Abstract
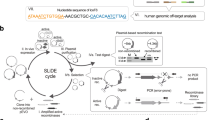
There is great interest in engineering human growth factors as potential therapeutic agonists and antagonists. We approached this goal with a synthetic DNA recombination method. We aligned a pool of “top-strand” oligonucleotides incorporating polymorphisms from mammalian genes encoding epidermal growth factor (EGF) using multiple polymorphic “scaffold” oligonucleotides. Top strands were then linked by gap filling and ligation. This approach avoided heteroduplex annealing in the linkage of highly degenerate oligonucleotides and thus achieved completely random recombination. Cloned genes from a human–mouse chimeric library captured every possible permutation of the parental polymorphisms, creating an apparently complete recombined gene-family library, which has not been previously described. This library yielded a chimeric protein whose agonist activity was enhanced 123-fold. A second library from five mammalian EGF homologs possessed the highest reported recombination density (1 crossover per 12.4 bp). The five-homolog library yielded the strongest-binding hEGF variant yet reported. In addition, it contained strongly binding EGF variants with antagonist properties. Our less biased approach to DNA shuffling should be useful for the engineering of a wide variety of proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mendelsohn, J. The epidermal growth factor receptor as a target for cancer therapy. Endocr. Relat. Cancer 8, 3–9 (2001).
Reddy, C.C., Niyogi, S.K., Wells, A., Wiley, H.S. & Lauffenburger, D.A. Engineering epidermal growth factor for enhanced mitogenic potency. Nat. Biotechnol. 14, 1696–1699 (1996).
Matsunami, R.K., Campion, S.R., Niyogi, S.K. & Stevens, A. Analogs of human epidermal growth factor which partially inhibit the growth factor-dependent protein-tyrosine kinase activity of the epidermal growth factor receptor. FEBS Lett. 264, 105–108 (1990).
Mullenbach, G.T. et al. Modification of a receptor-binding surface of epidermal growth factor (EGF): analogs with enhanced receptor affinity at low pH or at neutrality. Protein Eng. 11, 473–480 (1998).
Souriau, C., Gracy, J., Chiche, L. & Weill, M. Direct selection of EGF mutants displayed on filamentous phage using cells overexpressing EGF receptor. Biol. Chem. 380, 451–458 (1999).
Souriau, C. et al. A simple luciferase assay for signal transduction activity detection of epidermal growth factor displayed on phage. Nucleic Acids Res. 25, 1585–1590 (1997).
van de Poll, M.L., van Vugt, M.J., Lenferink, A.E. & van Zoelen, E.J. Identification of the minimal requirements for binding to the human epidermal growth factor (EGF) receptor using chimeras of human EGF and an EGF repeat of Drosophila Notch. J. Biol. Chem. 273, 16075–16081 (1998).
Van Zoelen, E.J., Stortelers, C., Lenferink, A.E. & Van de Poll, M.L. The EGF domain: requirements for binding to receptors of the ErbB family. Vitam. Horm. 59, 99–131 (2000).
Groenen, L.C., Nice, E.C. & Burgess, A.W. Structure–function relationships for the EGF/TGF-α family of mitogens. Growth Factors 11, 235–257 (1994).
Pompon, D. & Nicolas, A. Protein engineering by cDNA recombination in yeasts: shuffling of mammalian cytochrome P-450 functions. Gene 83, 15–24 (1989).
Stemmer, W.P. Rapid evolution of a protein in vitro by DNA shuffling. Nature 370, 389–391 (1994).
Crameri, A., Raillard, S.A., Bermudez, E. & Stemmer, W.P. DNA shuffling of a family of genes from diverse species accelerates directed evolution. Nature 391, 288–291 (1998).
Coco, W.M. et al. DNA shuffling method for generating highly recombined genes and evolved enzymes. Nat. Biotechnol. 19, 354–359 (2001).
Gibbs, M.D., Nevalainen, K.M. & Bergquist, P.L. Degenerate oligonucleotide gene shuffling (DOGS): a method for enhancing the frequency of recombination with family shuffling. Gene 271, 13–20 (2001).
Moore, G.L., Maranas, C.D., Lutz, S. & Benkovic, S.J. Predicting crossover generation in DNA shuffling. Proc. Natl. Acad. Sci. USA 98, 3226–3231 (2001).
Paabo, S., Irwin, D.M. & Wilson, A.C. DNA damage promotes jumping between templates during enzymatic amplification. J. Biol. Chem. 265, 4718–4721 (1990).
Levichkin, I.V., Shul'ga, A.A., Kurbanov, F.T. & Kirpichnikov, M.P. A new method of designing hybrid genes—the homolog recombination method. Mol. Biol. (Mosk) 29, 983–991 (1995).
Stemmer, W.P.C. Searching sequence space. Biotechnology 13, 549–553 (1995).
Stemmer, W.P. DNA shuffling by random fragmentation and reassembly: in vitro recombination for molecular evolution. Proc. Natl. Acad. Sci. USA 91, 10747–10751 (1994).
Ostermeier, M., Nixon, A.E., Shim, J.H. & Benkovic, S.J. Combinatorial protein engineering by incremental truncation. Proc. Natl. Acad. Sci. USA 96, 3562–3567 (1999).
Kikuchi, M., Ohnishi, K. & Harayama, S. An effective family shuffling method using single-stranded DNA. Gene 243, 133–137 (2000).
Voigt, C.A., Kauffman, S. & Wang, Z.G. Rational evolutionary design: the theory of in vitro protein evolution. Adv. Protein Chem. 55, 79–160 (2001).
Voigt, C.A., Mayo, S.L., Arnold, F.H. & Wang, Z.G. Computational method to reduce the search space for directed protein evolution. Proc. Natl. Acad. Sci. USA 98, 3778–3783 (2001).
Moore, J.C. & Arnold, F.H. Directed evolution of a para-nitrobenzyl esterase for aqueous-organic solvents. Nat. Biotechnol. 14, 458–467 (1996).
Coco, W.M. RACHITT: gene family shuffling by random chimeragenesis on transient templates. Methods Mol. Biol, in press (2003).
van de Poll, M.L., van Vugt, M.J., Lenferink, A.E. & van Zoelen, E.J. Insertion of Argos sequences into the B-loop of epidermal growth factor results in a low-affinity ligand with strong agonistic activity. Biochemistry 36, 7425–7431 (1997).
Acknowledgements
We thank Beat Blattmann, Steve Palmer, Maureen Downey, Gabe Villarreal, and Chanda Tavoloni for robotic screening assays, competition binding assays, and DNA sequencing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The work was funded by Enchira Biotechnology Corporation and was completed when the authors were employed by Enchira Biotechnology Corporation. The revised manuscript was submitted after the authors were no longer employed by the company and had no competing financial interests.
Rights and permissions
About this article
Cite this article
Coco, W., Encell, L., Levinson, W. et al. Growth factor engineering by degenerate homoduplex gene family recombination. Nat Biotechnol 20, 1246–1250 (2002). https://doi.org/10.1038/nbt757
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt757
This article is cited by
-
Optimizing Human Epidermal Growth Factor for its Endurance and Specificity Via Directed Evolution: Functional Importance of Leucine at Position 8
International Journal of Peptide Research and Therapeutics (2020)
-
Vector Design Tour de Force: Integrating Combinatorial and Rational Approaches to Derive Novel Adeno-associated Virus Variants
Molecular Therapy (2014)
-
Random mutagenesis and recombination of sam1 gene by integrating error-prone PCR with staggered extension process
Biotechnology Letters (2008)
-
Revealing biases inherent in recombination protocols
BMC Biotechnology (2007)
-
In vitro 'sexual' evolution through the PCR-based staggered extension process (StEP)
Nature Protocols (2006)