments. M.S. performed the m1A-seq optimizations and experiments, the luciferase assays, and the genetic perturbations. M.S. and A.S.-C. performed the targeted sequencing experiments and the mitochondrial RNA half-life experiments. M.S. and R.W. performed the polysomal fractionations. R.N. performed the oligoarray experiment. A.N. assisted in the GTEx analysis. S.S. and M.S. performed the computational analysis. S.S. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks Y. Motorin and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Analytical pipeline for identification of m1A sites.
a, Overview of the analytical pipeline, which was applied to each of the four data sets. The pipeline utilizes two key statistical tests to identify ‘A’ containing sites that either harbour misincorporations at higher levels following immunoprecipitation with an anti-m1A antibody compared with input samples or compared with Dimroth-treated samples. In addition, the pipeline requires that minimum levels of at least two distinct types of misincorporation be observed for a given site. b–d, Truncation and misincorporation rates are reproducible between replicates, both with SuperScript and with TGIRT enzymes. Shown are scatterplots based on 205 putative m1A sites, indicating the correlation between replicates of truncation rates using SuperScript (b) misincorporation rates using SuperScript (c), and misincorporation rates using TGIRT (d). e, Comparison of truncation rates using TGIRT (y axis) and SuperScript (x axis), on the basis of the 205 detected sites. Sites are colour-coded on the basis of the indicated classes of RNA. f, Comparison of misincorporation rates using TGIRT and SuperScript, plotted as in e. g, Comparison of misincorporation rates in IP samples compared with IP + Dimroth, in both cases using TGIRT. h, Comparison of misincorporation rates in IP samples compared with input, using TGIRT. i, Serial dilutions of RNA extracted from cells overexpressing TRMT6/TRMT61A within RNA extracted from cells depleted of TRMT6/TRMT61A. The x axis denotes the percentage of the RNA originating from cells overexpressing TRMT6/TRMT61A, and the y axis captures the misincorporation rate, as measured via targeted sequencing of the three presented sites.
Extended Data Figure 2 Misincorporation levels at putative sites on the basis of GTEx data set.
Shown are misincorporation levels across eight lncRNA + mRNA and two rRNA sites on the basis of a random sampling of 4,114 GTEx RNA-seq samples. Boxplots represent median, interquartile range, distribution, and outliers, as indicated in Fig. 2h.
Extended Data Figure 3 m1A detection as a function of gene expression, on the basis of analysis of sites identified following TRMT6/TRMT61A overexpression.
a, All genes were divided into five bins, on the basis of the indicated levels of expression. The number of m1A harbouring sites in each bin is plotted. b, As in a, but normalized by number of genes within each bin. Error bars, binomial confidence intervals.
Extended Data Figure 4 Peak detection following m1A-seq reveals a strong enrichment towards the 5′ terminus.
a, Representative meta-gene profiles of read coverage in m1A-IP (top) or input (bottom). Shown are the first 500 nt immediately following the transcription start site (left) and immediately preceding the annotated 3′ terminus of the gene (right). Genes are divided into five bins, on the basis of expression levels. b, Distribution of expression levels (on the basis of input samples) of all genes harbouring high-confidence peaks. On this distribution are overlaid, in red, the genes in which putative m1A sites were detected. c, Consistently identified peaks are highly enriched towards the 5′ terminus of the gene. Each peak was classified into one of five segments, as in ref. 41, in the following order: transcription start site (TSS) if the peak is present in the first 200 nt of the gene; 5′ UTR if in the 5′ UTR region but outside the transcription start site region; stop codon region, comprising 200 nt on both sides of the stop window; CDS region for peaks within the CDS; and 3′ UTR for remaining 3′ UTR peaks. Each peak is scored on the basis of the number of experiments in which it was detected, whereby more robustly identified peaks should be considered as ones of higher confidence. The stacked bar plots summarize the relative proportion of peaks in each segment, and the right-most bar plots the relative amount of ‘gene architecture’ taken up by each of these segments. d, Analysis of transcription start site peaks. The number of reads beginning (rather than overlapping) at each of the first 50 annotated transcribed bases was calculated across IP and input samples (Methods), and the log(fold change) between the two was derived. All fold changes were binned into six bins (as plotted) and the fraction of positions harbouring an ‘A’ are plotted as a function of this binned fold change, revealing that positions at the transcription start site that are enriched in IP samples over input samples are biased towards beginning with an ‘A’. Error bars, binomial error.
Extended Data Figure 5 Characterization of sequence and structure at m1A sites, and requirement of the TRMT6/TRMT61A machinery for its catalysis in the cytosol.
a, Predicted secondary structures of sequence environment surrounding putative m1A-containing sites in cytoplasmic mRNA and lncRNAs (see also Fig. 2a). b, Predicted secondary structure at the vicinity of mitochondrial m1A sites. A common secondary structure surrounding mitochondrial sites is depicted, featuring a ‘UAAA’ motif in the loop, stabilized by a stem structure. c, Misincorporation levels at the indicated sites, measured by targeted sequencing, in cells depleted of TRMT6/TRMT61A via siRNAs, compared with mock-treated controls. Error bars, binomial confidence intervals. d, e, Distributions of loop length (d) and stem length (e) among the 384 putative m1A sites in mRNA + lncRNAs, compared with randomly shuffled controls. These values are derived from RNAfold.
Extended Data Figure 6 Validation of putative m1A sites, via m6A-seq of Dimroth-converted RNA extracted from TRMT6/TRMT61A mRNA.
a, Coverage plot along the cytosolic 28S rRNA across the four indicated samples. The known m1A and m6A sites are indicated. b, Coverage plots as in a for the mitochondrial 16S rRNA. The normalized coverage levels are indicated to the left of the track (note the orders-of-magnitude higher coverage at the m1A sites upon m6A-IP in Dimroth-treated samples, compared with the controls). c, Quantification of percentage of coverage in a 100 nt centred region around putative m1A sites out of overall coverage of the gene in two biological replicates of mRNA extracted from cells overexpressing TRMT6/TRMT61A; this mRNA was subjected to Dimroth treatment followed by m6A-seq. Quantifications were obtained for 40 sites exhibiting misincorporation levels greater than 10% upon TRMT6/TRMT61A overexpression in addition to the 2 sites on the 16S and 28S rRNA molecules. d, Comparison of ‘% Coverage in peak’ (as in c) between RNA subjected to Dimroth treatment and m6A-IP compared with RNA only subjected to m6A-IP. e, Misincorporation levels across the indicated conditions across 8 sites (of the 42 tested) in which a significant P value (P < 0.05) was obtained when comparing Dimroth + m6A-IP samples with their corresponding input (significance indicated by ‘*’), and/or when comparing Dimroth + m6A-IP to no-Dimroth + m6A-IP (significance indicated by ‘#’). Error bars, binomial error after pooling of two biological replicates.
Extended Data Figure 7 ND5 methylation levels are genetically determined, are in part controlled by a SNP two bases upstream, and are increased in stable transcripts.
a, Snapshot of randomly sampled reads aligned to the ND5 locus. Reads originating from the heavy strand are depicted in red, reads from the light strand in purple. Top: IP sample; bottom: input sample. Misincorporations are apparent only in reads originating from the heavy strand. b, Misincorporation levels at the ND5 locus in DNA and RNA samples of five individuals. For RNA, the distribution of misincorporation reads are shown across all tissues; for DNA, the measurement consists of a single measurement available in GTEx. c, Misincorporation levels at ND5:1374 measured at the indicated time-points following ethidium-bromide-mediated transcriptional arrest (n = 3); Points, mean; error bars, s.e.m. d, Correlations between misincorporation levels at ND5:1374 in skin versus brain samples, from the same individuals. e, Histogram of all pairwise correlation coefficients between tissues (but from the same individuals); note that values are centred around 0.5, rather than around 0 if they were independent of each other. f, Misincorporation rates at ND5:1374 obtained via strand-specific targeted sequencing of the ND5 locus across six lymphoblastoid cell lines, two harbouring a G13708A SNP and four wild-type samples.
Extended Data Figure 8 m1A represses translation.
a, Representative sucrose gradient, indicating the division into fractions on the basis of the number of polysomes associated with them. b, Scheme of experimental design. The wild-type m1A-containing stretch from the PRUNE gene was cloned either in-frame and upstream of firefly luciferase (CDS construct) or as a 3′ UTR element (3′ UTR construct). Control or TRMT6/TRMT61A-overexpressing cells were co-transfected with each of these plasmids along with a plasmid expressing Renilla luciferase. c, d, Renilla-normalized firefly luciferase levels in TRMT6/TRMT61A-overexpressing cells, standardized by this value in the non-overexpressing (control) cells (n = 3). Note that for the CDS construct the presented data are identical to those in Fig. 4 and are re-plotted for convenience.
Extended Data Figure 9 Cytosolic m1A does not affect steady-state levels of mRNA or mRNA stability.
a, Comparison of expression levels of all genes acquiring robust levels of m1A (defined as misincorporation levels greater than 10%) upon overexpression of TRMT6/TRMT61A, compared with wild-type (non-overexpressing) counterparts. b, Misincorporation levels in the four indicated genes, in cells overexpressing TRMT6/TRMT61A, in a 6 h time course following cycloheximide treatment (n = 3). Error bars, s.e.m.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-10 and Supplementary References.
Supplementary Table 1
Alignment statistics for all four analysed datasets, including total number of reads and the percentage of aligned reads.
Supplementary Table 2
Dataset of 277 putative m1A sites, obtained after intersection of the four analysed datasets – see Supplementary Information document for full description.
Supplementary Table 3
Dataset of 690 consistently identified peaks, used in the context of analyses presented in Extended Data Fig. 4 revealing strong bias towards 5’ termini of genes. The dataset is in bed format.
Supplementary Table 4
Dataset of 495 putative m1A sites, obtained following overexpression of TRMT6/TRMT61A.
Supplementary Table 5
A list of primer sequences used in this study.
Rights and permissions
About this article
Cite this article
Safra, M., Sas-Chen, A., Nir, R. et al. The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution. Nature 551, 251–255 (2017). https://doi.org/10.1038/nature24456
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24456
This article is cited by
-
Research progress of N1-methyladenosine RNA modification in cancer
Cell Communication and Signaling (2024)
-
Diabetes-induced male infertility: potential mechanisms and treatment options
Molecular Medicine (2024)
-
RNA modifications in cellular metabolism: implications for metabolism-targeted therapy and immunotherapy
Signal Transduction and Targeted Therapy (2024)
-
N6-methyladenosine modification is not a general trait of viral RNA genomes
Nature Communications (2024)
-
Nucleic acid and protein methylation modification in renal diseases
Acta Pharmacologica Sinica (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.