Abstract
Desmoplastic melanoma is a rare subtype of melanoma characterized by dense fibrous stroma, resistance to chemotherapy and a lack of actionable driver mutations, and is highly associated with ultraviolet light-induced DNA damage1. We analysed sixty patients with advanced desmoplastic melanoma who had been treated with antibodies to block programmed cell death 1 (PD-1) or PD-1 ligand (PD-L1). Objective tumour responses were observed in forty-two of the sixty patients (70%; 95% confidence interval 57–81%), including nineteen patients (32%) with a complete response. Whole-exome sequencing revealed a high mutational load and frequent NF1 mutations (fourteen out of seventeen cases) in these tumours. Immunohistochemistry analysis from nineteen desmoplastic melanomas and thirteen non-desmoplastic melanomas revealed a higher percentage of PD-L1-positive cells in the tumour parenchyma in desmoplastic melanomas (P = 0.04); these cells were highly associated with increased CD8 density and PD-L1 expression in the tumour invasive margin. Therefore, patients with advanced desmoplastic melanoma derive substantial clinical benefit from PD-1 or PD-L1 immune checkpoint blockade therapy, even though desmoplastic melanoma is defined by its dense desmoplastic fibrous stroma. The benefit is likely to result from the high mutational burden and a frequent pre-existing adaptive immune response limited by PD-L1 expression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Shain, A. H. et al. Exome sequencing of desmoplastic melanoma identifies recurrent NFKBIE promoter mutations and diverse activating mutations in the MAPK pathway. Nat. Genet. 47, 1194–1199 (2015)
Ribas, A. et al. Association of pembrolizumab with tumor response and survival among patients with advanced melanoma. J. Am. Med. Assoc. 315, 1600–1609 (2016)
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015)
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015)
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016)
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016)
Han, D. et al. Clinicopathologic predictors of survival in patients with desmoplastic melanoma. PLoS One 10, e0119716 (2015)
Busam, K. J. et al. Cutaneous desmoplastic melanoma: reappraisal of morphologic heterogeneity and prognostic factors. Am. J. Surg. Pathol. 28, 1518–1525 (2004)
Alexandrov, L. B. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013)
Wiesner, T. et al. NF1 mutations are common in desmoplastic melanoma. Am. J. Surg. Pathol. 39, 1357–1362 (2015)
Krauthammer, M. et al. Exome sequencing identifies recurrent mutations in NF1 and RASopathy genes in sun-exposed melanomas. Nat. Genet. 47, 996–1002 (2015)
Hayward, N. K. et al. Whole-genome landscapes of major melanoma subtypes. Nature 545, 175–180 (2017)
Akbani, R. et al. Genomic classification of cutaneous melanoma. Cell 161, 1681–1696 (2015)
Shin, D. S. et al. Primary resistance to PD-1 blockade mediated by JAK1/2 mutations. Cancer Discov. 7, 188–201 (2017)
Roh, W. et al. Integrated molecular analysis of tumor biopsies on sequential CTLA-4 and PD-1 blockade reveals markers of response and resistance. Sci. Transl. Med. 9, eaah3560 (2017)
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015)
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N. Engl. J. Med. 375, 819–829 (2016)
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014)
Daud, A. I. et al. Programmed death-ligand 1 expression and response to the anti-programmed death 1 antibody pembrolizumab in melanoma. J. Clin. Oncol. 34, 4102–4109 (2016)
Frydenlund, N. et al. Tumoral PD-L1 expression in desmoplastic melanoma is associated with depth of invasion, tumor-infiltrating CD8 cytotoxic lymphocytes and the mixed cytomorphological variant. Mod. Pathol. 30, 357–369 (2017)
Ansell, S. M. et al. PD-1 blockade with nivolumab in relapsed or refractory Hodgkin’s lymphoma. N. Engl. J. Med. 372, 311–319 (2015)
Akbay, E. A. et al. Activation of the PD-1 pathway contributes to immune escape in EGFR-driven lung tumors. Cancer Discov. 3, 1355–1363 (2013)
Casey, S. C. et al. MYC regulates the antitumor immune response through CD47 and PD-L1. Science 352, 227–231 (2016)
Dorand, R. D. et al. Cdk5 disruption attenuates tumor PD-L1 expression and promotes antitumor immunity. Science 353, 399–403 (2016)
Kataoka, K. et al. Aberrant PD-L1 expression through 3′-UTR disruption in multiple cancers. Nature 534, 402–406 (2016)
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012)
Jiang, H. et al. Targeting focal adhesion kinase renders pancreatic cancers responsive to checkpoint immunotherapy. Nat. Med. 22, 851–860 (2016)
Nghiem, P. T. et al. PD-1 blockade with pembrolizumab in advanced Merkel-cell carcinoma. N. Engl. J. Med. 374, 2542–2552 (2016)
Hamid, O. et al. Safety and tumor responses with lambrolizumab (anti-PD-1) in melanoma. N. Engl. J. Med. 369, 134–144 (2013)
Rittmeyer, A. et al. Atezolizumab versus docetaxel in patients with previously treated non-small-cell lung cancer (OAK): a phase 3, open-label, multicentre randomised controlled trial. Lancet 389, 255–265 (2017)
Acknowledgements
This study was funded in part by the Grimaldi Family Fund, the Parker Institute for Cancer Immunotherapy, National Institutes of Health (NIH) grants R35 CA197633 and P01 CA168585, the Ressler Family Fund, the Samuels Family Fund and the Garcia-Corsini Family Fund (to A.R.). Z.E. was supported in part by the Moffitt Cancer Center NCI Skin SPORE (5P50CA168536) and Moffitt’s Total Cancer Care Initiative and Collaborative Data Services (P30-CA076292) for this work. J.M.Z. is part of the UCLA Medical Scientist Training Program supported by NIH training grant GM08042. S.H.-L. was supported by a Young Investigator Award and a Career Development Award from the American Society of Clinical Oncology (ASCO), a Tower Cancer Research Foundation Grant, and a Dr. Charles Coltman Fellowship Award from the Hope Foundation. We acknowledge the Translational Pathology Core Laboratory (TPCL) and R. Guo, W. Li, J. Pang and M. H. Macabali from UCLA for blood and biopsy processing, and X. Li, L. Dong, J. Yoshizawa, and J. Zhou from the UCLA Clinical Microarray Core for sequencing expertise. G.V.L. is supported by an NHMRC Fellowship and The University of Sydney Medical Foundation. R.A.S. is supported by an NHMRC Fellowship.
Author information
Authors and Affiliations
Contributions
Z.E., J.M.Z., S.H.-L. and A.R. developed the concepts. Z.E., S.H.-L, J.M.Z., and A.R. designed the experiments. Z.E., J.M.Z., S.H.-L. and A.R. interpreted the data. S.H.-L., I.P.S. and Z.E. performed IHC analyses. J.M.Z. performed genomic analyses. Z.E., A.R., B.C., D.W.K., A.A., D.B.J., E.L., B.K., R.M., S.R., J.A.S., R.J., M.A.P., M.S.C, W.-J.H., and G.V.L. clinically evaluated patients and contributed clinical data and tumour samples. R.A.S., J.M., and A.J.C. evaluated tumour samples. P.F.G. conducted the heat map analysis. X.W. performed statistical analyses. C.W. evaluated the non-DM clinical data. Z.E., J.M.Z., S.H.-L. and A.R. wrote the manuscript. S.H.-L. and A.R. supervised the project. All authors contributed to the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
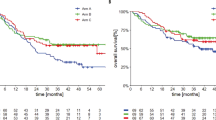
Extended Data Figure 1 Survival data for the DM cohort.
a, Progression-free survival (PFS), n = 60, median not reached. b, Overall survival (OS), n = 60, median not reached.
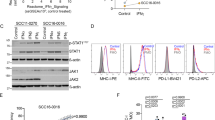
Extended Data Figure 3 Mutational analysis in the desmoplastic melanoma cohort.
a, Analysis of mutational load in samples obtained from primary locally advanced cases and metastatic lesions. Two-sided Wilcoxon Mann–Whitney rank sum test, P = 0.16 (95% CI, −171 to 1,175). b, Scores from the loss-of-function (LOF) SigRank algorithm1 show enrichment for LOF mutations (nonsense, frameshift, splice-site or damaging missense) compared to the expected number based on the rate of LOF mutations in the cohort. Solid line corresponds to observed/expected ratio of 1.0. c, Mutational load in the vanAllen16 anti-CTLA4 treated cohort separated by driver subtype and coloured by response. In the box plots, line is median, box is 25th to 75th percentile, whiskers show highest and lowest values within 1.5 × interquartile range.
Extended Data Figure 4 Mutations in antigen-presenting machinery or enriched by response in the DM cohort.
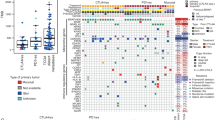
a, Mutations in genes enriched in responders (n = 12) (blue) or non-responders (n = 5) (red). Shown are genes with P < 0.05 by unadjusted two-sided Fisher’s exact test of samples with or without a non-synonymous mutation between responders and non-responders. None were significant after false-discovery rate adjustment. b, Mutations in antigen-presenting machinery genes. Tiling plot shows mutations in a given gene (rows) per sample (columns). Colour indicates mutation type, with truncating mutations (frameshift, nonsense, splice-site) in red, missense in green. Darker colour intensity indicates potentially homozygous mutations, with variant allele frequency more than 1.5 times the sample median.
Extended Data Figure 5 Patterns of CD8 infiltration and PD-L1 expression in biopsies from patients with DM and non-DM tumours.
a–e, Using cut off of >10% for high CD8 density in either parenchyma or invasive margins and >15% for high PD-L1 expression, five different patterns were identified. a, High CD8 density, high PD-L1 in tumour parenchyma higher than in invasive margins. b, High CD8 density, high PD-L1 in invasive margins higher than in tumour parenchyma. c, High CD8 density, high PD-L1 in the invasive margins only. d, Low CD8 density, high PD-L1. e, Low CD8 density, low PD-L1 expression. f, Yellow lines delineate the edges of tumour regions determined by positive S100 staining. Green or red lines mark the invasive margins around the tumour edges. All analysis was done with HALO software (Indica Labs). g, Heat map summary of patterns of CD8 and PD-L1 expression in biopsies from patients with DM and CM, based on their response to anti-PD-1 or anti-PD-L1 treatment. Intensity of colour coding indicates number of cases in each category. All calculations were based on scanned whole tumour images.
Extended Data Figure 6 CD8 density and PD-L1 expression in the tumour parenchyma and invasive margins in biopsies of patients with DM and non-DM tumours.
a, CD8 staining in the invasive margin. b, PD-L1 staining in the invasive margin. c, CD8 staining in the tumour centre. d, PD-L1 staining in the tumour centre. The percentage of positively stained cells in all nucleated cells is shown. CB, clinical benefit; PD, progressive disease. All calculations used two-sided Mann–Whitney rank sum test. See Supplementary Table for all statistical analyses. Asterisk indicates statistical significance. Tumour, tumour centre.
Extended Data Figure 7 Correlation between CD8 and PD-L1 in the invasive margin or tumour parenchyma in DM.
Black squares represent a sample from a patient who had a good response in the lesion biopsied (analysed) but was found to have brain metastasis shortly after treatment started. See Supplementary Table for further statistical analyses. IM, invasive margin.
Extended Data Figure 8 Hierarchical clustering of cases of DM and non-DM based on CD8 and PD-L1 expression in the invasive margin and tumour parenchyma.
a, Non-desmoplastic cutaneous melanomas (n = 13), with the y axis colour coded for response and mutational load. b, Desmoplastic melanomas (n = 19), with the additional information of differentiation between pure (red) and mixed (blue) histology on the y axis. For mutational load, darker squares correspond to higher mutational load. Gray squares are missing data points.
Supplementary information
Supplementary Information
Life Sciences Reporting Summary (PDF 214 kb)
Rights and permissions
About this article
Cite this article
Eroglu, Z., Zaretsky, J., Hu-Lieskovan, S. et al. High response rate to PD-1 blockade in desmoplastic melanomas. Nature 553, 347–350 (2018). https://doi.org/10.1038/nature25187
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25187
This article is cited by
-
Identifying microRNAs associated with tumor immunotherapy response using an interpretable machine learning model
Scientific Reports (2024)
-
Schwierig zu therapierende Melanome
hautnah dermatologie (2024)
-
Comprehensive analysis of necroptotic patterns and associated immune landscapes in individualized treatment of skin cutaneous melanoma
Scientific Reports (2023)
-
Immune microenvironment and lymph node yield in colorectal cancer
British Journal of Cancer (2023)
-
PDJ amplicon in triple negative breast cancer
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.