Abstract
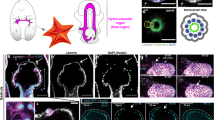
The mechanisms regulating the extension of small unicellular tubes remain poorly defined. Here we identify several steps in Caenorhabditis elegans excretory canal growth, and propose a model for lumen extension. Our results suggest that the basal and apical excretory membranes grow sequentially: the former extends first like an axon growth cone; the latter extends next as a result of an osmoregulatory activity triggering peri-apical vesicles (a membrane reservoir) to fuse with the lumen. An apical cytoskeletal web including intermediate filaments and actin crosslinking proteins ensures straight regular lumen growth. Expression of several genes encoding proteins mediating excretory lumen extension, such as the osmoregulatory STE20-like kinase GCK-3 and the intermediate filament IFB-1, is regulated by ceh-26 (here referred to as pros-1), which we found essential for excretory canal formation. Interestingly, PROS-1 is homologous to vertebrate Prox1, a transcription factor controlling lymphatic vessel growth. Our findings have potential evolutionary implications for the origin of fluid-collecting organs, and provide a reference for lymphangiogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Schmidt-Rhaesa, A. The Evolution of Organ Systems 169–190 (Oxford Univ. Press, 2007).
Schedl, A. Renal abnormalities and their developmental origin. Nat. Rev. Genet. 8, 791–802 (2007).
Tammela, T. & Alitalo, K. Lymphangiogenesis: molecular mechanisms and future promise. Cell 140, 460–476 (2010).
Andrew, D. J. & Ewald, A. J. Morphogenesis of epithelial tubes: insights into tube formation, elongation, and elaboration. Dev. Biol. 341, 34–55 (2010).
Kerman, B. E., Cheshire, A. M. & Andrew, D. J. From fate to function: the Drosophila trachea and salivary gland as models for tubulogenesis. Differentiation 74, 326–348 (2006).
Kamei, M. et al. Endothelial tubes assemble from intracellular vacuoles in vivo. Nature 442, 453–456 (2006).
Bagnat, M., Cheung, I. D., Mostov, K. E. & Stainier, D. Y. Genetic control of single lumen formation in the zebrafish gut. Nat. Cell Biol. 9, 954–960 (2007).
Su, T. et al. A kinase cascade leading to Rab11-FIP5 controls transcytosis of the polymeric immunoglobulin receptor. Nat. Cell Biol. 12, 1143–1153 (2010).
Cheshire, A. M., Kerman, B. E., Zipfel, W. R., Spector, A. A. & Andrew, D. J. Kinetic and mechanical analysis of live tube morphogenesis. Dev. Dyn. 237, 2874–2888 (2008).
Vasilyev, A. et al. Collective cell migration drives morphogenesis of the kidney nephron. PLoS Biol. 7, e9 (2009).
Buechner, M., Hall, D. H., Bhatt, H. & Hedgecock, E. M. Cystic canal mutants in Caenorhabditis elegans are defective in the apical membrane domain of the renal (excretory) cell. Dev. Biol. 214, 227–241 (1999).
Nelson, F. K., Albert, P. S. & Riddle, D. L. Fine structure of the Caenorhabditis elegans secretory-excretory system. J. Ultrastruct. Res. 82, 156–171 (1983).
Nelson, F. K. & Riddle, D. L. Functional study of the Caenorhabditis elegans secretory-excretory system using laser microsurgery. J. Exp. Zool. 231, 45–56 (1984).
Hedgecock, E. M., Culotti, J. G., Hall, D. H. & Stern, B. D. Genetics of cell and axon migrations in Caenorhabditis elegans. Development 100, 365–382 (1987).
Gao, J., Estrada, L., Cho, S., Ellis, R. E. & Gorski, J. L. The Caenorhabditis elegans homolog of FGD1, the human Cdc42 GEF gene responsible for faciogenital dysplasia, is critical for excretory cell morphogenesis. Hum. Mol. Genet. 10, 3049–3062 (2001).
Suzuki, N. et al. A putative GDP–GTP exchange factor is required for development of the excretory cell in Caenorhabditis elegans. EMBO Rep. 2, 530–535 (2001).
Van Furden, D., Johnson, K., Segbert, C. & Bossinger, O. The C. elegans ezrin-radixin-moesin protein ERM-1 is necessary for apical junction remodelling and tubulogenesis in the intestine. Dev. Biol. 272, 262–276 (2004).
Berry, K. L., Bulow, H. E., Hall, D. H. & Hobert, O. A C. elegans CLIC-like protein required for intracellular tube formation and maintenance. Science 302, 2134–2137 (2003).
Liegeois, S., Benedetto, A., Michaux, G., Belliard, G. & Labouesse, M. Genes required for osmoregulation and apical secretion in Caenorhabditis elegans. Genetics 175, 709–724 (2007).
Oka, T., Yamamoto, R. & Futai, M. Three vha genes encode proteolipids of Caenorhabditis elegans vacuolar-type ATPase. Gene structures and preferential expression in an H-shaped excretory cell and rectal cells. J. Biol. Chem. 272, 24387–24392 (1997).
Woo, W. M., Goncharov, A., Jin, Y. & Chisholm, A. D. Intermediate filaments are required for C. elegans epidermal elongation. Dev. Biol. 267, 216–229 (2004).
Solomon, A. et al. Caenorhabditis elegans OSR-1 regulates behavioral andphysiological responses to hyperosmotic environments. Genetics 167, 161–170 (2004).
Hahn-Windgassen, A. & Van Gilst, M. R. The Caenorhabditis elegans HNF4α Homolog, NHR-31, mediates excretory tube growth and function through coordinate regulation of the vacuolar ATPase. PLoS Genet. 5, e1000553 (2009).
Choe, K. P. & Strange, K. Evolutionarily conserved WNK and Ste20 kinases are essential for acute volume recovery and survival after hypertonic shrinkage in Caenorhabditis elegans. Am. J. Physiol. Cell. Physiol. 293, C915–C927 (2007).
Hisamoto, N. et al. Caenorhabditis elegans WNK-STE20 pathway regulates tube formation by modulating ClC channel activity. EMBO Rep. 9, 70–75 (2008).
Gobel, V., Barrett, P. L., Hall, D. H. & Fleming, J. T. Lumen morphogenesis in C. elegans requires the membrane-cytoskeleton linker erm-1. Dev. Cell 6, 865–873 (2004).
McKeown, C., Praitis, V. & Austin, J. sma-1 encodes a betaH-spectrin homolog required for Caenorhabditis elegans morphogenesis. Development 125, 2087–2098 (1998).
Husken, K. et al. Maintenance of the intestinal tube in Caenorhabditis elegans: the role of the intermediate filament protein IFC-2. Differentiation 76, 881–896 (2008).
Burglin, T. R. A Caenorhabditis elegans prospero homologue defines a novel domain. Trends Biochem. Sci. 19, 70–71 (1994).
Doe, C. Q., Chu-LaGraff, Q., Wright, D. M. & Scott, M. P. The prospero gene specifies cell fates in the Drosophila central nervous system. Cell 65, 451–464 (1991).
Wigle, J. T. & Oliver, G. Prox1 function is required for the development of the murine lymphatic system. Cell 98, 769–778 (1999).
Yousef, M. S. & Matthews, B. W. Structural basis of Prospero-DNA interaction: implications for transcription regulation in developing cells. Structure 13, 601–607 (2005).
Reece-Hoyes, J. S. et al. Insight into transcription factor gene duplication from Caenorhabditis elegans Promoterome-driven expression patterns. BMC Genom. 8, 27 (2007).
Mattingly, B. C. & Buechner, M. The FGD homologue EXC-5 regulates apical trafficking in C. elegans tubules. Dev. Biol. 359, 59–72 (2011).
Choe, K. P. & Strange, K. Molecular and genetic characterization of osmosensing and signal transduction in the nematode Caenorhabditis elegans. FEBS J. 274, 5782–5789 (2007).
Gobel, V. et al. Nat. Cell Biol. http://dx.doi.org/10.1038/ncb2656 (2012).
Choksi, S. P. et al. Prospero acts as a binary switch between self-renewal and differentiation in Drosophila neural stem cells. Dev. Cell 11, 775–789 (2006).
Hassan, B. et al. Prospero is a panneural transcription factor that modulates homeodomain protein activity. Proc. Natl Acad. Sci. USA 94, 10991–10996 (1997).
Sinha, B. et al. Cells respond to mechanical stress by rapid disassembly of caveolae. Cell 144, 402–413 (2011).
Armstrong, K. R. & Chamberlin, H. M. Coordinate regulation of gene expression in the C. elegans excretory cell by the POU domain protein CEH-6. Mol. Genet. Genom. 283, 73–87 (2010).
Burglin, T. R. & Ruvkun, G. Regulation of ectodermal and excretoryfunction by the C. elegans POU homeobox gene ceh-6. Development 128, 779–790 (2001).
Mah, A. K. et al. Transcriptional regulation of AQP-8, a Caenorhabditis elegans aquaporin exclusively expressed in the excretory system, by the POU homeobox transcription factor CEH-6. J. Biol. Chem. 282, 28074–28086 (2007).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Ryter, J. M., Doe, C. Q. & Matthews, B. W. Structure of the DNA binding region of prospero reveals a novel homeo-prospero domain. Structure 10, 1541–1549 (2002).
Kamath, R. S. et al. Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 421, 231–237 (2003).
Kolotuev, I., Schwab, Y. & Labouesse, M. A precise and rapid mapping protocol for correlative light and electron microscopy of small invertebrate organisms. Biol. Cell 102, 121–132 (2010).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Mastronarde, D. N. Dual-axis tomography: an approach with alignment methods that preserve resolution. J. Struct. Biol. 120, 343–352 (1997).
Strange, K., Christensen, M. & Morrison, R. Primary culture of Caenorhabditis elegans developing embryo cells for electrophysiological, cell biological and molecular studies. Nat. Protoc. 2, 1003–1012 (2007).
Acknowledgements
We thank M. Buechner (University of Kansas, USA), J. Culotti (Mt. Sinai Hospital, Toronto, Canada), I. Hope (University of Leeds, UK), V. Göbel (Massachusetts General Hospital, Boston, USA), K. Strange (Mount Desert Island Biological Laboratory, Salisbury Cove, USA), C. Eckmann (MPI-CBG, Dresden, Germany) and the CGC for strains, the Sanger Center for cosmids; P. Schultz for access to his EMPACT2 high-pressure system and Technai microscope for electron microscope tomography; L. Bianchetti, F. Klein and G. Garber for help with bioinformatics; M. Koch, P. Kessler and D. Hentsch for imaging advice; C. Spiegelhalter for assistance with electron microscopy; E. Troesch and D. Busso for directed mutagenesis; and M. Diem for technical assistance. We thank L. Broday, O. Pourquié, C. Gally and S. Quintin for critical reading of the manuscript; and L. Broday, C. Gally, I. Biryukova and G. Volohonsky for useful suggestions during the course of this work. This work was supported by grants from the Institut National du Cancer to M.L., and by a fellowship from the Fondation pour la Recherche Médicale to I.K. We wish to dedicate this work to R. Kolotueva.
Author information
Authors and Affiliations
Contributions
I.K. and M.L. designed the experiments. I.K. developed markers to monitor canal outgrowth, discovered the canal pearling response, identified the molecular nature of mc41, and characterized the role of IFB-1 in the canal. Y.S. and V.H. carried out most electron microscopy experiments with help from I.K. V.H. and D.R. examined the influence of osmotic shocks on growth and characterized PROS-1 potential targets with input from I.K. and M.L. M.L. wrote the manuscript on the basis of a draft by I.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1358 kb)
Supplementary Tables 1–3
Supplementary Information (XLS 204 kb)
Kinetics of excretory canal growth in a wild-type L1 larva.
Video of a young transgenic vha-1p::gfp L1 larva during excretory canal extension (see Fig. 1e for timing). (AVI 28 kb)
3D reconstruction of the left and right excretory canal extensions in a wild-type L1 larva.
3D models were constructed from 50 (V2) or 40 (V3) 60 nm-thick serial sections. Individual electron micrographs were aligned with Adobe Photoshop, exported as an image stack, then processed with the 3Dmod module of the Imod suite for manual segmentation of identifiable structures and 3D rendering (see also Fig. 1f–f’). (MPG 3982 kb)
3D reconstruction of the left and right excretory canal extensions in a wild-type L1 larva.
3D models were constructed from 50 (V2) or 40 (V3) 60 nm-thick serial sections. Individual electron micrographs were aligned with Adobe Photoshop, exported as an image stack, then processed with the 3Dmod module of the Imod suite for manual segmentation of identifiable structures and 3D rendering (see also Fig. 1f–f’). (MPG 2642 kb)
3D-tomography reconstruction of the excretory canal in a wild-type adult.
Electron tomograms were computed from tilt series using the etomo module of the Imod suite. The contours of the structures of interest were then manually traced and rendered with 3Dmod (see also Fig. 2d). (AVI 40113 kb)
Movement of vesicles within puffs after recovery from hypertonic conditions in a wild-type adult.
Video of a transgenic vha-1p::gfp adult recovering from a 500 mM NaCl osmotic shock on control plates with 50 mM NaCl. Images captures on a Leica-SP5 confocal microscope with 11 focal planes every 30 s (only one focal plane shown). Some vesicles are highly mobile within pearls (see also Supplementary Fig. S2b). (AVI 10756 kb)
3D reconstruction of the excretory canal in standard salt conditions.
Electron tomograms were processed as indicated in the legend to Supplementary Videos S4 (canaliculi vesicles connected to the lumen are shaded pale blue as the lumen; see also Fig. 3e–e’). (MPG 9541 kb)
3D reconstruction of the excretory canal on standard medium after recovery from high salt conditions.
(canaliculi vesicles connected to the lumen are shaded pale blue as the lumen; see also Fig. 3f–f’). (MPG 11421 kb)
Kinetics of excretory canal retraction after depletion of IFB-1 by RNAi.
Video of a double transgenic vha-5::mrfp vha-1p::gfp larva after ifb-1(RNAi) treatment during larval development; the movie shows only the green channel in false colours (see Fig. 4d for timing). (AVI 130 kb)
Rights and permissions
About this article
Cite this article
Kolotuev, I., Hyenne, V., Schwab, Y. et al. A pathway for unicellular tube extension depending on the lymphatic vessel determinant Prox1 and on osmoregulation. Nat Cell Biol 15, 157–168 (2013). https://doi.org/10.1038/ncb2662
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2662
This article is cited by
-
Pulsed stimulated Brillouin microscopy enables high-sensitivity mechanical imaging of live and fragile biological specimens
Nature Methods (2023)
-
Promiscuous Dimerization Between the Caenorhabditis elegans IF Proteins and a Hypothesis to Explain How Multiple IFs Persist Over Evolutionary Time
Journal of Molecular Evolution (2019)
-
The small GTPase ARF-1.2 is a regulator of unicellular tube formation in Caenorhabditis elegans
The Journal of Physiological Sciences (2019)
-
Overlapping cell population expression profiling and regulatory inference in C. elegans
BMC Genomics (2016)
-
CCM-3/STRIPAK promotes seamless tube extension through endocytic recycling
Nature Communications (2015)