Abstract
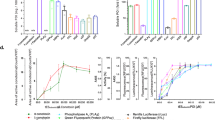
We have made an immobilized and reusable molecular chaperone system for oxidative refolding chromatography. Its three components—GroEL minichaperone (191–345), which can prevent protein aggregation; DsbA, which catalyzes the shuffling and oxidative formation of disulfide bonds; and peptidyl–prolyl isomerase—were immobilized on an agarose gel. The gel was applied to the refolding of denatured and reduced scorpion toxin Cn5. The 66–residue toxin, which has four disulfide bridges and a cis peptidyl–proline bond, had not previously been refolded in reasonable yield. We recovered an 87% yield of protein with 100% biological activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ellis, J.R. 1996. Chaperonins: introductory perspective, pp. 1–20 in The chaperonins. Buetow, D.E., Cameron, L., Padilla, G. Zimmerman, A.M. (eds.). Academic Press, New York, NY.
Bulleid, N.J. 1993. Protein disulfide–isomerase: role in biosynthesis of secretory proteins, pp. 125–148 in Accessory proteins. Lorimer, G., Anfinsen, C., Edsall, J., Richards, F., Eisenberg, D. (eds.). Academic Press, New York, NY.
Bardwell, J.J. 1993. The bonds that tie: catalysed disulfide bond formation. Cell 74:769–771.
Thomas, J.G., Ayling, A., and Baneyx, F. 1997. Molecular chaperones, folding catalysts, and the recovery of active recombinant proteins from E. coli. Appl. Biochem. Biotechnol. 66:197–238.
Hartl, F.U. 1996. Molecular chaperones in cellular protein folding. Nature 381:571–580.
Xu, Z., Horwich, A.L., and Sigler, P.B. 1997. The crystal structure of the asymmetric GroEL–GroES–(ADP)7 chaperonine complex. Nature 388:741–750.
Zahn, R., Buckle, A.M., Perrett, S., Johnson, C.M., Corrales, F.J., Golbik, R. et al. 1996. Chaperone activity and structure of monomeric polypeptide binding domains of GroEL. Proc. Natl. Acad. Sci. USA 93:15024–15029.
Buckle, A.M., Zahn, R., and Fersht, A.R. 1997. A structural model for GroEL–polypeptide recognition. Proc. Natl. Acad. Sci. USA. 94:3571–3575.
Golbik, R., Zahn, R., Harding, S.E., and Fersht, A.R. 1998. Thermodynamic stability and folding of GroEL minichaperones. J. Mol. Biol. 276:505–515.
Chatellier, J., Hill, F., Lund, P.A., and Fersht, A.R. 1998. In vivo activities of GroEL minichaperones. Proc. Natl. Acad. Sci. USA 95:9861–9866.
Altamirano, M.M., Golbik, R., Buckle, A.M., and Fersht, A.R. 1997. Refolding chromatography with immobilised mini–chaperones. Proc. Natl. Acad. Sci. USA 94:3576–3578 .
Freedman, R.B. 1989. Protein disulfide isomerase: multiple roles in the modification of nascent secretory proteins. Cell 57:1069–1072.
Bardwell, J.C.A., McGovern, K., and Beckwith, J. 1991. Identification of a protein required for disulfide bond formation in vivo. Cell 67:581–589.
Missiakas, D., Georgopoulos, C., and Raina, S. 1994. The Escherichia coli dsbc (xprA) gene encodes a periplasmic protein involved in disulfide bond formation. EMBO J. 13:2013–2020.
Lyles, M.M. and Gilbert., H.F. 1991. Catalysis of the oxidative folding of ribonuclease A by protein disulfide isomerase: dependence rate on the composition of the redox buffer. Biochemistry 30:613–619.
Bader, M., Muse, W., Zander, T., and Bardwell, J.C.A. 1998. Reconstitution of a protein disulfide catalytic system. J. Biol. Chem. 273:10302–10307.
Freedman, R.B. 1992. Protein folding in the cell, pp. 455–539 in Protein folding. Creighton, T.E. (ed.). Freeman, New York, NY.
Schmid, F.X., Mayr, L.M., Mücke, M., and Schönbrunner, R. 1993. Prolyl isomerase: role in protein folding, pp. 25–65 in Accessory proteins. Anfinsen, C., Edsall, J., Richards, F., Eisenberg, D. (eds.). Academic Press, New York, NY.
Wunderlich, M., Otto, A., Seckler, R., and Glockshuber, R. 1993. Bacterial protein disulfide isomerase: efficient catalysis of oxidative protein folding at acid pH. Biochemistry 32:12251–12256.
Martin, J.L., Bardwell, J.C.A., and Kuriyan, J. 1993. Crystal structure of the DsbA protein required for disulphide bond formation in vivo. Nature 365:464–468.
Nelson, J.W. and Creighton, T.E. 1994. Reactivity and ionization of the active site cysteine residues of DsbA, a protein required for disulfide bond formation in vivo. Biochemistry 33:5974–5983.
Zapun, A., Bardwell, J.C.A., and Creighton, T.E. 1993. The reactive and destabilizing disulfide bond of DsbA, a protein required for protein disulfide bond formation in vivo. Biochemistry 32:5083–5092.
Darby, N. and Creighton, T.E. 1995. Catalytic mechanism of DsbA and its comparison with that of protein disulfide isomerase. Biochemistry 34:3576–3587.
Wunderlich, M. and Glockshuber, R. 1993. Redox properties of protein disulfide isomerase (DsbA) from Escherichia coli. Protein Sci. 2:717–726.
Akiyama, Y., Kamitani, S., Kusukawa, N., and Ito, K. 1992. In vitro catalysis of oxidative folding of disulfide–bonded proteins by the Escherichia coli dsbA gene product. J. Biol. Chem. 267:22440–22445.
Joly, J.C. and Swartz, J.R. 1997. In vitro and in vivo redox states of the Escherichia coli periplasmic oxidoreductases DsbA and DsbC. Biochemistry 36:10067–10072.
Zapun, A., Cooper, L., and Creighton, T.E. 1994. Replacement of the active–site residues of DsbA, a protein required for disulfide bond formation in vivo. Biochemistry 33:1907–1914.
Matouschek, A., Rospert, S., Schmid, K., Glick, B.S., and Schatz, G. 1995. PPI catalyzes protein folding in yeast mitochondria. Proc. Natl. Acad. Sci. USA 92:6319–6323.
Freedman, R.B. 1991. Protein disulfide isomerase, pp. 204–214 in Conformation and forces in protein folding. Nall. B.T and Dill, K.A. (eds.). American Association for the Advancement of Science, Washington, DC.
Rudolph, R. and Lilie, H. 1996. In vitro folding of inclusion body proteins. FASEB J. 10:49–56.
De Bernandez Clark, E. 1998. Refolding of recombinant proteins. Curr. Opin. Biotechnol. 9:157–163.
Creighton, T.E. 1985. Folding of protein adsorbed reversibly to ion–exchange resins, pp. 249–258 in UCLA symposia on molecular and cellular biology new series, Vol. 39. Oxender, D.L. (ed.). New York, NY.
Stempfer, G., Holl–Neugebauer, B., and Rudolph, R. 1996. Improved refolding of an immobilized fusion protein. Nat. Biotechnol. 14:329–334.
Garcia, C., Becerril, B., Selisko, B., Delepierre, M., and Possani, L.D. 1997. Isolation, characterization and comparison of a novel crustacean toxin with a mammalian toxin from the venom of the scorpion Centruroides noxius Hoffmann. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 116:315–322 .
Zilberberg, N.G.D., Pelhate, M., Adams, M., Norris, T., Zlotkin, E., and Gurevitz, M. 1996. Functional expression and genetic alteration of an alpha scorpion neurotoxin. Biochemistry 35:10215–10222.
Florence, T.M., 1980. Degradation of protein disulphide bonds in dilute alkali. Biochem J. 189:507–520.
Hennecke, J., Sillen, A., Huber–Wunderlich, M., Engelborghs, Y., and Glockshuber, R. 1997. Quenching of tryptophan fluorescence by the active–site disulfide bridge in the DsbA protein from Escherichia coli. Biochemistry 36:6391–6400.
Holmgren, A. 1979. Thioredoxin catalyzes the reduction of insulin disulfides by dithiothreitol and dihydrolipoamide. J. Biol. Chem. 254:9627–9632.
Loret, E.P., Martin–Euclaire, M., and Mansuelle, P. 1991. An anti–insect toxin purified from the scorpion Androctonus australis Hector also acts on the α– and β–sites of the mammalian sodium channel: sequence and circular dichroism study. Biochemistry 30:633–640.
Morjana, N.A. and Gilbert., H.F., 1994. Catalysis of protein folding by agarose–immobilized protein disulfide isomerase. Protein Expr. Purif. 5:144–148.
Muronetz, V.I., Zhang, N.X., Bulatnikov, I., and Wang, C.C. 1998. Study on the interactions between protein disulfide isomerase and target proteins, using immobilization on solid support. FEBS Lett. 426:107–110.
Buchner, J. and Rudolph, R. 1991. Renaturation, purification and characterization of recombinant Fab fragments produced in Escherichia coli. Bio/Technology 9:157–162.
Buchner, J., Brinkmann, U., and Pastan, I. 1992. Renaturation of a single–chain immunotoxin facilitated by chaperones and protein disulfide isomerase. Bio/Technology 10:682–685.
Sabatier, J.M., Darbon, H., Fourquet, P., Rochat, H., and Rietschoten, J. 1987. Reduction and reoxidation of the neurotoxin II from the scorpion Androctonus australis Hector. Int. J. Pept. Protein Res 30:125–134.
Jasanoff, A., Davis, B., and Fersht A.R. 1994. Detection of an intermediate in the folding of the (ba) 8–barrel N–(5´–phosphoribosyl)anthranilate isomerase from Escherichia coli. Biochemistry 33:6350–6355.
Miroux, B. and Walker., J.E. 1996. Over–production of proteins in Escherichia coli: mutant hosts that allow synthesis of some membrane proteins and globular proteins at high levels. J. Mol. Biol. 260:289–298.
Altamirano, M.M., Lara–Lemus, G., Libreros, C., and Calcagno, M. 1989. Evidence for vicinal dithiols and their functional role in glucosamine–6–phosphate deaminase from Escherichia coli. Arch. Biochem. Biophys. 269:555–561.
Altamirano M.M., Plumbridge, J.A., and Calcagno, M.L. 1992. Identification of two cysteine residues forming a pair of vicinal thiols in glucosamine–6–phosphate deaminase from Escherichia coli and a study of their functional role by site–directed mutagenesis. Biochemistry 31:1153–1158.
Acknowledgements
M.M.A. was an EMBO fellow. This research was partially supported by a Howard Hughes Medical Institute grant (to L.D.P.). Fruitful discussions with Gilles Travé are also acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Altamirano, M., García, C., Possani, L. et al. Oxidative refolding chromatography: folding of the scorpion toxin Cn5. Nat Biotechnol 17, 187–191 (1999). https://doi.org/10.1038/6192
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/6192
This article is cited by
-
Towards a generic prototyping approach for therapeutically-relevant peptides and proteins in a cell-free translation system
Nature Communications (2022)
-
The versatile mutational “repertoire” of Escherichia coli GroEL, a multidomain chaperonin nanomachine
Biophysical Reviews (2018)
-
Protein recovery from inclusion bodies of Escherichia coli using mild solubilization process
Microbial Cell Factories (2015)
-
In vitro refolding and functional analysis of polyhistidine-tagged Buthus martensii Karsch antitumor-analgesic peptide produced in Escherichia coli
Biotechnology Letters (2015)
-
Oxidative refolding of lysozyme assisted by DsbA, DsbC and the GroEL apical domain immobilized in cellulose
Biotechnology and Bioprocess Engineering (2012)