Abstract
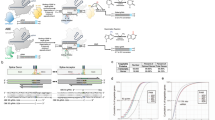
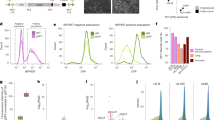
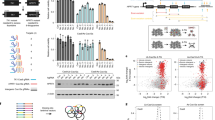
The functions of many long noncoding RNAs (lncRNAs) in the human genome remain unknown owing to the lack of scalable loss-of-function screening tools. We previously used pairs of CRISPR–Cas9 (refs. 1, 2, 3) single guide RNAs (sgRNAs) for small-scale functional screening of lncRNAs4. Here we demonstrate genome-wide screening of lncRNA function using sgRNAs to target splice sites and achieve exon skipping or intron retention. Splice-site targeting outperformed a conventional CRISPR library in a negative selection screen targeting 79 ribosomal genes. Using a genome-scale library of splicing-targeting sgRNAs, we performed a screen covering 10,996 lncRNAs and identified 230 that are essential for cellular growth of chronic myeloid leukemia K562 cells. Screening GM12878 lymphoblastoid cells and HeLa cells with the same library identified cell-type-specific differences in lncRNA essentiality. Extensive validation confirmed the robustness of our approach.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
16 January 2019
In the supplementary information originally posted, there were incorrect values for the GM12878 cell line in Supplementary Table 5. The error has been corrected online.
References
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Zhu, S. et al. Genome-scale deletion screening of human long non-coding RNAs using a paired-guide RNA CRISPR–Cas9 library. Nat. Biotechnol. 34, 1279–1286 (2016).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Wang, T., Wei, J.J., Sabatini, D.M. & Lander, E.S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Koike-Yusa, H., Li, Y., Tan, E.P., Del Castillo Velasco-Herrera, M. & Yusa, K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2014).
Zhou, Y. et al. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature 509, 487–491 (2014).
Ezkurdia, I. et al. Multiple evidence strands suggest that there may be as few as 19,000 human protein-coding genes. Hum. Mol. Genet. 23, 5866–5878 (2014).
Rinn, J.L. & Chang, H.Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 81, 145–166 (2012).
Quinn, J.J. & Chang, H.Y. Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 17, 47–62 (2016).
Kretz, M. et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493, 231–235 (2013).
Guttman, M. et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 477, 295–300 (2011).
Lin, N. et al. An evolutionarily conserved long noncoding RNA TUNA controls pluripotency and neural lineage commitment. Mol. Cell 53, 1005–1019 (2014).
Liu, S.J. et al. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells. Science 355, eaah7111 (2017).
Adamson, B., Smogorzewska, A., Sigoillot, F.D., King, R.W. & Elledge, S.J. A genome-wide homologous recombination screen identifies the RNA-binding protein RBMX as a component of the DNA-damage response. Nat. Cell Biol. 14, 318–328 (2012).
Yuan, P. et al. Chondroitin sulfate proteoglycan 4 functions as the cellular receptor for Clostridium difficile toxin B. Cell Res. 25, 157–168 (2015).
Ulitsky, I., Shkumatava, A., Jan, C.H., Sive, H. & Bartel, D.P. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147, 1537–1550 (2011).
Lim, L.P. & Burge, C.B. A computational analysis of sequence features involved in recognition of short introns. Proc. Natl. Acad. Sci. USA 98, 11193–11198 (2001).
Crooks, G.E., Hon, G., Chandonia, J.M. & Brenner, S.E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Ren, Q. et al. A Dual-reporter system for real-time monitoring and high-throughput CRISPR/Cas9 library screening of the hepatitis C virus. Sci. Rep. 5, 8865 (2015).
Peng, J., Zhou, Y., Zhu, S. & Wei, W. High-throughput screens in mammalian cells using the CRISPR-Cas9 system. FEBS J. 282, 2089–2096 (2015).
Zhu, S., Zhou, Y. & Wei, W. Genome-wide CRISPR/Cas9 screening for high-throughput functional genomics in human cells. Methods Mol. Biol. 1656, 175–181 (2017).
Matlin, A.J., Clark, F. & Smith, C.W. Understanding alternative splicing: towards a cellular code. Nat. Rev. Mol. Cell Biol. 6, 386–398 (2005).
Taggart, A.J., DeSimone, A.M., Shih, J.S., Filloux, M.E. & Fairbrother, W.G. Large-scale mapping of branchpoints in human pre-mRNA transcripts in vivo. Nat. Struct. Mol. Biol. 19, 719–721 (2012).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Xu, H. et al. Sequence determinants of improved CRISPR sgRNA design. Genome Res. 25, 1147–1157 (2015).
Heidari, N. et al. Genome-wide map of regulatory interactions in the human genome. Genome Res. 24, 1905–1917 (2014).
Muller, R.Y., Hammond, M.C., Rio, D.C. & Lee, Y.J. An Efficient method for electroporation of small interfering RNAs into ENCODE project tier 1 GM12878 and K562 cell lines. J. Biomol. Tech. 26, 142–149 (2015).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Cheng, J. et al. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science 308, 1149–1154 (2005).
Fang, Y. & Fullwood, M.J. Roles, functions, and mechanisms of long non-coding RNAs in cancer. Genomics Proteomics Bioinformatics 14, 42–54 (2016).
Joung, J. et al. Genome-scale activation screen identifies a lncRNA locus regulating a gene neighbourhood. Nature 548, 343–346 (2017).
Goyal, A. et al. Challenges of CRISPR/Cas9 applications for long non-coding RNA genes. Nucleic Acids Res. 45, e12 (2017).
Gilbert, L.A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Engreitz, J.M. et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016).
Li, B. & Dewey, C.N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Leng, N. et al. EBSeq: an empirical Bayes hierarchical model for inference in RNA-seq experiments. Bioinformatics 29, 1035–1043 (2013).
Jiao, X. et al. DAVID-WS: a stateful web service to facilitate gene/protein list analysis. Bioinformatics 28, 1805–1806 (2012).
Acknowledgements
We acknowledge the staff of the BIOPIC High-Throughput Sequencing Center (Peking University) for their assistance in next-generation sequencing analysis, the National Center for Protein Sciences Beijing (Peking University), and the core facilities at School of Life Sciences (Peking University) for help in fluorescence-activated cell sorting. We also acknowledge the High-performance Computing Platform of Peking University. This project was supported by funds from the National Science Foundation of China (NSFC31430025), the Beijing Advanced Innovation Center for Genomics at Peking University, and the Peking-Tsinghua Center for Life Sciences (W.W.).
Author information
Authors and Affiliations
Contributions
W.W. conceived and supervised the project. W.W., Y.L. and Z.C. designed the experiments. Y.L., Z.C., P.X. and Y.H. performed the experiments. Y.G. designed the oligonucleotides used for ribosomal gene mutagenesis and genome-wide lncRNA library, and Z.L. designed the pgRNAs used for individual validation. Y.W., Y.G. and P.Y. performed the bioinformatics analysis. Y.L., Z.C., Y.W. and W.W. wrote the manuscript with the help of all other authors.
Corresponding author
Ethics declarations
Competing interests
A patent has been filed relating to the data presented. W.W. is a founder and scientific advisor for EdiGene.
Supplementary information
Supplementary Figures
Supplementary Figures 1–14 (PDF 12615 kb)
Supplementary Table 1
sgRNAs of splicing-targeting library on essential ribosomal genes (XLSX 287 kb)
Supplementary Table 2
sgRNA read counts and phenotypes in ribosomal gene screening (XLSX 635 kb)
Supplementary Table 3
sgRNAs of whole-genome human lncRNA library (XLSX 6458 kb)
Supplementary Table 4
sgRNA read counts in splicing-targeting screen for lncRNAs in multiple cell lines (XLSX 18380 kb)
Supplementary Table 5
Screen scores of lncRNAs by splicing-targeting screen in multiple cell lines (XLSX 2586 kb)
Supplementary Table 6
Screen scores of lncRNAs by splicing-targeting screen in multiple cell lines (generated after filtering sgRNAs with potential off-target effects) (XLSX 2655 kb)
Supplementary Table 7
Individually cloned sgRNAs and pgRNAs for validation (XLSX 20 kb)
Supplementary Code
Source code for the computational analysis of lncRNA screens described in this paper. (ZIP 2285 kb)
Rights and permissions
About this article
Cite this article
Liu, Y., Cao, Z., Wang, Y. et al. Genome-wide screening for functional long noncoding RNAs in human cells by Cas9 targeting of splice sites. Nat Biotechnol 36, 1203–1210 (2018). https://doi.org/10.1038/nbt.4283
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4283
This article is cited by
-
Integration of transcription regulation and functional genomic data reveals lncRNA SNHG6’s role in hematopoietic differentiation and leukemia
Journal of Biomedical Science (2024)
-
Genome-scale pan-cancer interrogation of lncRNA dependencies using CasRx
Nature Methods (2024)
-
Targeting and engineering long non-coding RNAs for cancer therapy
Nature Reviews Genetics (2024)
-
Involvement of lncRNAs in cancer cells migration, invasion and metastasis: cytoskeleton and ECM crosstalk
Journal of Experimental & Clinical Cancer Research (2023)
-
Evidence for widespread existence of functional novel and non-canonical human transcripts
BMC Biology (2023)