Abstract
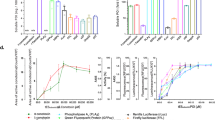
Structural genomics has the ambitious goal of delivering three-dimensional structural information on a genome-wide scale. Yet only a small fraction of natural proteins are suitable for structure determination because of bottlenecks such as poor expression, aggregation, and misfolding of proteins, and difficulties in solubilization and crystallization. We propose to overcome these bottlenecks by producing soluble, highly expressed proteins that are derived from and closely related to their natural homologs. Here we demonstrate the utility of this approach by using a green fluorescent protein (GFP) folding reporter assay to evolve an enzymatically active, soluble variant of a hyperthermophilic protein that is normally insoluble when expressed in Escherichia coli, and determining its structure by X-ray crystallography. Analysis of the structure provides insight into the substrate specificity of the enzyme and the improved solubility of the variant.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Christendat, D. et al. Structural proteomics: prospects for high throughput sample preparation. Prog. Biophys. Mol. Biol. 73, 339–345 (2000).
Gaasterland, T. Archaeal genomics. Curr. Opin. Microbiol. 2, 542–547 (1999).
Makrides, S.C. Strategies for achieving high-level expression of genes in Escherichia coli. Microbiol. Rev. 60, 512–538 (1996).
Armstrong, N., De Lencastre, A. & Gouaux, E. A new protein folding screen: application to the ligand binding domains of a glutamate and kainate receptor and to lysozyme and carbonic anhydrase. Protein Sci. 8, 1475–1483 (1999).
Rudolph, R. & Lilie, H. In vitro folding of inclusion body proteins. Faseb J. 10, 49–56 (1996).
Waldo, G.S., Standish, B.M., Berendzen, J. & Terwilliger, T.C. Rapid protein-folding assay using green fluorescent protein. Nat. Biotechnol. 17, 691–695 (1999).
Kim, C.A. et al. Polymerization of the SAM domain of TEL in leukemogenesis and transcriptional repression. EMBO J. 20, 4173–4182 (2001).
Skinner, M.M. & Terwilliger, T.C. Potential use of additivity of mutational effects in simplifying protein engineering. Proc. Natl. Acad. Sci. USA 93, 10753–10757 (1996).
Heinz, D.W., Baase, W.A. & Matthews, B.W. Folding and function of a T4 lysozyme containing 10 consecutive alanines illustrate the redundancy of information in an amino acid sequence. Proc. Natl. Acad. Sci. USA 89, 3751–3755 (1992).
Porath, J. Immobilized metal ion affinity chromatography. Prot. Exp. Purif. 3, 263–281 (1992).
Hagel, L., Lundstrom, H., Andersson, T. & Lindblom, H. Properties; in theory and practice; of novel gel-filtration media for standard liquid-chromatography. J. Chromato. 476, 329–344 (1989).
Janin, J. et al. Three-dimensional structure of nucleoside diphosphate kinase. J. Bioenerg. Biomemb. 32, 215–225 (2000).
Stemmer, W.P.C. Rapid evolution of a protein in vitro by DNA shuffling. Nature 370, 389–391 (1994).
De la Rosa, A., Williams, R.L. & Steeg, P.S. Nm23/nucleoside diphosphate kinase: toward a structural and biochemical understanding of its biological functions. Bioessays 17, 53–62 (1995).
Lascu, I. & Gonin, P. The catalytic mechanism of nucleoside diphosphate kinases. J. Bioenerg. Biomemb. 32, 237–246 (2000).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 233, 123–138 (1993).
Morera, S., Lacombe, M.L., Xu, Y.W., Lebras, G. & Janin, J. X-ray structure of human nucleoside diphosphate kinase B complexed with GDP at 2 Å resolution. Structure 3, 1307–1314 (1995).
Karamohamed, S., Nordstrom, T. & Nyren, P. Real-time bioluminometric method for detection of nucleoside diphosphate kinase activity. Biotechniques 26, 728 (1999).
Cherfils, J., Morera, S., Lascu, I., Veron, M. & Janin, J. X-ray structure of nucleoside diphosphate kinase complexed with thymidine diphosphate and Mg2+ at 2 Å resolution. Biochemistry 33, 9062–9069 (1994).
Dale, G.E., Broger, C., Langen, H., Darcy, A. & Stuber, D. Improving protein solubility through rationally designed amino acid replacements: solubilization of the trimethoprim-resistant type S1 dihydrofolate reductase. Protein Eng. 7, 933–939 (1994).
Williams, R.L. et al. Crystal structure of Myxococcus xanthus nucleoside diphosphate kinase and its interaction with a nucleotide substrate at 2 Å resolution. J. Mol. Biol. 234, 1230–1247 (1993).
Lascu, I., Giartosio, A., Ransac, S. & Erent, M. Quaternary structure of nucleoside diphosphate kinases. J. Bioenerg. Biomemb. 32, 227–236 (2000).
Lacombe, M.L., Milon, L., Munier, A., Mehus, J.G. & Lambeth, D.O. The human Nm23/nucleoside diphosphate kinases. J. Bioenerg. Biomemb. 32, 247–258 (2000).
Yang, J.K. et al. Crystallization and preliminary X-ray crystallographic analysis of the Rv2002 gene product from Mycobacterium tuberculosis, a β-ketoacyl carrier protein reductase homologue. Acta Crystallogr. D. Biol. Crystallogr. 58, 303–305 (2002).
Trivedi, V.D., Raman, B., Ramakrishna, T. & Rao, C.M. Detection and assay of proteases using calf lens β-crystallin aggregate as substrate. J. Biochem. Biophys. Meth. 40, 49–55 (1999).
Van Duyne, G.D., Standaert, R.F., Karplus, P.A., Schreiber, S.L. & Clardy, J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229, 105–124 (1993).
Collaborative Computational Project, N. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D. Biol. Crystallogr. 50, 760–763 (1994).
Terwilliger, T.C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D. Biol. Crystallogr. 55, 849–861 (1999).
Terwilliger, T.C. Reciprocal-space solvent flattening. Acta Crystallogr. D. Biol. Crystallogr. 55, 1863–1871 (1999).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D. Biol. Crystallogr. 54, 905–921 (1998).
Kraulis, P.J. Molscript: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Bacon, D.J. Photorealistic molecular graphics. Meth. Enzym. 277, 505–524 (1997).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281–296 (1991).
Acknowledgements
We thank Leon Flaks for his assistance in performing data collection. We also thank James Jett, Andrew Bradbury, and Kathleen Sandman for review of the manuscript, and the National Institutes of Health and University of California Campus Laboratory Collaboration Program for generous support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors have filed pending US patent application 09/410,889, “Method for determining and improving peptide/protein solubility.”
Rights and permissions
About this article
Cite this article
Pédelacq, JD., Piltch, E., Liong, E. et al. Engineering soluble proteins for structural genomics. Nat Biotechnol 20, 927–932 (2002). https://doi.org/10.1038/nbt732
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt732
This article is cited by
-
Computational analysis of the amino acid interactions that promote or decrease protein solubility
Scientific Reports (2018)
-
Periscope: quantitative prediction of soluble protein expression in the periplasm of Escherichia coli
Scientific Reports (2016)
-
The promises and challenges of fusion constructs in protein biochemistry and enzymology
Applied Microbiology and Biotechnology (2016)
-
Prediction and analysis of protein solubility using a novel scoring card method with dipeptide composition
BMC Bioinformatics (2012)
-
Large-scale experimental studies show unexpected amino acid effects on protein expression and solubility in vivo in E. coli
Microbial Informatics and Experimentation (2011)