Abstract
Considerable progress has been made in converting human pluripotent stem cells (hPSCs) into functional neurons. However, the protracted timing of human neuron specification and functional maturation remains a key challenge that hampers the routine application of hPSC-derived lineages in disease modeling and regenerative medicine. Using a combinatorial small-molecule screen, we previously identified conditions to rapidly differentiate hPSCs into peripheral sensory neurons. Here we generalize the approach to central nervous system (CNS) fates by developing a small-molecule approach for accelerated induction of early-born cortical neurons. Combinatorial application of six pathway inhibitors induces post-mitotic cortical neurons with functional electrophysiological properties by day 16 of differentiation, in the absence of glial cell co-culture. The resulting neurons, transplanted at 8 d of differentiation into the postnatal mouse cortex, are functional and establish long-distance projections, as shown using iDISCO whole-brain imaging. Accelerated differentiation into cortical neuron fates should facilitate hPSC-based strategies for disease modeling and cell therapy in CNS disorders.
Similar content being viewed by others
Main
Over the past few years, methods have been developed to convert hPSCs into early neural lineages. A particularly efficient strategy is the use of two small-molecule inhibitors of SMAD signaling (LDN193189 and SB431542; referred to as the LSB protocol) to trigger differentiation of human embryonic stem cells (hESCs) or human induced pluripotent stem cells (hiPSCs) into PAX6+ CNS neural precursors within 11 d of differentiation1. Neural subtype specification can be further modulated using additional small molecules targeting pathways such as WNT signaling. Timed exposure to compounds activating WNT signaling under dual SMAD inhibition conditions induces the neural crest lineage, marked by SOX10 expression. In contrast, inhibition of WNT signaling blocks the formation of neural crest cells and enhances the induction of forebrain precursors, marked by FOXG1 expression2,3,4. Although those manipulations efficiently specify defined neural precursor cell populations, further differentiation into functional neurons in vitro is a lengthy process that can extend over weeks if not months.
To accelerate neuronal fate acquisition, we have used two additional small molecules: SU5402, a potent inhibitor of fibroblast growth factor (FGF) signaling5, and DAPT, a γ-secretase inhibitor blocking Notch signaling6. Combinatorial application of those two inhibitors with dual SMAD inhibition and WNT activation yields 75% post-mitotic neurons in 11 d of differentiation7, the same time period required for neural precursor cell induction under standard dual SMAD inhibition conditions1. However, coexpression of BRN3A and ISL1 in those rapidly induced neurons defined them as peripheral sensory rather than PAX6-derived CNS neurons7. Therefore, it has remained unclear whether strategies to accelerate neuronal fate acquisition during sensory fate specification can be adapted for CNS fates. PAX6-derived cortical neurons are of particular interest for studies of human development and for modeling human neurodevelopmental and neurodegenerative disorders. While reliable protocols exist to derive cortical neurons from hPSCs, those conditions require 30–90 d of differentiation to yield both lower- and upper-layer cortical neurons8,9 and even longer time periods to achieve full maturation.
Here we sought to identify small-molecule-based conditions that accelerate human cortical neuron fate induction to facilitate the routine application of hPSC-derived neurons in disease modeling and regenerative medicine. We present modified small-molecule-based differentiation conditions that sharply accelerate the derivation of forebrain neurons in the presence of WNT pathway inhibitors. Phenotypic marker expression, electrophysiological analysis, and in vivo transplantation studies confirm neuronal identity and functional properties of the cells, indicating that strategies for accelerated neuronal fate acquisition can be adapted for derivation of both peripheral sensory and CNS fates.
Results
Development of an accelerated CNS neuron differentiation protocol
Given the critical roles of WNT signaling in determining fate choice between the CNS and neural crest3,10, we hypothesized that developing a combinatorial small-molecule approach that inhibits rather than activates WNT signaling may trigger rapid differentiation toward cortical neurons (Fig. 1a). To test this hypothesis, we replaced the GSK3β inhibitor CHIR99021 (C; WNT agonist) with the tankyrase inhibitor XAV939 (X; WNT antagonist), which acts to stabilize Axin11. All other inhibitors used previously for the derivation of sensory neurons (LSB, SU5402 (S; FGF antagonist), and DAPT (D; Notch antagonist)) remained unchanged for these initial studies aimed at rapidly inducing forebrain neuron fates (LSB+X/S/D protocol).
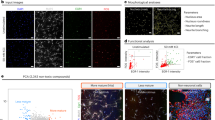
(a) Illustration of pathway manipulations used to generate neural crest (NC) versus forebrain fates and neural precursor versus accelerated neuronal fates. SOX10+ neural crest precursors, PAX6+ CNS precursors and BRN3A+ and ISL1+ sensory neurons have been previously reported, whereas the current study is focused on the rapid induction of cortical neurons from hPSCs. (b) Validation of hESC-based reporter lines for PAX6::H2B-GFP, SOX10::GFP and SIX1::H2B-GFP by assessment of colabeling with matched protein markers. (c) Time-course quantitative analysis of CNS (PAX6::H2B-GFP, left), NC (SOX10::GFP, middle) and placode (SIX1::H2B-GFP, right) induction under the various protocols. LSB, dual SMAD inhibition; XAV, XAV939; CHIR, CHIR99021; S, SU5402; D, DAPT. N = 3 independent batches of cell cultures per condition and cell line. (d) Immunocytochemistry for PAX6, SOX10 and TUJ1 during CNS- versus NC-based hPSC differentiation. (e) CNS- and NC-based hPSC differentiation in combination with S/D exposure to accelerate neuronal fate acquisition. (f) Quantification of TUJ1 expression data from d and e at day 13 of differentiation by intracellular flow. Black dots represent values from independent experiments. From left to right, N = 3, 3, 5, 3. Statistical analysis was done by unpaired t-test with Welch's correction (two-tailed). XAV vs. XAV+S/D: t = 9.510, df = 2.160, P = 0.0084. CHIR+S/D vs. XAV+S/D: t = 13.40, df = 4.718, P < 0.0001. (g) P, S single and combinatorial dose response analysis based on quantification of the percentages of Nestin+ and TUJ1+ cells by intracellular flow cytometry (top) and an estimate (bottom) of the total neurons in culture per cm2 (for every hPSC plated at day 0 under P1S5D and P8S10D conditions, we obtained ∼0.7 and 0.2 neurons, respectively, at day 13). P = PD0325901, ERK/MEK inhibitor. The digits following P, S, D represent the respective concentrations in μM. Each column represents N = 2 technical replicates per marker screened. (h) Immunocytochemistry for PAX6 and TUJ1 in P1S5D and P8S10D cultures at day 13. (i) Quantification of TUJ1+ cells by intracellular flow cytometry at day 13. Black dots represent values from independent experiments. From left to right, N = 4, 5, 5, 4. Statistical analysis was done using unpaired t-test with Welch's correction (two-tailed). XAV+P1D vs. XAV+P1S5D: t = 4.057, df = 6.045, P = 0.0066. XAV+P1S5 vs. XAV+P1S5D: t = 20.62, df = 4.880, P < 0.0001. XAV+P1S5D vs. XAV+P8S10D: t = 13.26, df = 6.397, P < 0.0001. Scale bars, 100 μm (b) or 200 μm (d,e,h). Error bars represent s. e. m. **P < 0.01, ****P < 0.0001.
In light of our past experience in unexpectedly triggering a CNS to peripheral nervous system (PNS) fate switch during rapid neuronal induction7, we first assessed the impact of the LSB+X/S/D condition on early ectodermal lineage choice using three engineered hESC reporter lines: PAX6::H2B-GFP (CNS lineage), SOX10::GFP2,7 (neural crest fate), and SIX1::H2B-GFP line12 (cranial placode fate). Faithfulness of reporter expression was validated after directed in vitro differentiation into the respective fates1,2,13 (Fig. 1b). Consistent with previous work, both LSB and LSB+X conditions gave rise to a near-uniform population (>95%) of PAX6+ cells, with few cells expressing SOX10 or SIX1 (Fig. 1c). In contrast, LSB+C or LSB+C/S/D (also referred to as 3i (ref. 7) or PNS sensory neuron protocol) gave rise to only a few cells expressing PAX6, but a large percentage of SOX10+ neural crest precursors, consistent with the important role of WNT signaling in neural crest induction7. LSB+X/S/D, our candidate protocol for rapid induction of CNS neurons, gave rise to an almost pure population (>98%) of PAX6+ cells as early as day 6 of differentiation. This accelerated timing was consistent with the role for FGF inhibition in exiting pluripotency in hPSCs14 (Fig. 1c).
Next, we assessed whether the LSB+X/S/D condition can induce putative CNS neurons with efficiencies similar to those reported for the rapid induction of sensory neurons7. Intracellular flow cytometry for β-III tubulin (TUJ1), a pan-neuronal marker, showed that the LSB+X/S/D condition gave rise to only 10% TUJ1+ neurons at day 13 of differentiation compared to >40% in the 3i condition7 (Fig. 1d–f). These data indicate that inhibition of WNT signaling in the presence of SU5402 and DAPT rapidly induced CNS lineage but triggered neuronal differentiation at only moderate efficiency compared to PNS sensory neuron conditions.
In an effort to enhance the neuronal conversion efficiency, we performed a targeted small-molecule screen using the CNS differentiation condition without acceleration (LSB+X). We selected molecules targeting signaling pathways involved in neural precursor cell proliferation, such as SHH (cyclopamine, Cur-61414, purmorphamine), PI3K and PDGFR (LY-294002, imatinib), MYC/bromodomain proteins (JQ1), retinoid signaling (all-trans retinoic acid), TGF-β activation (IDE-1), HMG-CoA reductase inhibition (lovastatin) and the nicotinamide phosphoribosyltransferase inhibition (P7C3), as well as signaling pathways downstream of FGF receptor activation, including ERK signaling (PD0325901). Only PD0325901, an orally bioavailable, potent inhibitor for mitogen-activated protein kinase (MAPK/ERK kinase or MEK)10, enhanced neuronal differentiation under LSB+X. Inhibition of ERK1/2 in the mouse causes premature neuronal differentiation during cortical development15, and ERK inhibition has been previously proposed as a strategy to enhance overall neuronal differentiation in hPSCs16. PD0325901 could boost the yield of TUJ1+ neurons to >50%, a value approximating the efficiency of the 3i protocol for generating sensory neurons, but only when used at high concentrations (Fig. 1g, top), resulting in a low yield in total cell numbers (Fig. 1g, bottom). Improved yields in total neuron numbers were obtained when lower PD0325901 concentrations were combined with SU5402 exposure in an effort to balance neuronal induction efficiency with overall cell loss. Additional treatment with DAPT did not affect overall neuron yield but further increased the efficiency of neuronal induction. To understand whether reduced yield is due to rapid cell cycle exit or direct toxicity, we measured phospho-histone 3 and cleaved caspase 3 as markers of cell proliferation and death, respectively. Exposure to both PD0325901 (P) and SU5402 reduced cell proliferation as early as 24 h after P/S treatment, whereas cell death was observed only at high doses of SU5402 (Supplementary Fig. 1a–f), implicating restriction of precursor cell proliferation as a key factor in the rapid neuronal differentiation response. We selected two conditions for all subsequent studies that produced high percentages and high total yields of neurons: (i) PD0325901 (1 μM) + SU5402 (5 μM) + D (10 μM) is referred to as the 'P1S5D condition'; and (ii) PD0325901 (8 μM) + SU5402 (10 μM) + D (10 μM) as the 'P8S10D condition' (Fig. 1h,i).
Phenotypic analysis of CNS and cortical neuron identity
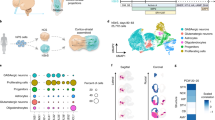
We next determined the efficiency of P1S5D and P8S10D as cocktails for the rapid induction of CNS neurons (Fig. 2a). The P8S10D condition resulted in the most dramatic acceleration of neuronal fate acquisition (Fig. 2b and Supplementary Fig. 2a), generating 70% of TUJ1+ neurons by 13 d of differentiation. Gene expression analysis confirmed downregulation of the pluripotency marker OCT4 and induction of neural and neuronal markers PAX6, FOXG1 and DCX, as well as markers of early-born cortical neurons, including TBR1 (preplate, subplate and layer VI) and REELIN, in LSB+X/P/S/D conditions. In contrast, the sensory neuron (3i) and the CNS induction protocol without acceleration (LSB+X) showed lack of cortical or neuronal marker induction, respectively (Fig. 2c). Given the weak induction of the forebrain marker FOXG1 under the P8S10D condition, we tested the impact of each small molecule on FOXG1 expression (Supplementary Fig. 2b,c). In particular, the exposure to PD0325901 dramatically reduced the efficiency of FOXG1 induction. However, TBR1 was expressed in >50% of TUJ1+ cells at day 13 in both P1S5D and P8S10D conditions (Fig. 2d–f), with similar expression of the layer VI marker TLE4 (Supplementary Fig. 3a).
(a) Differentiation scheme for KSR-based P1S5D and P8S10D neuronal induction protocol. hPSCs were plated 1 d before differentiation at 200,000 cells per cm2 in conditioned hESC medium supplemented with 10 ng/ml FGF2 and 10 μM ROCK-inhibitor Y-27632. Small molecules were added in the presence of dual SMAD inhibition (LDN193189, SB431542) and XAV939 treatment (LSB+X). Optimized timing for the application of PD0325901, SU5402 and DAPT (P/S/D) is shown. NB, neurobasal medium; BCA, BDNF, cAMP and ascorbic acid. (b) Time-course analysis of neuronal induction efficiency by intracellular flow cytometry (N = 4 independent batches of cell cultures), unpaired t-test with Welch's correction (two-tailed) to compare mean difference between each group at day 13. LSBX vs. P1S5D: t = 20.79, df = 3.042, P = 0.0002. 3i vs. P1S5D: t = 0.8277, df = 5.013, P = 0.4455, not significant. P1S5D vs. P8S10D: t = 12.79, df = 5.994, P < 0.0001. (c) Time-course qRT-PCR analysis at days 5, 8, 11 and 13 of differentiation. OCT4 (POU5F1), human pluripotency marker; PAX6, dorsal cortical progenitor marker; FOXG1, forebrain marker; DCX, pan-neuron marker; TBR1, preplate, subplate and cortical layer VI neuron marker; REELIN, cortical layer I (Cajal-Retzius cell) neuron marker; FC, fold change. N = 3 independent batches of cell cultures. (d–f) TBR1 and TUJ1 expression analyzed by immunocytochemistry at day 13 (d) and quantification of the percentage of TBR1+ cells among total cells (e) (t = 3.151, df = 7.820, P = 0.0140, two-tailed) or among TUJ1+ neurons (f) (t = 1.094, df = 9.997, P = 0.2994, two-tailed, not significant). Black dots represent values from quantification of uniform random selection of 6 150 μm × 150 μm areas from 3 independent batches of cell cultures. Statistical analysis was done using unpaired t-test with Welch's correction. (g) Alternative scheme of accelerated neuronal differentiation protocol using P/S/D in E6 medium. (h) Validation of P/S/D protocol in E6 by immunocytochemistry of PAX6, TBR1 and TUJ1 expression at day 13. Scale bars, 50 μm (d and h, right) or 100 μm (h, left and middle). Error bars represent s.e.m. *P < 0.05, ***P < 0.001, ****P < 0.0001.
To identify the remaining neurons that were negative for TBR1 and TLE4, we screened a panel of additional markers at day 13 (Supplementary Table 1 and Supplementary Fig. 3b). Surprisingly, 15–20% neurons expressed BRN3A but only very few neurons expressed ISL1, suggesting the presence of a contaminating BRN3A+ CNS lineage (Supplementary Table 1 and Supplementary Fig. 3c,d). On the basis of the expression of both BRN3A and GSX2 (Supplementary Fig. 3b), those noncortical neurons may correspond to an early thalamic lineage (http://www.gensat.org/; http://developingmouse.brain-map.org/). Characterization of the TUJ1− fraction showed the presence of cells positive for TBR2, BLBP and CUX2 (Supplementary Table 1 and Supplementary Fig. 3a,b), consistent with cortical precursor cell identity. We further observed upregulation (compared to LSB+X) of other anterior CNS and cortical progenitor and neuron markers in LSB+X/P/S/D conditions (Supplementary Fig. 3b). However, we did not detect the expression of ventral forebrain, cortical interneuron or other GABAergic neuron fates (Supplementary Table 1 and Supplementary Fig. 3b).
To determine whether LSB+X/P/S/D conditions were robust across multiple lines, we tested six independent hiPSC lines derived from two healthy individuals. By 13 d of differentiation, all lines were enriched for TBR1+/TUJ1+ neurons and displayed morphologies similar to those obtained from the WA09 hESC line (Supplementary Fig. 4a,b). Quantification of the percentages of neurons showed similar efficiency to that observed for WA09, although there was some variability across lines (Supplementary Fig. 4c). In an effort to translate the accelerated protocol to GMP-compatible culture conditions, the protocol was further adapted to an Essential 6 medium (E6)-based induction platform (Fig. 2g). We observed efficient PAX6 induction and accelerated generation of highly enriched populations of TBR1+ post-mitotic neurons by 13 d of differentiation (Fig. 2h). Thus, our rapid induction strategy can be applied across hiPSC lines and adapted to GMP-compatible culture conditions.
During corticogenesis, projection neurons are produced in an inside-out manner17. The enrichment of TBR1+ neurons suggested a potential bias toward generating the earliest-born deep-layer cortical neurons. However, further maintenance of P1S5D or P8S10D cultures in the absence of FGF-ERK and Notch inhibition (days 13–55) (Fig. 3a) enabled generation of neurons expressing markers representing a broader range of cortical layers, such as FOXP2 (layer V–VI), CTIP2 (layer V), SATB2 (layer II–III, V), and RGS4 (layers II–III, V) (Fig. 3b and Supplementary Fig. 5), as well as generation of upper layer CUX2+ (layers II–IV) neurons monitored by using a tamoxifen-inducible CUX2 reporter hESC line (Supplementary Fig. 6a–d). P1S5D- or P8S10D-treated cells started to produce CUX2+ post-mitotic neurons with mature morphologies as early as day 33, compared to day 55 using a protocol without acceleration (Supplementary Fig. 6e–g). While cortical neurogenesis was considerably accelerated, no upregulation of glial markers, such as GFAP, AQP4 or OLIG2 was observed. Similarly, there was no induction of retinal fate markers such as CHX10 (Supplementary Fig. 5). The quantification of TBR1+, CTIP2+ and SATB2+ neurons (Fig. 3c) suggested that in vitro–derived neurons may follow a temporal order of marker expression consistent with in vivo corticogenesis. To further address the specific timing of neuron subtype derivation in vitro, we performed birth-dating experiments (Fig. 3a). 5-Ethynyl-2-deoxyuridine (EdU) colabeling with layer-specific markers showed successive waves of cell birth (Fig. 3d,e). Thus, our data demonstrate highly efficient induction of layer VI and indicate the feasibility of accelerated derivation of upper-layer neurons using a modified small-molecule timing regimen.
(a) Illustration of long-term culture protocols. In vitro differentiation before day 8 is the same as described in Figure 2a. For long-term culture, P1S5D and P8S10D cells were passaged at day 8 of differentiation at 150,000/cm2 and 300,000/cm2, respectively, in NB/B27+BCA medium without additional inhibitors thereafter. Cells were fixed at various time points and processed for immunocytochemistry or RNA extraction and qRT-PCR analysis. (b) Immunocytochemistry analysis after long-term maintenance of P1S5D and P8S10D cells shows neurons constituting distinct cortical layer fates: FOXP2 (layer V–VI), TLE4 (layer VI), CTIP2 (layer V), SATB2 (layer II–III, V), RGS4 (layer II–III, layer V), CUX2 (cortical progenitors and layer II–IV). (c) Quantification of TBR1+, CTIP2+ and SATB2+ cells in total cell population using P1S5D differentiation. N = quantification of 6 randomly selected photo frames captured using a 20× objective from 2 independent batches of cell cultures. (d) Representative image showing colabeling of EdU and TBR1 and CTIP2 at day 40 of P1S5D differentiation. (e) Quantification of the percentage of EdU+ cells among marker-positive cells at day 40 of P1S5D differentiation. EdU was added to the cultures for 48 h at various time points of differentiation (as indicated on x axis), and the cultures were fixed at day 40. Dots represent values of quantification results of individual photo frames from 2 independent batches of cell cultures. From left to right, for TBR1, CTIP2, SATB2, respectively, N = 3,4,4; 0,0,0; 3,4,3; 3,3,4; 3,5,1; 1,3,4. Scale bars, 50 μm. Error bars represent s.e.m.
Rapid induction of neuronal function
We next addressed whether rapid induction of neuronal markers is paralleled by rapid in vitro maturation, such as the ability to spontaneously fire repetitive action potentials. Functional maturation of hPSC-derived neurons has been previously demonstrated8,9, with firing of action potentials typically occurring at about 50–100 d of differentiation. To investigate maturation, we cultured cells under P1S5D or P8S10D conditions for 8 d followed by an additional 8 d in (i) basal medium without any small molecules, (ii) addition of DAPT only or (iii) addition of DAPT with SU5402, PD0325901 and CHIR99021 (P/S/D/C) (Fig. 4a). The WNT agonist CHIR99021 was included for this final differentiation step, as it exerted a strong pro-survival effect on cultures maintained in P/S/D, and had been previously shown to promote neuronal differentiation including axonal outgrowth and synapse formation by activating canonical WNT signaling18,19. Notably, P8S10D cells maintained with P/S/D/C for only 8 d (day 16 of differentiation from pluripotent state) yielded neurons with mature electrophysiological properties characterized by the spontaneous firing of trains of action potentials at rest membrane potential or upon induced hyperpolarization after −10 pA current injection (Fig. 4b). In all conditions, 70–80% of the neurons recorded were capable of firing, with ∼20–30% neurons showed more mature firing patterns (∼train of ten action potential firing peaks) in P8S10D cells with P/S/D/C (Fig. 4c,d). Additional parameters of neuronal maturation include resting membrane potential, action potential half-width and rise rate (Tau) of initial firing, input resistance and maximum firing frequency (Fig. 4e). While maintaining P8S10D cells in P/S/D/C resulted in the most mature neuronal properties, even the mildest condition (P1S5D cells with basal medium) resulted in neurons with mature firing patterns by day 37 (Supplementary Fig. 7). This time frame is considerably faster than that achieved by most previous hPSC-based cortical neuron differentiation methods8,9 without the use of specialized neuronal recording media20, which may accelerate the onset of neuronal activity.
(a) Illustration of six different treatment conditions for neuronal differentiation and maintenance. In vitro differentiation protocol up to day 8 is the same as described in Figure 2a. P1S5D and P8S10D cells were then passaged at 150,000/cm2 and 300,000/cm2, respectively, at day 8 in NB/B27+BCA. Both passaged P1S5D and P8S10D cells were further maintained in three conditions: without inhibitors (none), with DAPT (D) and with PD0315901 (1 μM), SU5402 (5 μM), DAPT and CHIR99021 (3 μM) (PSDC), making 6 different treatments of cells in total. (b) Action potential firing traces at day 16 from cells representing the 6 differentiation conditions. (c,d) Quantification of percentage of cells with indicated firing frequencies at day 16 of differentiation without current injection (c) and with −10 pA current injection (d). From left to right, N = 15, 18, 11, 16, 17, 12 from 4 batches of independent cell cultures. (e) Quantification of auto-firing electrophysiological properties at day 16 (without current injection). Black dots represent values of individual cells. From left to right, N = 15, 18, 11, 16, 17, 12 from 4 batches of independent cell cultures. (f) Voltage-dependent sodium channel responses of P1S5D neurons without inhibitors at day 37 by whole-cell patch clamp. Responses could be blocked by TTX, which specifically blocks the sodium channel. Inset, protocol used for triggering sodium channel currents. (g) Spontaneous excitatory postsynaptic currents (sEPSCs) recorded in P1S5D neurons without inhibitors at day 40. sEPSCs could be blocked by NBQX, which selectively blocks AMPA receptors, indicating excitatory synaptic currents. Error bars represent s.e.m.
We observed robust voltage-dependent sodium channel responses that could be blocked by tetrodotoxin (TTX) (Fig. 4f). In addition, cells exhibited spontaneous excitatory postsynaptic currents that could be inhibited by NBQX, a specific AMPA receptor antagonist (Fig. 4g), indicating the formation of functional excitatory synapses. While those functional maturation data were obtained in the absence of any astrocyte co-culture, we tested whether the addition of astrocytes would further accelerate the maturation or promote the maintenance of the neurons. Indeed, culturing P8S10D-derived neurons on mouse astrocytes or in astrocyte-conditioned medium in the presence of DAPT (Supplementary Fig. 8a–c) improved overall neuronal survival and enabled long-term maintenance of rapidly induced neurons from 70 d to beyond 90 d (Supplementary Fig. 8d). Cultures on astrocytes further yielded neurons with decreased input resistance and enhanced morphological complexity, indicating increased neuronal maturity (Supplementary Fig. 8e–g).
In vivo studies using iDISCO whole-brain analysis
The in vitro data demonstrate that our combinatorial small-molecule protocols can rapidly induce cortical neurons with functional electrophysiological properties. However, to assess long-term survival and the capacity for axonal projections and integration into host circuitry, we performed in vivo transplantation studies. Immature neurons derived from day 8 cultures using an hESC line constitutively expressing EGFP were grafted into the somatosensory cortex of P2 NOD-SCID IL2Rgc–/– mice. Brains of the grafted animals were collected at 1–6 months after grafting, and subjected to whole-brain immunofluorescence imaging following the iDISCO21 clearing and whole-mount immunohistochemistry protocol (Fig. 5a). Most transplantation studies were carried out using the P1S5D-treated cells, which showed robust in vivo survival up to 6 months after transplantation, the latest time point tested in our study. P8S10D neurons showed more variable in vivo survival, with evidence of engraftment and axonal projections in only a subset of the animals at 1 month after transplantation (Supplementary Fig. 9a). Matched day 8 cells from the LSB+X condition showed extensive graft overgrowth with minimal evidence of neuronal differentiation or graft integration (Supplementary Fig. 9b). These data are reminiscent of previous results suggesting that early neuroepithelial, 'rosette-stage' cells result in tumor-like overgrowth22. Therefore, differentiation of neuroepithelial cells toward later-stage neural precursors or neurons is critical in reducing the risk of neural overgrowth.
(a) Analysis of the grafted brain at 1–6 months after grafting using whole-brain immunohistochemistry and imaging by light-sheet microscopy. Dorsal view of the graft core and its cortical projections at 1.5 months. (b) Projections (100 μm thick) showing the graft core morphology and the major projection regions (frontal cortex and corpus callosum). Dotted line, midline; arrow, aberrant longitudinal projections in the corpus callosum. (c) iDISCO-based imaging of half-brains showing the morphology of the graft projections at 1.5, 3 and 6 months. Top, views of the half-brain showing the graft cores and their major cortical projections. Middle, projections (100 μm thick) showing the details of the fiber morphology from the boxed regions and their increased branching over time. Bottom, projections (100 μm thick) showing human-specific synaptophysin (hSyn) colabeled with GFP in the hippocampus. (d) Immunohistochemical analysis of markers of cortical identity in graft-derived neurons identified by human-specific cytoplasm marker (SC121) or GFP expression at 1 month after grafting. N = 5 animals for 1, 1.5 and 3 months, and 2 animals for 6 months. Scale bars, 500 μm (a, b and c, top and middle) or 50 μm (c, bottom and d).
Analysis of brains grafted with P1S5D neurons at 1 and 1.5 months after grafting allowed visualization of the graft core and neuronal projections (Fig. 5b and Supplementary Video 1). After 1 month, GFP+-grafted cells developed extensive defasciculated projections across all cortical layers. A few long dense bundles were also consistently seen in cortical layer VI. Most of the projections terminated in the prefrontal motor cortex and frontal cortex, although many axons were also traced in the ipsilateral hippocampus and contralateral cortex through the corpus callosum (Fig. 5b and Supplementary Video 1). Very sparse graft-derived fibers were observed in the striatum, suggesting that grafted neurons preferred projecting across cortical regions rather than targeting subpallial regions. Using autofluorescence to map host axonal pathways (Supplementary Fig. 10a), we observed that the majority of graft-derived fiber bundles followed endogenous tracts (Supplementary Fig. 10b). However, some fibers projected outside of the host 'descending tracts' (Supplementary Fig. 10c). Overall, P1S5D grafts at 1 and 1.5 months after grafting showed enlarged terminal structures reminiscent of growth cones, a pattern characteristic of ongoing pathfinding with only limited terminal arborization (Fig. 5c, left). In contrast, by 3 months after transplantation and most pronounced at 6 months (Fig. 5c, middle and right, and Supplementary Video 2), there was extensive terminal arborization of human axons in matched target areas (Fig. 5c, middle versus left panel). Concomitant with extensive arborization, we also observed a dense network of human synaptophysin-positive structures colocalized with the GFP+ fibers in several target areas, such as the host hippocampus (Fig. 5c, bottom right). The grafted neurons exhibited a range of morphologies, with unipolar, bipolar, multipolar and pyramidal shapes (Supplementary Fig. 10d).
iDISCO was complemented with conventional immunohistochemical analyses, which confirmed in vivo cortical marker expression in human cells. These markers included general forebrain marker FOXG1, and layer-specific markers such as REELIN, SATB2 and CTIP2 (Fig. 5d). Expression of SATB2+ in the grafted neurons was consistent with commissural neuron identity, matching the presence of commissural axonal projections in the iDISCO studies (Fig. 5c and Supplementary Video 1). In addition, we gained preliminary evidence of in vivo function of grafted cells by electrophysiology (Supplementary Fig. 11). Our data indicated that P1S5D-induced neural cells at day 8 of differentiation were capable of in vivo survival and extensive axonal projections within the cortex. While P8S10D-induced neurons showed overall reduced graft size and viability, animals with surviving grafts showed extensive fiber outgrowth and arborization already at 1.5 months.
Discussion
We present a protocol to generate early-born cortical neurons, in particular of layer VI identity, that have mature electrophysiological properties by day 16 of differentiation and are capable of in vivo engraftment and long-distance projections in the postnatal mouse cortex (Fig. 6). This time frame is more than twice as fast as that demonstrated in previous cortical neuron differentiation protocols8,9,23,24,25, faster than most extrinsic-factor-based strategies for other hPSC-derived neuronal subtypes24,25,26 and similar in speed to protocols that rely on the forced expression of neurogenic transcription factors such as NGN2 (ref. 26). The use of small molecules avoids genetic modification and may offer greater flexibility than transcription factor–based methods. The cortical neurons derived under the current P1S5D or P8S10D conditions are biased toward deep cortical layers. Although our study was focused on manipulating the timing of neuron induction and maturation as independent parameters in hPSC differentiation protocols, we provide preliminary evidence that the culture conditions can be adapted to upper-layer neurons, albeit at a slower time scale.
Dual SMAD inhibition by LSB inhibits trophectoderm, mesendoderm and non-neural ectoderm cell fates, promoting CNS fates. XAV939 promotes anterior CNS identity, and SU5402 and PD0325901 (SU5402/PD0325901) accelerate exit from pluripotency toward neuroectodermal fates. A highly transient anterior neuroectodermal precursor state is driven toward post-mitotic cortical fates in the presence of DAPT and SU5402/PD0325901. Immature cortical neurons can acquire functional maturity in vitro by day 16 of differentiation, and day 8 neurons, grafted into neonatal mouse host brain, show widespread axonal projections and integration in cortex.
To our knowledge, there have been no previous reports using iDISCO to map hPSC-derived graft survival, axonal projections or host innervation. The iDISCO data include whole-brain immunohistochemistry and imaging for GFP, as well as imaging of human-specific markers such as human synaptophysin. This technology should be suitable for use with most human-specific markers to monitor graft biology. In future studies, it may be particularly interesting to apply iDISCO to map region-specific projections of defined hPSC-derived cortical lineages, such cells with selective cortical area and layer identity22, or to directly compare in vivo survival and projection patterns of distinct hPSC-derived neuronal subtypes injected at identical grafting sites. The assay could also serve as a tool to define neurons of related lineages but distinct projection patterns, such as midbrain dopamine neurons of A9 (substantia nigra) versus A10 (ventral tegmental area) identity, and to map terminal projection patterns of neurons placed at heterotopic27 versus orthotopic locations28.
Further studies are needed to optimize the derivation of layer-specific neurons and to understand the timing for generating deep versus upper layers. We propose that similar small-molecule acceleration strategies may also be developed for additional neuron subtypes. Such rapid, directed differentiation protocols will facilitate the generation of specific neuron subtypes relevant to diverse applications, including disease modeling, drug discovery and cell therapy.
Methods
hESC lines and hiPSC line generation.
hESCs (WA09; passages 32–60) were obtained from WiCell and maintained up to passage 60. The hESC SOX10::GFP bacterial artificial chromosome reporter line (WA09; passage 40–70) was generated as reported previously7. Constitutive EGFP+ hESC line (WA09; passage 35–60) was generated as reported29. RUES2 cell line used for deriving CUX2 conditional reporter line was generated at the Rockefeller University as described at http://rues.rockefeller.edu. For hiPSC induction, fibroblasts were prepared by digesting skin-punch biopsies following a protocol shared by M. Sheldon. Briefly, skin punches were digested in a mixture of collagenase (1%) and dispase (1 U/ml; Stem Cell Technology) in DMEM+10% FBS for 16–18 h at 37 °C in a tissue culture incubator. After digestion, the epidermal layer was discarded and the partially digested dermal layer was quartered onto the surface of a dry tissue culture dish and was left undisturbed for 2–5 min to encourage adhesion to the dish. DMEM+10% FBS was carefully added to the well so as not to detach the dermal layer. Cultures were fed every 3 d until confluent foci covered ∼2/3 of the well. Once confluent, cultures were passaged by trypsinization and expanded for 4–5 passages before reprogramming. iPSCs were made using the original CytoTune iPS Reprogramming Kit (A1378002; Life Technologies) using the manufacturer's protocol with a few modifications. Human ES medium containing 1 mM valproic acid (EMD Millipore) was added from days 2–9. After 2–3 weeks, individual iPS clones were picked and propagated as iPS lines. To verify that each of the three iPS subclones from a given individual were truly nonclonal, we picked colonies from 3 different wells that derived from 3 separate transductions. Each line was propagated for 10 passages before performing quality control assays. We first confirmed expression of OCT4, NANOG, SSEA-3, SSEA-4 and Tra-1-81. Clones that expressed all pluripotency markers were verified to have a normal karyotype by the Molecular Cytogenetics Core Facility at Memorial Sloan Kettering Cancer Center (MSKCC). The amount of Sendai vector present after 10 passages was quantified using the TaqMan iPSC Sendai Detection Kit (A13640; Life Technologies), and only clones with less than 0.01% Sendai virus amplicon (Mr04269880_mr) were used. All the hPSCs lines used for this study were tested for mycoplasma contamination every 2 weeks, and hPSC line identity was authenticated by STR analysis.
Generation of PAX6::H2B-GFP and SIX1::H2B-GFP lines (passages 40–65).
The PAX6-P2A-H2B-GFP and SIX1-P2A-H2B-GFP donor constructs were generated by performing In-Fusion cloning (Clontech) into the pUC19 backbone. Homology arms were generated by using genomic DNA, H2B:GFP was a gift from G. Wahl (plasmid #11680; Addgene), Pgk-Puro was amplified from the AAVS1 hPgk-PuroR-pA donor plasmid (a gift from R. Jaenisch (plasmid #22072; Addgene)). TALE nucleases were generated using the TALE-Toolbox provided by F. Zhang via Addgene30. Sequences targeting the stop codon of PAX6 were TGTCCTGTATTGTACCACT and TGTATACAAAGGTCCTTGT; for SIX1, were TCTCTGCTCGGCCCCCTCA and TTGGGGTCCTAAGTGGGGA. Briefly, 25 μg of donor plasmid and 5 μg of each TALEN were nucleofected into 10 × 106 WA09 hESCs. Puromycin selection was applied 72 h after nucleofection to isolate resistant clones. Clones were amplified and genomic PCRs confirming targeting were performed. All positive clones used had normal karyotype.
Generation of transgenic CUX2 conditional reporter line.
The CUX2::CreERT2/AAVS1-CAG::FLEX/tdTomato line was created in the RUES2 background by two sequential nucleofection and selection cycles. In the first round, 2 μg of CUX2::CreERT2/FRT-Puro-FRT-TK homology donor was electroporated into 2 × 106 early passage hESCs together with TALENs targeting the CUX2 initiation codon. Nucleofection was carried out using Amaxa nucleofector solution L (Lonza). Single cells were obtained by treating cultures with Accutase (Innovative Cell Technology), and cells were maintained in the ROCK-inhibitor Y-27632 (10 μM; Tocris) after nucleofection for 3 d. Nucleofected cells were subsequently grown for 2 weeks in puromycin selection medium maintained for the initial 10 d. Ganciclovir (2 μM) was also added for negative selection of random integrations. After 2 weeks, 22 clones were selected for further characterization by PCR genotyping, sequencing and karyotyping. One clone, which satisfied all criteria, was expanded and subjected to a second round of nucleofection with 2 μg of AAVS1 CAG::FLEX tdTomato/BSD homology donor, 0.5 μg each of AAVS1 right and left TALENs (Addgene), and 2 μg pCAG-Flpe (Addgene). The Flp recombinase was added to excise the FRT-Puro-FRT cassette from the transgene in the CUX2 locus. The nucleofected cells were then grown for 2 weeks in blasticidin selection. 12 clones were subsequently expanded for PCR genotyping and confirmed for excision of the FRT-Puro-FRT cassette. Out of the clones that were found to carry the transgene, one clone was karyotyped and chosen for further experiments. A list of primers used for genotyping is provided in Supplementary Table 2.
Culture of undifferentiated cells and neuronal induction (days 0–13 of differentiation).
hPSC lines were maintained with mouse embryonic fibroblasts (MEFs; Globalstem) pre-plated at 16,000 cells/cm2 on gelatin-coated tissue culture plate. Medium contained DMEM/F12, 20% (v/v) knockout serum replacement, 1 mM L-glutamine, 100 μM MEM nonessential amino acids and 0.1 mM β-mercaptoethanol (Life Technologies). 10 ng/ml FGF2 (R&D Systems) was added after sterile filtration. Cells were fed daily and passaged weekly using 6 U/ml dispase. For neural differentiation, cells were disassociated with Accutase and pre-plated as reported1 at the density of 200,000 cells/cm2 supplemented with 10 μM Y-27632 on Matrigel-coated plates and started differentiation the next day when confluent. KSR medium (820 ml of knockout DMEM, 150 ml knockout serum replacement, 1 mM L-glutamine, 100 μM MEM nonessential amino acids and 0.1 mM β-mercaptoethanol) was used to start differentiation. Inhibitors used in LSB+X/P/S/D induction included LDN193189 (250 nM; Stemgent), SB431542 (10 μM; Tocris), XAV939 (5 μM; Tocris), PD0325901 (1 μM in P1S5D, 8 μM in P8S10D; Tocris), SU5402 (5 μM in P1S5D, 10 μM in P8S10D; Biovision), DAPT (10 μM; Tocris). More inhibitors used in other induction described in the paper include CHIR99021 (6 μM in LSBC, 3 μM in LSB+C/S/D; Stemgent). N2 medium1 with B27 supplement (N2/B27; Life Technologies) was added in increasing 1/3 increment every other day from day 4, until reaching 100% neurobasal/B27/L-glutamine containing medium (NB/B27; Life Technologies) supplemented with BDNF (20 ng/ml; R&D), dibutyryl cAMP (0.5 mM; Sigma-Aldrich) and ascorbic acid (0.2 mM; Sigma-Aldrich) (BCA) at day 8. An outline of the P1S5D and P8S10D differentiation scheme (days 0–13 of differentiation) is presented in Figure 2a, with detailed daily feeding instructions shown in Supplementary Table 3 and Supplementary Methods. We tested 3 different lots of KSR, which gave consistent results in neuronal yield by day 13.
Rapid neuronal differentiation in Essential 6 medium (E6). The hPSC line (WA-09) was maintained in vitronectin (VTN-N; Thermo Fisher Scientific) coated culture plates in Essential 8 medium (with supplement E8). Cells were fed daily and passaged every 5 d with EDTA solution. For neural induction, cells were dissociated and pre-plated in E8 the same way as described for KSR based induction. Differentiation was started the next day when cells were confluent. Inhibitors used in LSB+X induction in E6 included LDN193189 (100 nM) and SB431542 (10 μM) for treatment of 10 d, and XAV939 (2 μM) for treatment of 3 d. E6 was used for the initial 10 d, and was switched to N2/B27 starting at day 10. For accelerated induction, cells were treated with LSB+X at concentration above in E6 from day 0 for 3 d. Then starting from day 3, LDN193189 (50 nM), SB431542 (5 μM), XAV939 (1 μM), PD0325901 (0.4 μM), SU5402 (2 μM) and DAPT (5 μM) were added into E6. N2/B27 medium was added to E6 at 1/3 (v/v) from day 5, with 1/3 increment every other day. Inhibitors in N2/B27 include LSB+X+P/S/D at the same concentration as P1S5D in KSR/N2-based induction. LSB+X were withdrawn at day 7 while P/S/D remained. 100% NB/B27+BCA was used from day 9. Inhibitors used in NB/B27 include PD0325901 (1 μM), SU5402 (5 μM) and DAPT (10 μM). An outline of the accelerated differentiation scheme in E6 (days 0–13 of differentiation) is presented in Figure 2g.
Long-term culture beyond day 13 for generation of deep- and upper-layer neurons.
The long-term culture protocol for the generation of deep- and upper-layer cortical neurons is schematically illustrated in Figure 3a, with detailed daily feeding instructions presented in Supplementary Table 3 and in Supplementary Methods. hPSCs were induced by P1S5D or P8S10D from day 0 as described in Figure 2a, and passaged on day 8 of differentiation by Accutase-mediated dissociation for 0.5–1 h at 37 °C. Cells were replated at 150,000 cells/cm2 or 300,000 cells/cm2 for P1S5D or P8S10D groups, respectively, onto pre-coated culture dishes. For pre-coating, dishes were exposed to polyornithine (PO; 15 μg/ml; Sigma-Aldrich) diluted in PBS for 24 h at 37 °C; after washing with PBS for three times, the culture dishes were further treated with mouse laminin I (1 μg/ml; R&D system) and fibronectin (2 μg/ml; Sigma-Aldrich) diluted in PBS for 12 h at 37 °C. Laminin and fibronectin were removed immediately before use. Medium used for both passaging and long-term culture was NB/B27+BCA as described above. Medium was changed every 3–4 d, and 1 μg/ml laminin was added weekly for maintaining attachment of neurons. The cells were then assessed at various in vitro time points for electrophysiological recordings, immunocytochemistry and RNA extraction. For results shown in Supplementary Figure 8, passaged P8S10D cells were co-cultured with mouse astrocytes or in the presence of astrocyte conditioned medium. The isolation and maintenance of astrocytes for those studies was carried out as described previously31. For collecting conditioned medium, astrocytes were fed with NB/B27+BCA, and conditioned medium was collected every 2–3 d. The conditioned medium was then filtered through 0.22 μm membrane pore vacuum filter (Corning) to get rid of cell contamination.
EdU labeling and quantification of cells.
EdU was added to the cultures at 5 μM for a window of 48 h each starting at various time points of differentiation (days 8, 13, 18, 23, 28 and 33), and the cells were fixed at day 40 with 4% paraformaldehyde for 20 min. EdU was detected with the Click-iT EdU Imaging Kit (Invitrogen) according to the specifications of the manufacturer. Quantification of EdU positive and cortical-layer marker-positive neurons in the EdU-labeling experiments, and the quantification of marker-positive neurons and total cells in the long-term culture was carried out using ImageJ with ITCN plugin for nuclei quantification, combined with manual counting. Six uniform randomly selected image frames from 2 independent batches of cell cultures were captured using a 20× objective and used for quantification. Areas containing clusters could not be properly resolved for colabeling analysis (EdU, cortical-layer markers) were avoided. Quantification of phospho-histone 3 and cleaved caspase 3 positive cells was also carried out using ImageJ with ITCN plugin. Per culture plate, 4 uniform randomly selected image frames were captured with 10× objective and used for quantification from 2 independent batches of cell cultures. All quantification results were plotted in Prism (version 6.0, GraphPad).
RNA extraction and qRT-PCR.
Cells were lysed with TRIzol Reagent (Life Technologies) and stored at −20 °C. Total RNA was extracted using phenol–chloroform and isopropanol precipitation, and dissolved in ddH2O. cDNA was generated using the QuantiTech Reverse Transcription Kit (Qiagen). qRT-PCR was performed using the Mastercycler Realplex2 (Eppendorf), and GAPDH was used as the housekeeping gene control for normalization. ΔΔCt and fold changes were calculated and results were plotted in Prism.
Immunocytochemistry.
Cells were fixed with 4% (v/v) paraformaldehyde for 20 min, washed with PBS, permeabilized and blocked using 0.3% (v/v) Triton X-100 in PBS with 1% (w/v) BSA for 1 h. For immunocytochemistry, cells were incubated with primary antibodies diluted in the same blocking buffer at 4 °C overnight. A list of the primary antibodies used in this study is provided as Supplementary Table 4. Following several washes, cells were incubated with appropriate Alexa Fluor secondary antibodies (1:500; Molecular Probes) and DAPI (1:1,000; Thermo Fisher) diluted in the blocking buffer for 1 h at room temperature. After washing, cells were taken images by Olympus IX71 microscope using a Hamamatsu ORCA CCD camera. For histological analysis of in vivo studies, the fixed brains were sectioned into 60-μm thick slices using a vibratome (Leica VT1200S) and stored in PBS with 0.02% NaN3 afterwards for up to 1 week. For immunocytochemistry, slices were permeabilized and blocked using 0.3% Triton X-100 in PBS with 1% BSA for 2 h and incubated with the primary antibodies diluted in the same blocking buffer for 3–5 d at 4 °C. Secondary antibody staining was performed the same as on cells. Images were acquired by either Olympus IX81 microscope with the same setting as above, or confocal laser scanning microscope (Olympus FV1000) at 2 μm with Z-series. Confocal images were taken under water immersion lenses (10× and 40×) and analyzed using FluoView (Olympus) and Photoshop (Adobe Systems).
iDISCO whole-brain immunofluorescence and imaging.
Brains were processed as described in the iDISCO protocol21, with modifications described in the updated online protocol (https://idisco.info, January 2015). The primary antibodies used were chicken anti-GFP (1:1,000; Aves GFP-1020), and mouse anti-hSynaptophysin (1:1,000; Enzo Life Science). Secondary antibodies used were donkey anti-chicken Alexa 647 (1:1,000; Jackson Immunoresearch) and donkey anti-mouse Alexa568 (1:1,000; Life Technologies). The cleared samples were imaged on a light sheet microscope (Ultramicroscope II; LaVision Biotec) equipped with a sCMOS camera (Andor Neo) and a 2×/0.5 NA objective lens equipped with a 6-mm working distance dipping cap.
Flow cytometry.
Cells were disassociated with Accutase for 0.5–1 h at 37 °C. After washing, cells were resuspended in 1× PBS with propidium iodide (2 μg/ml), and sorted by FACScalibur platform (BD Biosciences). GFP+ perecentage was determined within the propidium iodide–negative population. For intracellular flow cytometry, cells were disassociated and washed, and fixed with 4% (v/v) paraformaldehyde for 20 min. Fixed cells were then permeabilized and stained using 1× BD Perm/Wash Buffer (BD Biosciences) following the manufacturer's instructions. Primary conjugated antibodies for flow cytometry used were Nestin–Alexa 647 (1:50; 560341; BD Pharmingen) and TUJ1–Alexa 488 (1:50; 560338; BD Pharmingen). Cells were sorted using FACScalibur. Results were analyzed using FlowJo (Version 7.6).
Electrophysiology.
Cells were replated at day 8 of differentiation and maintained on 35-mm diameter Petri dishes (Falcon) in NB/B27+BCA medium supplemented with or without small molecules. On day 16, 23, 30, 37 and 40, electrophysiology was performed with preincubation in DMEM (11965; Life Technologies) at 37 °C for 2 h before recording. Only those cells with neuron-like morphologies were chosen for recording. For in vivo recording of EGFP+ grafted cells, NOD-SCID IL2Rgc−/− mice transplanted with EGFP+ H9-derived cells were anesthetized with Avertin and decapitated. The brain was removed, and 350 μm coronal brain slices were sectioned on a vibratome (Leica Microsystems) in ice-cold choline chloride–based cutting solution containing (in mM): 120 choline chloride, 26 NaHCO3, 2.6 KCl, 1.25 NaH2PO4, 7 MgSO4, 0.5 CaCl2, 1.3 ascorbate acid and 15 D-glucose, bubbled with 95% O2 and 5% CO2. Slices were transferred into artificial cerebral spinal fluid (ACSF) containing (in mM): 126 NaCl, 3 KCl, 1.2 NaH2PO4, 1.3 MgSO4, 2.4 CaCl2, 26 NaHCO3 and 10 D-glucose, bubbled with 95% O2 and 5% CO2, and recovered in an interface chamber at 32 °C for at least 1 h, and then kept at room temperature before being transferred to a recoding chamber containing ACSF at 34 °C. An infrared-DIC microscope (Olympus BX51) equipped with epifluorescence illumination, a CCD camera, and two water immersion lenses (10× and 60×) were used to visualize and target recording electrodes to EGFP+ grafted cells and H9 derived neurons in vitro. Glass recording electrodes (7–9 MΩ resistance) were filled with an intracellular solution consisting of (in mM): 126 potassium-gluconate, 2 KCl, 2 MgCl2, 0.2 EGTA, 10 HEPES, 4 Na2ATP, 0.4 Na2GTP and 0.5% neurobiotin (Invitrogen) (pH 7.25 and 295 mOsm/kg). Recordings data were collected using a Multiclamp 700B amplifier and pCLAMP10 software (Molecular Devices). The firing events were picked up, and the kinetics of firing was analyzed using Clampfit 10.2. The input resistance of a cell at the point of a small hyperpolarization current injection (–5 pA) pulse was given by Ohm′s law from the membrane potential change after it has reached plateau. Spontaneous-PSCs were analyzed using mini Analysis Program (Synaptosoft Inc.).
Transplantation into neonatal mouse.
All procedures were performed following NIH guidelines and were approved by the local Institutional Animal Care and Use Committee (IACUC), the Institutional Biosafety Committee (IBC) and the Embryonic Stem Cell Research Committee (ESCRO). P1S5D cells were disassociated with Accutase on day 8 of differentiation and filtered with a 40 μm cell strainer (Falcon). Cells were washed once and resuspended in ice-cold PBS at the density of 100,000 cells/μl and were then taken by a 10 μl syringe (Hamilton) with a 33 gauge sharp needle. A total of 2 μl cells were injected at the speed of 1 μl/min into the somatosensory cortex of P2 neonatal NOD-SCID IL2Rgc−/− mice (Jackson Laboratory) with the aid of stereotactic apparatus and electrical pump (Boston Scientific) to drive the syringe. Fully anesthetized mice were transcardially perfused with PBS containing heparin (20 units/ml) at 1 month, 1.5 months, 3 months and 6 months after grafting and followed by 20 ml of 4% paraformaldehyde. Mouse brain was then extracted and post-fixed by 4% paraformaldehyde overnight.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Lee, G. et al. Modelling pathogenesis and treatment of familial dysautonomia using patient-specific iPSCs. Nature 461, 402–406 (2009).
Mica, Y., Lee, G., Chambers, S.M., Tomishima, M.J. & Studer, L. Modeling neural crest induction, melanocyte specification, and disease-related pigmentation defects in hESCs and patient-specific iPSCs. Cell Rep. 3, 1140–1152 (2013).
Menendez, L., Yatskievych, T.A., Antin, P.B. & Dalton, S. Wnt signaling and a Smad pathway blockade direct the differentiation of human pluripotent stem cells to multipotent neural crest cells. Proc. Natl. Acad. Sci. USA 108, 19240–19245 (2011).
Maroof, A.M. et al. Directed differentiation and functional maturation of cortical interneurons from human embryonic stem cells. Cell Stem Cell 12, 559–572 (2013).
Sun, L. et al. Design, synthesis, and evaluations of substituted 3-[(3- or 4-carboxyethylpyrrol-2-yl)methylidenyl]indolin-2-ones as inhibitors of VEGF, FGF, and PDGF receptor tyrosine kinases. J. Med. Chem. 42, 5120–5130 (1999).
Dovey, H.F. et al. Functional gamma-secretase inhibitors reduce beta-amyloid peptide levels in brain. J. Neurochem. 76, 173–181 (2001).
Chambers, S.M. et al. Combined small-molecule inhibition accelerates developmental timing and converts human pluripotent stem cells into nociceptors. Nat. Biotechnol. 30, 715–720 (2012).
Espuny-Camacho, I. et al. Pyramidal neurons derived from human pluripotent stem cells integrate efficiently into mouse brain circuits in vivo. Neuron 77, 440–456 (2013).
Shi, Y., Kirwan, P., Smith, J., Robinson, H.P. & Livesey, F.J. Human cerebral cortex development from pluripotent stem cells to functional excitatory synapses. Nat. Neurosci. 15, 477–486 S1 (2012).
Barrett, S.D. et al. The discovery of the benzhydroxamate MEK inhibitors CI-1040 and PD 0325901. Bioorg. Med. Chem. Lett. 18, 6501–6504 (2008).
Huang, S.M. et al. Tankyrase inhibition stabilizes axin and antagonizes Wnt signalling. Nature 461, 614–620 (2009).
Zimmer, B. et al. Derivation of Diverse Hormone-Releasing Pituitary Cells from Human Pluripotent Stem Cells. Stem Cell Reports 6, 858–872 (2016).
Dincer, Z. et al. Specification of functional cranial placode derivatives from human pluripotent stem cells. Cell Rep. 5, 1387–1402 (2013).
Lanner, F. & Rossant, J. The role of FGF/Erk signaling in pluripotent cells. Development 137, 3351–3360 (2010).
Pucilowska, J., Puzerey, P.A., Karlo, J.C., Galán, R.F. & Landreth, G.E. Disrupted ERK signaling during cortical development leads to abnormal progenitor proliferation, neuronal and network excitability and behavior, modeling human neuro-cardio-facial-cutaneous and related syndromes. J. Neurosci. 32, 8663–8677 (2012).
Greber, B. et al. FGF signalling inhibits neural induction in human embryonic stem cells. EMBO J. 30, 4874–4884 (2011).
Greig, L.C., Woodworth, M.B., Galazo, M.J., Padmanabhan, H. & Macklis, J.D. Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 14, 755–769 (2013).
Lie, D.C. et al. Wnt signalling regulates adult hippocampal neurogenesis. Nature 437, 1370–1375 (2005).
Salinas, P.C. Wnt signaling in the vertebrate central nervous system: from axon guidance to synaptic function. Cold Spring Harb. Perspect. Biol. 4, (2012).
Bardy, C. et al. Neuronal medium that supports basic synaptic functions and activity of human neurons in vitro. Proc. Natl. Acad. Sci. USA 112, E2725–E2734 (2015).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Michelsen, K.A. et al. Area-specific reestablishment of damaged circuits in the adult cerebral cortex by cortical neurons derived from mouse embryonic stem cells. Neuron 85, 982–997 (2015).
Du, Z.W. et al. Generation and expansion of highly pure motor neuron progenitors from human pluripotent stem cells. Nat. Commun. 6, 6626 (2015).
Maury, Y. et al. Combinatorial analysis of developmental cues efficiently converts human pluripotent stem cells into multiple neuronal subtypes. Nat. Biotechnol. 33, 89–96 (2015).
Borghese, L. et al. Inhibition of notch signaling in human embryonic stem cell-derived neural stem cells delays G1/S phase transition and accelerates neuronal differentiation in vitro and in vivo. Stem Cells 28, 955–964 (2010).
Gao, P., Sultan, K.T., Zhang, X.J. & Shi, S.H. Lineage-dependent circuit assembly in the neocortex. Development 140, 2645–2655 (2013).
Kriks, S. et al. Dopamine neurons derived from human ES cells efficiently engraft in animal models of Parkinson's disease. Nature 480, 547–551 (2011).
Grealish, S. et al. Human ESC-derived dopamine neurons show similar preclinical efficacy and potency to fetal neurons when grafted in a rat model of Parkinson's disease. Cell Stem Cell 15, 653–665 (2014).
Hoya-Arias, R., Tomishima, M., Perna, F., Voza, F. & Nimer, S.D. L3MBTL1 deficiency directs the differentiation of human embryonic stem cells toward trophectoderm. Stem Cells Dev. 20, 1889–1900 (2011).
Sanjana, N.E. et al. A transcription activator-like effector toolbox for genome engineering. Nat. Protoc. 7, 171–192 (2012).
Kaech, S. & Banker, G. Culturing hippocampal neurons. Nat. Protoc. 1, 2406–2415 (2006).
Acknowledgements
We would like to thank G. Wahl (Salk Institute) and R. Jaenisch (Whitehead Institute) for sharing plasmids, F. Zhang (MIT) for sharing the TALE-toolbox, and G. Ciceri and G. Cederquist (MSKCC) for their valuable input on experimental design and feedback on the manuscript. We thank M. Sheldon (Rutgers University) for sharing protocol for fibroblast preparation. This work was supported in part through grants from the Starr Foundation (L.S. and A.H.B.) and grants NS084334 and R01NS072381(L.S.) and by NYSTEM contracts C030137 (S.S., & L.S.) and C028128 (A.H.B.) and private funds from the Rockefeller University. The Molecular Cytogenetics Core Facility at MSKCC as well as other MSKCC facilities and investigators are supported by the NIH Cancer Center support grant P30 CA008748. Some of the images were obtained using instrumentation at The Rockefeller University Bio-Imaging Resource Center. The SKI Stem Cell Research Facility is supported by NYSTEM grants C029153 and C024175 and The Starr Foundation. X.-J.Z. and B.Z. were supported by NYSTEM fellowships (C026879).
Author information
Authors and Affiliations
Contributions
Y.Q.: conception and study design, hESC manipulation, differentiation and characterization, in vitro and in vivo analyses and data interpretation and writing of manuscript. X.-J.Z.: electrophysiological recordings, in vivo transplantation, data analysis, interpretation and writing of manuscript. N.R. and Z.W.: iDISCO analysis of grafted animals, data analysis, interpretation and writing of manuscript. T.A. and Z.S.: iPSC differentiation studies, in vitro functional and electrophysiological analyses. M.Z.O. and A.H.B.: generation of the CUX2-tdTomato reporter line and writing of the manuscript. J.T. and B.Z.: generation of PAX6 and SIX1 reporter lines, data analysis. F.F. and N.Z.: neural crest differentiation protocols and data analysis. Y.G.: transplantation studies. R.A.: iDISCO analysis. M.K. and J.G.: iPSC differentiation studies, data interpretation. M.T.: iPSC induction and characterization, data analysis. M.T.-L.: design and interpretation of iDISCO studies, writing of manuscript.S.-H.S.: conception and study, data analysis and interpretation, writing of manuscript. L.S.: conception and study design, data analysis and interpretation, writing of manuscript.
Corresponding author
Ethics declarations
Competing interests
The Memorial Sloan-Kettering Cancer Center has filed a provisional patent application (US PRO 62/287821) on the methods described in the manuscript.
Integrated supplementary information
Supplementary Figure 1 Dosage-dependent response of proliferation and viability upon P/S/D treatment.
(a) Percentage of mitotic cells expressing phospho-histone 3 (pH3) among total cells at one day after P/S/D treatment (day 3 of differentiation). (b) Percentage of apoptotic cells expressing cleaved caspase 3 (CC3) among total cells at one day after P/S/D treatment (day 3 of differentiation). The conditions highlighted in the dashed line boxes in a,b are the P1 dosage groups aligned by ascending order of S concentration. For a,b, N = 4 randomly selected photo frames from each of the 2 independent batches of cell cultures. Statistical analysis was carried out using the Dunnett’s multiple comparison test to compare each dosage with LSB+X at day 3 of differentiation. Only those comparisons that are significantly different from LSB+X are marked on the graph. (c) Summary of grouped results in a following the order of increasing concentration of P, or increasing concentration of S (d). (e) Summary of grouped results in b following the order of increasing concentration of P, or increasing concentration of S (f). For c-f, statistical analysis was carried out using the Dunnett’s multiple comparison test to compare each dosage of P with the no P group (c,e) and each dosage of S with the no S dosage groups (d,f). (g) Nomenclature for the various dosage groups shown in c-f. Black dots represent values from quantification of individual photos frames. Error bars represent s. e. m. * P<0.05, ** P<0.01, *** P<0.001, **** P<0.0001.
Supplementary Figure 2 Gating of flow cytometry and impact of various small-molecule manipulations on FOXG1 expression.
(a) Gating of intracellular flow cytometry of TUJ1+ neuron population at day 13 for the various culture protocols used (blue: TUJ1 stain. red: isotype control). (b) Immunocytochemistry for FOXG1/TUJ1 co-expression at day 13. (c) Quantification of FOXG1 transcript expression level by qRT-PCR under P1S5D conditions but after systematic removal of one of the small molecules each. N = 3 independent batches of cell cultures. Scale bars: 100 μm. Error bars represent s. e. m.
Supplementary Figure 3 Characterization of additional fate markers at day 13 of differentiation.
(a) Expression of layer VI marker TLE4 in post-mitotic neurons, and CUX2 expression in progenitors at day 13 of differentiation in both P1S5D and P8S10D induction. (b) Quantification of mRNA expression at day 13 of markers other than TBR1 representing different brain areas and fates. N = 3 independent batches of cell cultures. Cortical progenitor marker OTX2, ZNF521, BRN2 and COUPTF1, layer V cortical neuron marker CTIP2, and the vesicular glutamate transporter VGLUT1 are upregulated compared to LSB+X. (c) Expression of BRN3A+ and ISL1+ cells in both P1S5D and P8S10D cultures at day 13. (d) Quantification of BRN3A+ cells at day 13. Scale bars: 50 μm. Error bars represent s. e. m.
Supplementary Figure 4 Rapid cortical neuronal induction in hiPSC lines.
(a) Validation of P1S5D and (b) P8S10D protocols on hiPSC lines by immunocytochemistry of TBR1/TUJ1 expression at day 13. (c) Quantification of neuronal induction efficiency at day 13 by intracellular flow cytometry for various hiPSC lines tested. N = 3 independent batches of cell cultures for line 1.1, 1.2, 1.3, and N = 1 for line 7.1, 7.2 and 7.4. Scale bars: 50 μm. Error bars represent s. e. m.
Supplementary Figure 5 Molecular characterization of long-term culture beyond day 13.
(a) Quantification of mRNAs expression in P1S5D and P8S10D treated culture (Fig. 3a) compared to LSB+X treated cultures. RGS4: cortical layer II-III,V marker. CHX10: retinal marker. GFAP, AQP4: astrocyte marker. OLIG2: oligodendrocyte precursor marker. For long-term culture of LSB+X cells, cells were maintained in N2 medium without re-adding small-molecule inhibitors. N = 3 independent batches of cell cultures. Error bars represent s. e. m.
Supplementary Figure 6 Generation of the CUX2-CreERT2 conditional reporter hPSC line and early generation of CUX2+ neurons in both P1S5D and P8S10D treated cells.
(a) Design of the homology donor targeting the CUX2 first exon. The selection cassette was excised upon expression of Flp recombinase after transgenesis. The location of primer sequences to confirm targeting is shown. (b) Schematic illustration of the targeted alleles. The transgenic lines express CreERT2 from the CUX2 locus and the FLEX-tdTomato conditional reporter under the CAG promoter at the AAVS1 safe harbor locus. Expression of CUX2 in the presence of 4OHT induces recombination at the reporter locus and tdTomato expression. (c) PCR confirmation of targeted transgenesis. Lanes 1-2 confirm 5’ and 3’ CreERT2 insertions at the CUX2 genomic locus, respectively, while lanes 3-4 confirm 5’ and 3’ CAG-FLEX/tdTomato insertions at the AAVS1 genomic locus, respectively. (d) Genomic DNA sequencing at the CUX2 locus confirms successful targeting of one allele and a wild-type non-targeted allele. (e) tdTomato positive post-mitotic neurons at day 70 in culture after 4OHT induction (i). Typical pyramidal morphology and lengthy projections can be observed upon higher magnification (ii). (f) CUX2+ neurons with mature morphologies observed in P1S5D and P8S10D culture at day 33 of differentiation. (g) Pyramidal morphology at day 33 suggests cortical projection neuron identify of P1S5D and P8S10D neurons. Scale bars in e (i) represents 100 μm, while those in others represent 50 μm.
Supplementary Figure 7 Summary of electrophysiological parameters for P1S5D cultures in long-term culture maintained in the absence of small molecules.
(a) Illustration of the P1S5D+none treatment analyzed in b,c for increasing levels of maturation upon further differentiation. (c) Time course quantitative analysis of electrophysiological properties of P1S5D+none conditions through day 37. Note that as time proceeded, resting membrane potential became hyperpolarized, input resistance decreased, Na+ channel current increased, action potential threshold decreased and the maximum firing frequency increased. Statistics was carried out first using ordinary one-way ANOVA to determine if statistically significant differences exist among the means of each group: F=0.3222, P=0.8093, R2=0.0248 (REM); F=7.554, P=0.0023, R2=0.5862 (half-width); F=0.7654, P=0.5209, R2=0.0560 (rising Tau); F=4.88, P=0.006, R2=0.2891 (input resistance); F=7.364, P=0.0005, R2=0.3676 (frequency). Then the Dunnett’s multiple comparison test was used to compare mean values of each group to day 16. Only those comparisons that are significant were marked on the graph. Error bars represent s. e. m. * P<0.05, ** P<0.01, *** P<0.001.
Supplementary Figure 8 Co-culture of hPSC-derived neurons with astrocyte or astrocyte conditioned medium.
(a) Schematic illustration of long-term maintenance of P8S10D+D neurons with astrocyte co-culture or conditioned media. (b) Bright field images of P8S10D+D neurons co-cultured with astrocytes or conditioned media at day 25 and day 35. (c) Representative traces of action potential firings of P8S10D+D neurons co-cultured with astrocytes or conditioned media at day 25 and day 35, evoked by current injection from -30 to +100 pA. (d) Bright field images of P8S10D+D neurons co-cultured with astrocytes at day 90. N = 15, 23, 7, 5 cells recorded for cultures with astrocytes at day 25, 35, and cultures in conditioned medium at day 25, 35. (e) Quantitative analysis of passive membrane properties and action potential properties. (f) MAP2ab staining of P8S10D+D neurons with astrocytes co-culture shows increased complexity of dendrite branching with time in culture. (g) Sholl analysis at day 36 of P8S10D+D neurons co-cultured with astrocytes, compared with neurons cultured with conditioned medium alone. Scale bars: 50 μm. Error bars represent s. e. m. *** P<0.001.
Supplementary Figure 9 iDISCO based whole brain immunofluorescence analyses of P8S10D and LSB+XAV grafts at 1 month after transplantation.
(a) P8S10D grafted half brain, stained for GFP (whole view and details of the frontal cortical region). P8S10D grafts showed inconsistent survival after transplantation into neonatal mouse cortex. However, animals with surviving graft showed long fiber projections across cortical regions. GFP+ cells devoid of axons were abundantly detected outside of the graft (boxed region). (b) LSB+X grafted half brain, stained for GFP. Whole view (side and dorsal) and detail of the graft margin (boxed region). LSB+X grafts showed massive overgrowth in host brain resulting in tumor-like structures with very limited evidence of neuronal differentiation and maturation. Only a few short fiber tracts can be seen at the margin of the graft (boxed region). N = 2 animals for P8S10D condition, and 4 animals for LSB+XAV condition analyzed. Scale bars: 500 μm.
Supplementary Figure 10 Trajectories and morphologies of P1S5D grafted neurons at 1.5 months after transplantation.
(a) Landmarks of the adult mouse brain are identified by tissue autofluorescence. The image represents 100-μm thick maximum projection of optical sections taken at the center of an adult mouse brain after iDISCO processing, and shows the major myelinated tracts from 488 nm laser excitation via endogenous fluorescence. (b) Examples of axons from grafted neurons following major pathways. iDISCO treated 1.5 months old mouse brain grafted at birth, stained for GFP. In the hippocampus, grafted neurons that are present in CA3 (arrow) and are sending axons along the fimbria tract toward the septum. In the cortex, grafted neuron project fibers across the hemisphere following callosal axons. (c) Examples of axons from grafted neurons not following major pathways. Maximum projections, 100 μm thick, of whole iDISCO treated 1.5 months old mouse brains grafted at birth, stained for GFP. In the striatum, GFP+ axons descending from the grafted neurons in the cortex are seen mainly outside of the main descending tracts. In the cortex, large bundles of GFP+ axons are observed in aberrant position. (d) A few representative morphologies seen in the grafted brains: most human neurons are clustered in the graft core, and therefore their dendritic morphologies are masked. However, a few GFP+ neurons are found outside of the graft core and exhibit diverse morphologies as shown. N = 3. Scale bars: 1 mm (a), 500 μm (b,c), 200 μm (d).
Supplementary Figure 11 in vivo electrophysiological properties of P1S5D grafted neurons.
(a, b) Recordings of action potentials and firing patterns as well as sEPSCs from a GFP+ graft at P10 (a) and P30 (b), respectively, with unusually mature properties. (c,d) Most GFP+ cells in grafts exhibited more immature firing patterns as illustrated by representative action potentials in c and sEPSCs in d recorded at P15 and P45. N = 4 animals for P1S5D condition analyzed, and 1 animal for P8S10 condition analyzed.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11, Supplementary Tables 1–4, and Supplementary Methods (PDF 2765 kb)
41587_2017_BFnbt3777_MOESM30_ESM.mp4
iDISCO-based whole-brain analysis of graft core and projections at 1 and 1.5 months post transplantation. (MP4 83155 kb)
Rights and permissions
About this article
Cite this article
Qi, Y., Zhang, XJ., Renier, N. et al. Combined small-molecule inhibition accelerates the derivation of functional cortical neurons from human pluripotent stem cells. Nat Biotechnol 35, 154–163 (2017). https://doi.org/10.1038/nbt.3777
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3777
This article is cited by
-
Human neuronal maturation comes of age: cellular mechanisms and species differences
Nature Reviews Neuroscience (2024)
-
An epigenetic barrier sets the timing of human neuronal maturation
Nature (2024)
-
Induced pluripotent stem cells (iPSCs): molecular mechanisms of induction and applications
Signal Transduction and Targeted Therapy (2024)
-
Affinity-matured DLL4 ligands as broad-spectrum modulators of Notch signaling
Nature Chemical Biology (2023)
-
Enrichment of human embryonic stem cell-derived V3 interneurons using an Nkx2-2 gene-specific reporter
Scientific Reports (2023)