Abstract
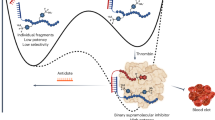
More effective therapies are urgently needed against hepatitis C virus (HCV), a major cause of viral hepatitis. We used in vitro protein expression and microfluidic affinity analysis to study RNA binding by the HCV transmembrane protein NS4B, which plays an essential role in HCV RNA replication. We show that HCV NS4B binds RNA and that this binding is specific for the 3′ terminus of the negative strand of the viral genome with a dissociation constant (Kd) of ∼3.4 nM. A high-throughput microfluidic screen of a compound library identified 18 compounds that substantially inhibited binding of RNA by NS4B. One of these compounds, clemizole hydrochloride, was found to inhibit HCV RNA replication in cell culture that was mediated by its suppression of NS4B's RNA binding, with little toxicity for the host cell. These results yield new insight into the HCV life cycle and provide a candidate compound for pharmaceutical development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Liang, T.J., Rehermann, B., Seeff, L.B. & Hoofnagle, J.H. Pathogenesis, natural history, treatment, and prevention of hepatitis C. Ann. Intern. Med. 132, 296–305 (2000).
Reed, K.E. & Rice, C.M. Overview of hepatitis C virus genome structure, polyprotein processing, and protein properties. Curr. Top. Microbiol. Immunol. 242, 55–84 (2000).
Rice, C.M. Flaviviridae: The viruses and their replication. in Fields Virology (eds. Fields, B.N., Knipe, D.M. & Howley, P.M.) 991–1041 (Lippincott-Raven Publications, Philadelphia, 1996).
Egger, D. et al. Expression of hepatitis C virus proteins induces distinct membrane alterations including a candidate viral replication complex. J. Virol. 76, 5974–5984 (2002).
Elazar, M., Liu, P., Rice, C.M. & Glenn, J.S. An N-terminal amphipathic helix in hepatitis C virus (HCV) NS4B mediates membrane association, correct localization of replication complex proteins, and HCV RNA replication. J. Virol. 78, 11393–11400 (2004).
Gosert, R. et al. Identification of the hepatitis C virus RNA replication complex in Huh-7 cells harboring subgenomic replicons. J. Virol. 77, 5487–5492 (2003).
El-Hage, N. & Luo, G. Replication of hepatitis C virus RNA occurs in a membrane-bound replication complex containing nonstructural viral proteins and RNA. J. Gen. Virol. 84, 2761–2769 (2003).
Kusov, Y.Y., Probst, C., Jecht, M., Jost, P.D. & Gauss-Müller, V. Membrane association and RNA binding of recombinant hepatitis A virus protein 2C. Arch. Virol. 143, 931–944 (1998).
Rodríguez, P.L. & Carrasco, L. Poliovirus protein 2C contains two regions involved in RNA binding activity. J. Biol. Chem. 270, 10105–10112 (1995).
Echeverri, A.C. & Dasgupta, A. Amino terminal regions of poliovirus 2C protein mediate membrane binding. Virology 208, 540–553 (1995).
Einav, S., Elazar, M., Danieli, T. & Glenn, J.S. A nucleotide binding motif in hepatitis C virus (HCV) NS4B mediates HCV RNA replication. J. Virol. 78, 11288–11295 (2004).
Overington, J.P. Al-Lazikani, B. & Hopkins AL. How many drug targets are there? Nat. Rev. Drug Discov. 5, 993–996 (2006).
Lundin, M., Monné, M., Widell, A., Von Heijne, G. & Persson, M.A.A. Topology of the membrane-associated hepatitis C virus protein NS4B. J. Virol. 77, 5428–5438 (2003).
Hadd, A.G., Raymond, D.E., Halliwell, J.W., Jacobson, S.C. & Ramsey, J.M. Microchip device for performing enzyme assays. Anal. Chem. 69, 3407–3412 (1997).
Maerkl, S.J. & Quake, S.R. A systems approach to measuring the binding energy landscapes of transcription factors. Science 315, 233–237 (2007).
Toepke, M.W. & Beebe, D.J. PDMS absorption of small molecules and consequences in microfluidic applications. Lab Chip 6, 1484–1486 (2006).
Lee, J.N., Park, C. & Whitesides, G.M. Solvent compatibility of poly(dimethylsiloxane)-based microfluidic devices. Anal. Chem. 75, 6544–6554 (2003).
Whitesides, G.M. The origins and the future of microfluidics. Nature 442, 368–373 (2006).
Kang, L., Chung, B.G., Langer, R. & Khademhosseini, A. Microfluidics for drug discovery and development: from target selection to product lifecycle management. Drug Discov. Today 13, 1–13 (2008).
Myer, V.E., Fan, X.C. & Steitz, J.A. Identification of HuR as a protein implicated in AUUUA-mediated mRNA decay. EMBO J. 16, 2130–2139 (1997).
Park, S., Myszka, D.G., Yu, M., Littler, S.J. & Laird-Offringa, I.A. HuD RNA recognition motifs play distinct roles in the formation of a stable complex with AU-rich RNA. Mol. Cell. Biol. 20, 4765–4772 (2000).
Park-Lee, S., Kim, S. & Laird-Offringa, I.A. Characterization of the interaction between neuronal RNA-binding protein HuD and AU-rich RNA. J. Biol. Chem. 278, 39801–39808 (2003).
Glenn, J.S., Taylor, J.M. & White, J.M. In vitro-synthesized hepatitis delta virus RNA initiates genome replication in cultured cells. J. Virol. 64, 3104–3107 (1990).
Glenn, J.S., Watson, J.A., Havel, C.M. & White, J.M. Identification of a prenylation site in delta virus large antigen. Science 256, 1331–1333 (1992).
Burd, C.G. & Dreyfuss, G. Conserved structures and diversity of functions of RNA-binding proteins. Science 265, 615–621 (1994).
Wung, C.H. et al. Identification of the RNA-binding sites of the triple gene block protein 1 of bamboo mosaic potexvirus. J. Gen. Virol. 80, 1119–1126 (1999).
Spångberg, K., Wiklund, L. & Schwartz, S. HuR, a protein implicated in oncogene and growth factor mRNA decay, binds to the 3′ ends of hepatitis C virus RNA of both polarities. Virology 274, 378–390 (2000).
Tscherne, D.M. et al. Time- and temperature-dependent activation of hepatitis C virus for low-pH-triggered entry. J. Virol. 80, 1734–1741 (2006).
Lindenbach, B.D. et al. Complete replication of hepatitis C virus in cell culture. Science 309, 623–626 (2005).
Elazar, M. et al. Amphipathic helix-dependent localization of NS5A mediates hepatitis C virus RNA replication. J. Virol. 77, 6055–6061 (2003).
Tong, X. et al. Identification and analysis of fitness of resistance mutations against the HCV protease inhibitor SCH 503034. Antiviral Res. 70, 28–38 (2006).
Dimitrova, M., Imbert, I., Kieny, M.P. & Schuster, C. Protein-protein interactions between hepatitis C virus nonstructural proteins. J. Virol. 77, 5401–5414 (2003).
Blight, K.J., Kolykhalov, A.A. & Rice, C.M. Efficient initiation of HCV RNA replication in cell culture. Science 290, 1972–1974 (2000).
Huang, L. et al. Hepatitis C virus nonstructural protein 5A (NS5A) is an RNA-binding protein. J. Biol. Chem. 280, 36417–36428 (2005).
Roosild, T.P. et al. NMR structure of Mistic, a membrane-integrating protein for membrane protein expression. Science 307, 1317–1321 (2005).
Schilders, G., Raijmakers, R., Raats, J.M.H. & Pruijn, G.J.M. MPP6 is an exosome-associated RNA-binding protein involved in 5.8S rRNA maturation. Nucleic Acids Res. 33, 6795–6804 (2005).
Acknowledgements
This work was supported by a Burroughs Wellcome Fund Clinical Scientist Award in Translational Research (to J.S.G.) and National Institutes of Health (NIH) RO1 DK066793, an NIH Director's Pioneer Award (to S.R.Q.) and NIH 1RO1 HG002644-01A1. D.G. was supported in part by the Fulbright Foundation. S.E. is a recipient of an American Liver Foundation Postdoctoral Fellowship Award. We also wish to thank the Stanford University SPARK Program, High-Throughput Bioscience Center and the Microfluidic Foundry. The plasmid FL-J6/JFH-5′C19Rluc2AUbi that consists of the full-length HCV genome and expresses Renilla luciferase was a gift from Charles M. Rice, Rockefeller University.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Stanford University has filed a patent application on the results from this work.
Supplementary information
Supplementary Text and Figures
Figures 1–6, Data (PDF 217 kb)
Rights and permissions
About this article
Cite this article
Einav, S., Gerber, D., Bryson, P. et al. Discovery of a hepatitis C target and its pharmacological inhibitors by microfluidic affinity analysis. Nat Biotechnol 26, 1019–1027 (2008). https://doi.org/10.1038/nbt.1490
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1490
This article is cited by
-
Systematic analysis of low-affinity transcription factor binding site clusters in vitro and in vivo establishes their functional relevance
Nature Communications (2022)
-
Microfluidic device for real-time formulation of reagents and their subsequent encapsulation into double emulsions
Scientific Reports (2018)
-
Chemical genetics-based development of small molecules targeting hepatitis C virus
Archives of Pharmacal Research (2017)
-
SELMAP - SELEX affinity landscape MAPping of transcription factor binding sites using integrated microfluidics
Scientific Reports (2016)