Abstract
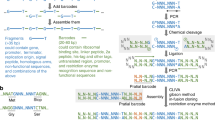
Here we describe two methods for generating DNA fragments with single-stranded overhangs, like those generated by the activity of many restriction enzymes, by simple methods that do not involve DNA digestion. The methods, RNA-overhang cloning (ROC) and DNA-overhang cloning (DOC), generate polymerase chain reaction (PCR)1 products composed of double-stranded (ds) DNA flanked by single-stranded (ss) RNA or DNA overhangs. The overhangs can be used to recombine DNA fragments at any sequence location, creating “perfect” chimeric genes composed of DNA fragments that have been joined without the insertion, deletion, or alteration of even a single base pair. The ROC method entails using PCR primers that contain regions of RNA sequence that cannot be copied by certain thermostable DNA polymerases. Using such a chimeric primer in PCR would yield a product with a 5′ overhang identical to the sequence of the RNA component of the primer, which can be used for directional ligation of the amplified product to other preselected DNA molecules. This method provides complete control over both the length and sequence of the overhangs, and eliminates the need for restriction enzymes as tools for gene engineering.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Saiki, R.K. et al. Enzymatic amplification of β-Globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230, 1350–1354 (1985).
Perler, F.B., Kumar, S. & Kong, H. Thermostable DNA polymerases. Adv. Protein Chem. 48, 377–435 (1996).
Myers, T.W. & Gelfand, D.H. Reverse transcription and DNA amplification by a Thermus thermophilus DNA polymerase. Biochemistry 30, 7661–7666 (1991).
Cline, J., Braman, J. C. & Hogrefe, H.H. PCR fidelity of Pfu DNA polymerase and other thermostable DNA polymerases. Nucleic Acids Res. 24, 3546–3551 (1996).
Krainer, A.R., Maniatis, T., Ruskin, B. & Green, M.R. Normal and mutant human β-globin pre-mRNAs are faithfully and efficiently spliced in vitro. Cell 36, 993–1005 (1984).
Sommer, B. et al. Flip and flop: a cell-specific functional switch in glutamate-operated channels of the CNS. Science 249, 1580–1585 (1990).
Mead, D.A., Kristy Pey, N., Herrnstadt, C., Marcil, R.A. & Smith, L.M. A universal method for the direct cloning of PCR amplified nucleic acid. Biotechnology (NY) 9, 657–663 (1991).
Acknowledgements
We thank Brenda Jarrell, Thomas Schaal, David Welch, and Tom Maniatis for editing of the manuscript, and Birgitte Møbes for technical assistance. This work was supported by NSF grant MCB9604458.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Coljee, V., Murray, H., Donahue, W. et al. Seamless gene engineering using RNA- and DNA-overhang cloning. Nat Biotechnol 18, 789–791 (2000). https://doi.org/10.1038/77363
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/77363
This article is cited by
-
Rapid and Efficient Gene Splicing Using Megaprimer-Based Protocol
Molecular Biotechnology (2008)
-
Creation of DNA overhangs by using modified DNA overhang cloning method
Applied Microbiology and Biotechnology (2007)
-
Description of a PCR-based technique for DNA splicing and mutagenesis by producing 5' overhangs with run through stop DNA synthesis utilizing Ara-C
BMC Biotechnology (2005)