Abstract
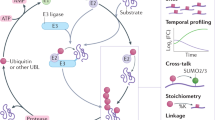
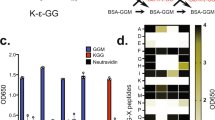
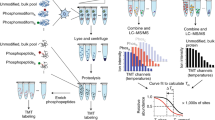
There is a growing need for techniques that can identify and characterize protein modifications on a large or global scale. We report here a proteomics approach to enrich, recover, and identify ubiquitin conjugates from Saccharomyces cerevisiae lysate. Ubiquitin conjugates from a strain expressing 6xHis-tagged ubiquitin were isolated, proteolyzed with trypsin and analyzed by multidimensional liquid chromatography coupled with tandem mass spectrometry (LC/LC-MS/MS) for amino acid sequence determination. We identified 1,075 proteins from the sample. In addition, we detected 110 precise ubiquitination sites present in 72 ubiquitin-protein conjugates. Finally, ubiquitin itself was found to be modified at seven lysine residues providing evidence for unexpected diversity in polyubiquitin chain topology in vivo. The methodology described here provides a general tool for the large-scale analysis and characterization of protein ubiquitination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Zhou, H., Watts, J.D. & Aebersold, R. A systematic approach to the analysis of protein phosphorylation. Nat. Biotechnol. 19, 375–378 (2001).
Oda, Y., Nagasu, T. & Chait, B.T. Enrichment analysis of phosphorylated proteins as a tool for probing the phosphoproteome. Nat. Biotechnol. 19, 379–382 (2001).
Ficarro, S.B. et al. Phosphoproteome analysis by mass spectrometry and its application to Saccharomyces cerevisiae. Nat. Biotechnol. 20, 301–305 (2002).
Weissman, A.M. Themes and variations on ubiquitylation. Nat. Rev. Mol. Cell. Biol. 2, 169–178 (2001).
Glickman, M.H. & Ciechanover, A. The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol. Rev. 82, 373–428 (2002).
Layfield, R., Alban, A., Mayer, R.J. & Lowe, J. The ubiquitin protein catabolic disorders. Neuropathol. Appl. Neurobiol. 27, 171–179 (2001).
Spence, J. et al. Cell cycle-regulated modification of the ribosome by a variant multiubiquitin chain. Cell 102, 67–76 (2000).
Finley, D. et al. Inhibition of proteolysis and cell cycle progression in a multiubiquitination-deficient yeast mutant. Mol. Cell Biol. 14, 5501–5509 (1994).
Gygi, M.P., Licklider, L.J., Peng, J. & Gygi, S.P. Combining two-dimensional chromatography and mass spectrometry for the separation of complex peptide mixtures. in Protein analysis: A Laboratory Manual (ed. Simpson, R.) (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2003), in the press.
Peng, J. & Gygi, S.P. Proteomics: the move to mixtures. J. Mass Spectrom. 36, 1083–1091 (2001).
Eng, J., McCormack, A.L. & Yates, J.R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Rotin, D., Staub, O. & Haguenauer-Tsapis, R. Ubiquitination and endocytosis of plasma membrane proteins: role of Nedd4/Rsp5p family of ubiquitin-protein ligases. J. Membr. Biol. 176, 1–17 (2000).
Hicke, L. Gettin' down with ubiquitin: turning off cell-surface receptors, transporters and channels. Trends Cell Biol. 9, 107–112 (1999).
Pickart, C.M. Ubiquitin in chains. Trends Biochem. Sci. 25, 544–548 (2000).
Mastrandrea, L.D., You, J., Niles, E.G. & Pickart, C.M. E2/E3-mediated assembly of lysine 29-linked polyubiquitin chains. J. Biol. Chem. 274, 27299–27306 (1999).
Spence, J., Sadis, S., Haas, A.L. & Finley, D. A ubiquitin mutant with specific defects in DNA repair and multiubiquitination. Mol. Cell Biol. 15, 1265–1273 (1995).
Baboshina, O.V. & Haas, A.L. Novel multiubiquitin chain linkages catalyzed by the conjugating enzymes E2EPF and RAD6 are recognized by 26 S proteasome subunit 5. J. Biol. Chem. 271, 2823–2831 (1996).
Finley, D. Signal transduction. An alternative to destruction. Nature 412, 283, 285–286 (2001).
Hoege, C., Pfander, B., Moldovan, G.L., Pyrowolakis, G. & Jentsch, S. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419, 135–141 (2002).
Ciechanover, A., Orian, A. & Schwartz, A.L. Ubiquitin-mediated proteolysis: biological regulation via destruction. Bioessays 22, 442–451 (2000).
Sloper-Mould, K.E., Jemc, J.C., Pickart, C.M. & Hicke, L. Distinct functional surface regions on ubiquitin. J. Biol. Chem. 276, 30483–30489 (2001).
Cook, W.J., Jeffrey, L.C., Kasperek, E. & Pickart, C.M. Structure of tetraubiquitin shows how multiubiquitin chains can be formed. J. Mol. Biol. 236, 601–609 (1994).
Goldknopf, I.L. et al. Isolation and characterization of protein A24, a “histone-like” non-histone chromosomal protein. J. Biol. Chem. 250, 7182–7187 (1975).
Gygi, S.P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Washburn, M.P., Wolters, D. & Yates, J.R. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001).
Link, A.J. et al. Direct analysis of protein complexes using mass spectrometry. Nat. Biotechnol. 17, 676–682 (1999).
Licklider, L.J., Thoreen, C.C., Peng, J. & Gygi, S.P. Automation of nanoscale microcapillary liquid chromatography–tandem mass spectrometry with a vented column. Anal. Chem. 74, 3076–3083 (2002).
Gygi, S.P., Rist, B., Griffin, T.J., Eng, J. & Aebersold, R. Proteome analysis of low abundance proteins using multidimensional chromatography and isotope coded affinity tags. J. Proteome Res. 1, 47–54 (2002).
Acknowledgements
We thank T. Yao and R. Cohen for sharing ubiquitin proteins and J. Rush and M. Comb for phosphopeptide synthesis. We also thank A. Goldberg, R. Cohen, D. Moazed, S. Gerber and L. Licklider for encouraging discussions. This work was supported in part by National Institutes of Health grants HG00041 (S.P.G.), GM67945 (S.P.G.), GM43601 (D.F.), Giovani-Armenise Harvard Foundation (S.P.G.), Jane Coffin Childs Memorial Fund for Medical Research (J.P.) and a European Molecular Biology Organization long-term fellowship (J.R.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Peng, J., Schwartz, D., Elias, J. et al. A proteomics approach to understanding protein ubiquitination. Nat Biotechnol 21, 921–926 (2003). https://doi.org/10.1038/nbt849
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt849
This article is cited by
-
Severity of Autism Spectrum Disorder Symptoms Associated with de novo Variants and Pregnancy-Induced Hypertension
Journal of Autism and Developmental Disorders (2024)
-
Targeting DCAF5 suppresses SMARCB1-mutant cancer by stabilizing SWI/SNF
Nature (2024)
-
Neutron-encoded diubiquitins to profile linkage selectivity of deubiquitinating enzymes
Nature Communications (2023)
-
Validation of catalytic site residues of Ubiquitin Specific Protease 2 (USP2) by molecular dynamic simulation and novel kinetics assay for rational drug design
Molecular Diversity (2023)
-
The emerging role of ubiquitin-specific protease 20 in tumorigenesis and cancer therapeutics
Cell Death & Disease (2022)