Abstract
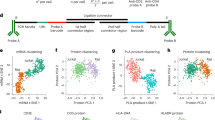
Ligand-induced clustering of type I cytokine receptor subunits leads to trans-phosphorylation and activation of associated cytosolic janus kinases (JAKs). In turn, JAKs phosphorylate tyrosine residues in the receptor tails, leading to recruitment and activation of signalling molecules1. Among these, signal transducers and activators of transcription (STATs) are important in the direct transmission of signals to the nucleus2. Here, we show that incorporation of an interaction trap in a signalling-deficient receptor allows the identification of protein–protein interactions, using a STAT-dependent complementation assay. Mammalian protein–protein interaction trap (MAPPIT) adds to existing yeast two-hybrid procedures, as originally explored by Fields and Song3, and permits the detection of both modification-independent and of phosphorylation-dependent interactions in intact human cells. We also demonstrate that MAPPIT can be used to screen complex complementary DNA libraries, and using this approach, we identify cytokine-inducible SH2-containing protein (CIS) and suppressor of cytokine signalling-2 (SOCS-2) as interaction partners of the phosphotyrosine 402 (Tyr 402)-binding motif in the erythropoietin receptor (EpoR). Importantly, this approach places protein–protein interactions in their normal physiological context, and is especially applicable to the in situ analysis of signal transduction pathways.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Darnell, J. E. Jr, Kerr, I. M. & Stark, G. R. Science 264, 1415–1421 (1994).
Bromberg, J. & Darnell, J. E. Jr. Oncogene 19, 2468–2473 (2000).
Fields, S. & Song, O. Nature 340, 245–246 (1989).
Tartaglia, L. A. et al. Cell 83, 1263–1271 (1995).
Carpenter, L. R. et al. Proc. Natl Acad. Sci. USA 95, 6061–6066 (1998).
Li, C. & Friedman, J. M. Proc. Natl Acad. Sci. USA 96, 9677–9682 (1999).
Eyckerman, S. et al. Eur. Cytokine Netw. 10, 549–556 (1999).
Eyckerman, S., Broekaert, D., Verhee, A., Vandekerckhove, J. & Tavernier, J. FEBS Lett. 486, 33–37 (2000).
Schmitz, J. et al. J. Immunol. 164, 848–854 (2000).
Schmitz, J., Weissenbach, M., Haan, S., Heinrich, P. C. & Schaper, F. J. Biol. Chem. 275, 12848–12856 (2000).
Nicholson, S. E. et al. Proc. Natl Acad. Sci. USA 97, 6493–6498 (2000).
Waelput, W. et al. Biochem. J. 348, 55–61 (2000).
Li, B. & Fields, S. FASEB J. 7, 957–963 (1993).
Starr, R. et al. Nature 387, 917–921 (1997).
Yoshimura, A. et al. EMBO J. 14, 2816–2826 (1995).
Kitamura, T. et al. Proc. Natl Acad. Sci. USA 92, 9146–9150 (1995).
Kojima, T. & Kitamura, T. Nature Biotechnol. 17, 487–490 (1999).
Verdier, F. et al. J. Biol. Chem. 273, 28185–28190 (1998).
Johnsson, N. & Varshavsky, A. Proc. Natl Acad. Sci. USA 91, 10340–10344 (1994).
Aronheim, A., Zandi, E., Hennemann, H., Elledge, S. J. & Karin, M. Mol. Cell Biol. 17, 3094–3102 (1997).
Miyawaki, A. et al. Nature 388, 882–887 (1997).
Rossi, F., Charlton, C. A. & Blau, H. M. Proc. Natl Acad. Sci. USA 94, 8405–8410 (1997).
Zozulya, S., Lioubin, M., Hill, R. J., Abram, C. & Gishizky, M. L. Nature Biotechnol. 17, 1193–1198 (1999).
Pandey, A. & Mann, M. Nature 405, 837–846 (2000).
Brent, R. & Finley, R. L. Annu. Rev. Genet. 31, 663–704 (1997).
Pattyn E. et al. J. Biol. Chem. 274, 34838–34845 (1999).
Acknowledgements
We thank G. Nolan and S. Kinoshita for the pBMN-Z retroviral vector, the mEcoR expression vector pM5Neo and the ΦNX-Eco packaging cell line, N. Nicola for Flag-tagged SOCS vectors, and S. Nordeen for pXP2d2. We are indebted to T. Van den Driessche and M. Chuah Khim for help in setting up the retroviral system, D. Lavens for the construction of the pXP2d2-rPAP1-luci reporter vector, and D. Defeau and M. Goethals for technical assistance. We thank D. Broekaert for critical reading of the manuscript and D. Iserentant and R. Dekeyser for suggestions. S.E. is a fellow with the Vlaams Instituut voor de Bevordering van het Wetenschappelijk-Technologisch Onderzoek in de Industrie (IWT). A.V. was supported by a grant from the Geconcerteerde Onderzoeksacties (GOA 12051296). This work was funded by the Fonds voor Wetenschappelijk Onderzoek Vlaanderen (FWO-V G.014999) and by the Interuniversity Attractions Poles (IUAP P4/23).
Author information
Authors and Affiliations
Supplementary information
Rights and permissions
About this article
Cite this article
Eyckerman, S., Verhee, A., der Heyden, J. et al. Design and application of a cytokine-receptor-based interaction trap. Nat Cell Biol 3, 1114–1119 (2001). https://doi.org/10.1038/ncb1201-1114
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1201-1114
This article is cited by
-
Next-generation large-scale binary protein interaction network for Drosophila melanogaster
Nature Communications (2023)
-
Bcl-xL acts as an inhibitor of IP3R channels, thereby antagonizing Ca2+-driven apoptosis
Cell Death & Differentiation (2022)
-
A reference map of the human binary protein interactome
Nature (2020)
-
Structural insights into substrate recognition by the SOCS2 E3 ubiquitin ligase
Nature Communications (2019)
-
YIPF2 is a novel Rab-GDF that enhances HCC malignant phenotypes by facilitating CD147 endocytic recycle
Cell Death & Disease (2019)