Abstract
Fluorescence resonance energy transfer (FRET)1 detection in fusion constructs consisting of green fluorescent protein (GFP)2 variants linked by a sequence that changes conformation upon modification by enzymes or binding of ligands has enabled detection of physiological processes such as Ca2+ ion release3, and protease4,5,6,7 and kinase activity8. Current FRET microscopy techniques are limited to the use of spectrally distinct GFPs such as blue or cyan donors in combination with green or yellow acceptors. The blue or cyan GFPs have the disadvantages of less brightness and of autofluorescence. Here a FRET imaging method is presented that circumvents the need for spectral separation of the GFPs by determination of the fluorescence lifetime of the combined donor/acceptor emission by fluorescence lifetime imaging microscopy (FLIM)9,10,11,12. This technique gives a sensitive, reproducible, and intrinsically calibrated FRET measurement that can be used with the spectrally similar and bright yellow and green fluorescent proteins (EYFP/EGFP), a pair previously unusable for FRET applications. We demonstrate the benefits of this approach in the analysis of single-cell signaling by monitoring caspase activity in individual cells during apoptosis.
Similar content being viewed by others
Main
Ligand activation of death domain-containing receptors triggers an activation cascade of cysteine proteases (caspases) that catalyze the cleavage of a number of target proteins13,14,15,16, ultimately resulting in cell death. The activation of these apoptotic proteolytic pathways can be ascertained by the detection of selective substrate cleavage. We developed a construct in which an EYFP/EGFP pair was linked by a caspase-3-sensitive cleavage site that incorporates the peptide motif DEVD (ref. 17). The functionality of this linker region was tested in vitro at enzyme concentrations known to yield complete proteolytic cleavage of the synthetic fluorogenic substrate Ac-DEVD-AMC (PharMingen, San Diego, CA) and was found to be specific for caspase-3 (Fig. 1A, C), because recombinant human caspases-6, -7, and -8 did not result in cleavage. Caspase-3 cleavage of His-EYFP-DEVD-EGFP resulted in the differential migration of the individual fluorescent products (Fig. 1A) in denaturing, nonreducing sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE). Fluorescence lifetime measurements of these gel bands by FLIM identified the upper and lower bands as EGFP and EYFP, by virtue of the difference in their fluorescence lifetimes (Fig. 2C)18. The close proximity of these fluorescent proteins in the construct gives rise to FRET that is disrupted upon caspase cleavage. As illustrated in Figure 1C, the fluorescence spectral changes upon cleavage are too subtle to be useful for intensity-based ratiometric imaging. However, excellent resolution for measuring FRET is provided by the fluorescence decays of the combined GFPs in the construct. This is because the transfer of energy causes a temporal increase in the acceptor fluorescence on the nanosecond time scale, which is referred to as acceptor in-growth (Fig. 2A). When the EGFP and EYFP fluorescence emissions are collected simultaneously, this FRET-induced in-growth is manifested by an increase in the fluorescence lifetime of the combined fluorescence. This effect can be imaged in cells using FLIM in which two separate determinations for the fluorescence lifetime, τφ and τm, are obtained at each pixel of an image, from the phase shift and de-modulation of the fluorescence emission, respectively9,10,11,12. The loss of FRET upon cleavage of the caspase construct results in a reduction of τφ and τm that is apparent in the calculated downward trajectory of their parametric plot (Fig. 2B, solid line). Under conditions of high FRET efficiency in a construct, the relationship between the lifetimes, in which τm normally exceeds τφ, is reversed (Fig. 2B, dashed line). This “inversion”, with τφ now exceeding τm, can then be used as a simple telltale diagnostic for FRET. In our caspase-3 construct, the occurrence of FRET is evident in the two-dimensional histogram of τφ and τm, in which longer lifetimes were observed before cleavage as measured on native gel and in solution (Fig. 2C, D). The lifetimes of the caspase-3 construct expressed in resting Cos7 cells were shorter compared with in vitro values. This indicates the presence of basal proteolytic activity in Cos7 cells because phase and modulation lifetimes provide an absolute reproducible measure for the relative amount of cleaved construct. Upon induction of apoptosis with Fas ligand, these values further dropped to that of fully cleaved construct (Fig. 2E). Based on these measured lifetimes, a FRET efficiency of 10–15% was calculated for the intact construct. Given the large R0 distance for 50% FRET efficiency between EGFP and EYFP (5.47 nm), higher FRET efficiencies are expected. The relatively low apparent efficiency can be caused by unfavorable orientation of the chromophore transition dipoles or by a large fraction of nonfluorescent EYFP. This demonstrates that detection of acceptor in-growth by combined donor–acceptor fluorescence lifetimes is sensitive even to relatively low FRET efficiencies. For this system we calculated an error of ∼10% in the determination of the relative concentration of cleaved construct (see Experimental Protocol). Accuracy can be further improved by engineering a construct that exhibits increased FRET efficiency.
(A) Schematic representation of His-tagged EYFP–DEVD–EGFP, indicating the linkage of EGFP to EYFP with the intervening peptide sequence encoding a glycine linker and the caspase-3 recognition motif, DEVD. The SDS–PAGE gel demonstrates in vitro digestion of His-tagged EYFP–DEVD–EGFP by caspase-3, and the designated bands correspond to the respective cleavage products, identified by their individual fluorescence lifetimes determined by “in-gel” measurements (see text). (B) Absorption spectrum of recombinant His-tagged EYFP–DEVD–EGFP (solid line) in comparison with recombinant His-tagged EGFP (dotted and dashed line), or EYFP (dotted line). (C) In vitro digestion of recombinant His-tagged EYFP–DEVD–EGFP with caspase-3 but not caspases-6, -7 or -8. Emission spectra of recombinant His-tagged EYFP–DEVD–EGFP in the presence or absence of 50 ng/ml caspases-3, -6, -7, or -8 at an excitation wavelength of 457 nm.
(A) The amplitude of the in-growth in the time-resolved fluorescence intensity (I, in arbitrary units, AU) of the donor/acceptor EGFP/EYFP system is dependent on FRET efficiency. The time-resolved kinetics calculated for FRET efficiencies between 10 and 70% are presented (lower to upper traces, respectively). (B) Changes in phase (τφ) and modulation (τm) fluorescence lifetimes obtained in the frequency domain are predicted following cleavage of EYFP–DEVD–EGFP with caspase-3. Parametric plot of the predicted trajectories of τφ and τm before (high values) and after caspase-3 cleavage (low values) for constructs with 10% (solid line) and 70% (dashed line) FRET efficiency. (C) Experimental demonstration of loss of FRET due to cleavage of EYFP–DEVD–EGFP in vitro. Two-dimensional histograms of τφ and τm of purified EYFP–DEVD–EGFP with (green) or without (red) cleavage by caspase-3 (16 h, 50 ng/ml) following resolution on a native polyacrylamide gel (pH 8.8). The measured τφ and τm of cleaved products resolved on a denaturing gel are also depicted for His-tagged EYFP (yellow) and EGFP (blue). (D) Cleavage of EYFP–DEVD–EGFP by caspase-3 in solution. The τφ and τm of purified His-tagged EYFP–DEVD–EGFP in droplets were imaged on coverslips, before (red) and after 16 h treatment with 50 ng/ml caspase-3 (green). (E) An apoptotic stimulus induces cleavage of EYFP–DEVD–EGFP in vivo. The τφ and τm of Cos7 cells transfected with pEYFP–DEVD–EGFP were imaged before (red, n = 5 cells) and after 4 h treatment with 50 μg/ml recombinant Fas ligand (green, n = 4 cells).
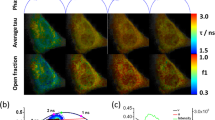
To demonstrate the ability of the method to screen on an individual cell basis in a population of cells that is heterogeneous in terms of the proteolytic events preceding apoptosis, FLIM measurements were carried out on Rat-1 fibroblasts expressing a 4-hydroxytamoxifen-dependent conditional c-myc protein19. In the absence of exogenous survival factors, the induction of c-myc expression drives cells into apoptosis. Therefore, the treatment of the Rat-1 fibroblasts with tamoxifen and Fas ligand synergistically, rapidly induces apoptosis20. Fluorescence lifetime images were acquired after expression of the caspase-3 construct in the Rat-1 fibroblasts, before (Fig. 3A) and after tamoxifen and Fas ligand treatment (Fig. 3B). In most cells a clear Fas-induced loss of FRET in the probe is observed. However, induction of apoptosis does not fully cleave the construct in every cell as visible in the cumulative two-dimensional histogram (Fig. 3C). Individual cells can now be evaluated for the onset of the asynchronous apoptotic program within a cell population, enabling the correlation with other cell state parameters on a single-cell level (Fig. 3B). The use of FLIM to detect acceptor in-growth as a measure of FRET expands the applicability of spectral variants of GFP to include pairs that cannot be separated by optical filtering in the microscope. Because the typical blue and cyan donor fluorophores also fluoresce in a spectral region containing high background autofluorescence of cells, FRET was typically measured by the FRET-induced increase in the fluorescence emission of the acceptor alone7. This intensity-based approach requires large sample statistical averaging with negative controls, or elaborate image processing to account for signals arising from direct excitation of the acceptor and bleed-through from the donor. These disadvantages are overcome in our approach that also obviates the need for strict filtering of the donor fluorescence to exclude acceptor fluorescence.
Apoptosis was induced with 50 ng/ml Fas ligand in combination with 100 nM 4-hydroxytamoxifen. Images were taken before (A) and 1 h after treatment (B). Excitation was at 457 nm and emission was detected with a 530–585 nm bandpass filter. Fluorescence intensity (I), phase lifetime (τφ), and modulation lifetime (τm) images and corresponding two-dimensional histograms are depicted for 2 out of 10 experiments for each of the untreated (A) or treated (B) situations. The cumulative two-dimensional histograms of (A) (red dots) and (B) (green dots) are shown in panel (C) for all 20 experiments.
Experimental protocol
Plasmid construction and recombinant protein purification.
The mammalian caspase-3 substrate construct comprises the fusion EYFP–GDEVDAIGGGGGGT–EGFP in the pEYFP vector (Clontech, Palo Alto, CA), referred to as EYFP–DEVD–EGFP. pRSETB was utilized to generate the histidine-tagged bacterial expression construct. pRSETB–EYFP–DEVD–EGFP was transformed into BL21(pLys)DE3 Escherichia coli, and cultures were induced with 1 mM isopropyl-β-D-thiogalactoside. Cultures were incubated for an additional 16 h at room temperature. Bacterial pellets were resuspended in distilled water, snap frozen on a bath of dry ice–methanol, and thawed at 37°C, three times. The lysate was normalized to 50 mM Tris, 300 mM NaCl (pH 10). The clarified supernatant was bound to ProBond (Invitrogen, Groningen, The Netherlands) resin equilibrated in 50 mM Tris, 300 mM NaCl (pH 10). The bound fusion protein was either maintained on the resin in 50 mM Tris, 300 mM NaCl (pH 10) or eluted with 50 mM Tris, 300 mM NaCl (pH 10) containing 350 mM imidazole.
In vitro and in vivo caspase activity assays.
For caspase digestion His-tagged EYFP–DEVD–EGFP was exchanged into caspase assay buffer (20 mM 1,4-piperazinediethanesulfonic acid, 100 mM NaCl, 1 mM ethylenediamine tetraacetic acid, 10 mM dithiothreitol, 0.1% CHAPS detergent, 10% sucrose, pH 8.0), and incubated with caspase-3, caspase-6, caspase-7, or caspase-8 (PharMingen) at concentrations as indicated in Figure 1, in the absence or presence of 200 nM caspase-3 inhibitor I (Calbiochem, La Jolla, CA), for 16 h at 37°C. Digestions were analyzed by nonreducing SDS–PAGE, native PAGE, or fluorimetry.
Fluorimetry.
Emission spectra of recombinant His-tagged EYFP–DEVD–EGFP were collected using a PTI QM1 Quantamaster T fluorimeter (Photon Technologies International, Monmouth Junction, NJ). The donor fluorophore (EGFP) was preferentially excited at an excitation wavelength of 457 nm, and the combined donor and acceptor (EYFP) emission spectra were recorded between 470 nm and 550 nm both before and after treatment with recombinant caspase-3, -6, -7, or -8.
Cell preparation.
Rat-1/c-MycER fibroblasts were cultured in Dulbecco's modified Eagle medium (DMEM) containing 10% (vol/vol) fetal calf serum (FCS) and 5 μg/ml puromycin (Sigma-Aldrich, St. Louis, MO). Apoptosis was induced with 100 nM 4-hydroxytamoxifen in combination with 50 ng/ml Fas ligand following 16 h serum starvation. Cos7 cells were cultured in DMEM containing 10% FCS.
Fluorescence microscopy.
All FLIM measurements were done at 37°C with an excitation wavelength of 457 nm and a 530–585 nm bandpass emission filter using instrumentation that had been described9. Fluorescence lifetimes were directly measured within immobilized gel slices of native polyacrylamide gels. His-tagged EYFP–DEVD–EGFP in the absence or presence of recombinant caspase-3 (50 ng/ml) was resolved on a 12% native gel at pH 8.8, and the relevant fluorescent protein bands were excised and mounted on microscope slides under coverslips using 4% agarose as sealant. The individual cleavage products were resolved on denaturing, nonreducing SDS–PAGE gels. Measurements in solution were performed on 20 μl droplets spotted on MatTek coverslip dishes (MatTek Corporation, Ashland, MA) in a humidified atmosphere.
Live cell FLIM measurements were performed on cells plated onto MatTek coverslip dishes or chamber coverslips (Labtek; Nalge Nunc International, Rochester, NY). Cells were transfected and, before microscopy, washed and immersed in HEPES-buffered, phenol red-, and riboflavin-free DMEM.
Error analysis.
The width of the distributions in the two-dimensional histograms in Figure 2C is a measure for the lifetime error (0.05 ns) because they are determined from gel bands that contain a unique species of molecule (fully cleaved or intact construct). The pixel-by-pixel error in the population of cleaved product was estimated from the lifetime error divided by the length of the lifetime trajectory from intact to cleaved product in the two-dimensional histogram (0.37 ns) and amounts to ∼14%. For an ensemble of pixels (a whole cell), this error depends on the reproducibility of the center-of-mass for each lifetime cloud in the two-dimensional histogram. For our equipment this reproducibility was 0.036 ns (n = 10, measured over 1 h) amounting to an error of ∼10%.
References
Förster, T. Zwischenmolekulare Energiewanderung und Fluoreszenz. Ann. Phys. (Leipz.) 2, 55–75 (1948).
Pollok, B.A. & Heim, R. Using GFP in FRET-based applications. Trends Cell Biol. 9, 57–60 (1999).
Miyawaki, A. et al. Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388, 882–887 (1997).
Heim, R. & Tsien, R.Y. Engineering green fluorescent protein for improved brightness, longer wavelengths and fluoresence resonance energy transfer. Curr. Biol. 6, 178–182 (1996).
Mitra, R.D., Silva, C.M. & Youvan, D.C. Fluorescence resonance energy transfer between blue-emitting and red-shifted excitation derivatives of the green fluorescent protein. Gene 173, 13–17 (1996).
Xu, X. et al. Detection of programmed cell death using fluorescence energy transfer. Nucleic Acids Res. 26, 2034–2035 (1998).
Mahajan, N.P., Harrison-Shostak, D.C., Michaux, J. & Herman, B. Novel mutant green fluorescent protein protease substrates reveal the activation of specific caspases during apoptosis. Chem. Biol. 401–409 (1999).
Nagai, Y. et al. A fluorescent indicator for visualizing cAMP-induced phosphorylation in vivo. Nat. Biotechnol. 18, 313–316 (2000).
Squire, A. & Bastiaens, P.I.H. Three dimensional image restoration in fluorescence lifetime imaging. J. Microsc. 193, 36–49 (1999).
Clegg, R.M., Feddersen, B.A., Gratton, E. & Jovin, T.M. Time resolved imaging fluorescence microscopy. Proc. SPIE 1640, 448–460 (1992).
Gadella Jr, T.W.J., Jovin, T.M. & Clegg, R.M. Fluorescence lifetime imaging microscopy (FLIM)—spatial resolution of microstructures on the nanosecond time-scale. Biophys. Chem. 48, 221–239 (1993).
Lakowicz, J.R. & Berndt, K. Lifetime-selective fluorescence imaging using an rf phase-sensitive camera. Rev. Sci. Instrum. 62, 1727–1734 (1991).
Tan, X., Martin, S.J., Green, D.R. & Wang, J.Y.J. Degradation of retinoblastoma protein in tumor necrosis factor- and CD95-induced cell death. J. Biol. Chem. 272, 9613–9616 (1997).
Martin, S.J., O'Brien, G.A., Nishioka, W.K. & McGahon, A.J. Proteolysis of fodrin (non-erythroid spectrin) during apoptosis. J. Biol. Chem. 270, 6425–6428 (1995).
Lazebnik, Y.A., Kaufmann, S.H., Desnoyers, S. & Poirier, G.G. Cleavage of poly(ADP-ribose) polymerase by a proteinase with properties like ICE. Nature 371, 346–347 (1994).
Nicholson, D.W., Ali, A., Thornberry, N.A. & Vaillancourt, J.P. Identification and inhibition of the ICE/CED-3 protease necessary for mammalian apoptosis. Nature 376, 37–43 (1995).
Casciola-Rosen, L.A., Miller, D.K., Anhalt, G.J. & Rosen, A. Specific cleavage of the 70-kDa protein component of the U1 small nuclear ribonucleoprotein is a characteristic biochemical feature of apoptotic cell death. J. Biol. Chem. 269, 30757–30760 (1994).
Pepperkok, R., Squire, A., Geley, S. & Bastiaens, P.I.H. Simultaneous detection of multiple green fluorescent proteins in live cells. Curr. Biol. 9, 269–272 (1999).
Littlewood, T.D., Hancock, D.C., Danielian, P.S., Parker, M.G. & Evan, G.I. A modified oestrogen receptor ligand-binding domain as an improved switch for the regulation of heterologous proteins. Nucleic Acids Res. 23, 1686–1690 (1995).
Hueber, A.O. et al. Requirement for the CD95 receptor–ligand pathway in c-Myc-induced apoptosis. Science 278, 1305–1309 (1997).
Acknowledgements
The authors would like to thank Gerard Evan and David Hancock of the Biochemistry of the Cell Nucleus Laboratory, Imperial Cancer Research Fund, for the Rat-1/c-mycER fibroblasts.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Harpur, A., Wouters, F. & Bastiaens, P. Imaging FRET between spectrally similar GFP molecules in single cells. Nat Biotechnol 19, 167–169 (2001). https://doi.org/10.1038/84443
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/84443