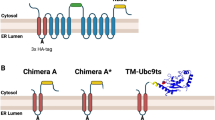
Abstract
To safeguard proteomic integrity, cells rely on the proteasome to degrade aberrant polypeptides, but it is unclear how cells remove defective proteins that have escaped degradation owing to proteasome insufficiency or dysfunction. Here we report a pathway termed misfolding-associated protein secretion, which uses the endoplasmic reticulum (ER)-associated deubiquitylase USP19 to preferentially export aberrant cytosolic proteins. Intriguingly, the catalytic domain of USP19 possesses an unprecedented chaperone activity, allowing recruitment of misfolded proteins to the ER surface for deubiquitylation. Deubiquitylated cargos are encapsulated into ER-associated late endosomes and secreted to the cell exterior. USP19-deficient cells cannot efficiently secrete unwanted proteins, and grow more slowly than wild-type cells following exposure to a proteasome inhibitor. Together, our findings delineate a protein quality control (PQC) pathway that, unlike degradation-based PQC mechanisms, promotes protein homeostasis by exporting misfolded proteins through an unconventional protein secretion process.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Wolff, S., Weissman, J. S. & Dillin, A. Differential scales of protein quality control. Cell 157, 52–64 (2014).
Bukau, B., Weissman, J. & Horwich, A. Molecular chaperones and protein quality control. Cell 125, 443–451 (2006).
Sontag, E. M., Vonk, W. I. & Frydman, J. Sorting out the trash: the spatial nature of eukaryotic protein quality control. Curr. Opin. Cell Biol. 26, 139–146 (2014).
Buchberger, A., Bukau, B. & Sommer, T. Protein quality control in the cytosol and the endoplasmic reticulum: brothers in arms. Mol. Cell 40, 238–252 (2010).
Chen, B., Retzlaff, M., Roos, T. & Frydman, J. Cellular strategies of protein quality control. Cold Spring Harb. Perspect. Biol. 3, a004374 (2011).
Clague, M. J. et al. Deubiquitylases from genes to organism. Physiol. Rev. 93, 1289–1315 (2013).
Eletr, Z. M. & Wilkinson, K. D. Regulation of proteolysis by human deubiquitinating enzymes. Biochim. Biophys. Acta 1843, 114–128 (2014).
Hassink, G. C. et al. The ER-resident ubiquitin-specific protease 19 participates in the UPR and rescues ERAD substrates. EMBO Rep. 10, 755–761 (2009).
Lee, J. G., Kim, W., Gygi, S. & Ye, Y. Characterization of the deubiquitinating activity of USP19 and its role in endoplasmic reticulum-associated degradation. J. Biol. Chem. 289, 3510–3517 (2014).
Wiles, B. et al. USP19 deubiquitinating enzyme inhibits muscle cell differentiation by suppressing unfolded-protein response signaling. Mol. Biol. Cell 26, 913–923 (2015).
Mei, Y., Hahn, A. A., Hu, S. & Yang, X. The USP19 deubiquitinase regulates the stability of c-IAP1 and c-IAP2. J. Biol. Chem. 286, 35380–35387 (2011).
Lu, Y. et al. USP19 deubiquitinating enzyme supports cell proliferation by stabilizing KPC1, a ubiquitin ligase for p27Kip1. Mol. Cell Biol. 29, 547–558 (2009).
Combaret, L. et al. USP19 is a ubiquitin-specific protease regulated in rat skeletal muscle during catabolic states. Am. J. Physiol. Endocrinol. Metab. 288, E693–E700 (2005).
Knowles, T. P., Vendruscolo, M. & Dobson, C. M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol. 15, 384–396 (2014).
Ryno, L. M., Wiseman, R. L. & Kelly, J. W. Targeting unfolded protein response signaling pathways to ameliorate protein misfolding diseases. Curr. Opin. Chem. Biol. 17, 346–352 (2013).
Hetz, C. & Mollereau, B. Disturbance of endoplasmic reticulum proteostasis in neurodegenerative diseases. Nat. Rev. Neurosci. 15, 233–249 (2014).
Brundin, P., Melki, R. & Kopito, R. Prion-like transmission of protein aggregates in neurodegenerative diseases. Nat. Rev. Mol. Cell Biol. 11, 301–307 (2010).
Baixauli, F., Lopez-Otin, C. & Mittelbrunn, M. Exosomes and autophagy: coordinated mechanisms for the maintenance of cellular fitness. Front. Immunol. 5, 403 (2014).
Mohamed, N. V., Herrou, T., Plouffe, V., Piperno, N. & Leclerc, N. Spreading of tau pathology in Alzheimer’s disease by cell-to-cell transmission. Eur. J. Neurosci. 37, 1939–1948 (2013).
Saman, S. et al. Exosome-associated tau is secreted in tauopathy models and is selectively phosphorylated in cerebrospinal fluid in early Alzheimer disease. J. Biol. Chem. 287, 3842–3849 (2012).
Nickel, W. & Rabouille, C. Mechanisms of regulated unconventional protein secretion. Nat. Rev. Mol. Cell Biol. 10, 148–155 (2009).
Lo Cicero, A., Stahl, P. D. & Raposo, G. Extracellular vesicles shuffling intercellular messages: for good or for bad. Curr. Opin. Cell Biol. 35, 69–77 (2015).
Rabouille, C., Malhotra, V. & Nickel, W. Diversity in unconventional protein secretion. J. Cell Sci. 125, 5251–5255 (2012).
Zhang, M. & Schekman, R. Cell biology. Unconventional secretion, unconventional solutions. Science 340, 559–561 (2013).
Nickel, W. Pathways of unconventional protein secretion. Curr. Opin. Biotechnol. 21, 621–626 (2010).
Malhotra, V. Unconventional protein secretion: an evolving mechanism. EMBO J. 32, 1660–1664 (2013).
Steringer, J. P., Muller, H. M. & Nickel, W. Unconventional secretion of fibroblast growth factor 2—A novel type of protein translocation across membranes? J. Mol. Biol. 427, 1202–1210 (2015).
Fujiwara, T., Oda, K., Yokota, S., Takatsuki, A. & Ikehara, Y. Brefeldin A causes disassembly of the Golgi complex and accumulation of secretory proteins in the endoplasmic reticulum. J. Biol. Chem. 263, 18545–18552 (1988).
Baneyx, F. & Mujacic, M. Recombinant protein folding and misfolding in Escherichia coli. Nat. Biotechnol. 22, 1399–1408 (2004).
Lee, J. G. & Ye, Y. Bag6/Bat3/Scythe: a novel chaperone activity with diverse regulatory functions in protein biogenesis and degradation. BioEssays 35, 377–385 (2013).
Tsunemi, T., Hamada, K. & Krainc, D. ATP13A2/PARK9 regulates secretion of exosomes and α-synuclein. J. Neurosci. 34, 15281–15287 (2014).
Hasegawa, T. et al. The AAA-ATPase VPS4 regulates extracellular secretion and lysosomal targeting of α-synuclein. PLoS ONE 6, e29460 (2011).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Nagano, Y. et al. Siah-1 facilitates ubiquitination and degradation of synphilin-1. J. Biol. Chem. 278, 51504–51514 (2003).
Friedman, J. R., Dibenedetto, J. R., West, M., Rowland, A. A. & Voeltz, G. K. Endoplasmic reticulum-endosome contact increases as endosomes traffic and mature. Mol. Biol. Cell 24, 1030–1040 (2013).
Zhang, M., Kenny, S. J., Ge, L., Xu, K. & Schekman, R. Translocation of interleukin-1β into a vesicle intermediate in autophagy-mediated secretion. eLife 4, e11205 (2015).
Kinseth, M. A. et al. The Golgi-associated protein GRASP is required for unconventional protein secretion during development. Cell 130, 524–534 (2007).
Duran, J. M., Anjard, C., Stefan, C., Loomis, W. F. & Malhotra, V. Unconventional secretion of Acb1 is mediated by autophagosomes. J. Cell Biol. 188, 527–536 (2010).
Hansen, C. et al. α-Synuclein propagates from mouse brain to grafted dopaminergic neurons and seeds aggregation in cultured human cells. J. Clin. Invest. 121, 715–725 (2011).
Oh, S. H. et al. Mesenchymal stem cells inhibit transmission of α-synuclein by modulating clathrin-mediated endocytosis in a Parkinsonian model. Cell Rep. 14, 835–849 (2016).
Desplats, P. et al. Inclusion formation and neuronal cell death through neuron-to-neuron transmission of α-synuclein. Proc. Natl Acad. Sci. USA 106, 13010–13015 (2009).
Vernace, V. A., Arnaud, L., Schmidt-Glenewinkel, T. & Figueiredo-Pereira, M. E. Aging perturbs 26S proteasome assembly in Drosophila melanogaster. FASEB J. 21, 2672–2682 (2007).
Andersson, V., Hanzen, S., Liu, B., Molin, M. & Nystrom, T. Enhancing protein disaggregation restores proteasome activity in aged cells. Aging 5, 802–812 (2013).
Bence, N. F., Sampat, R. M. & Kopito, R. R. Impairment of the ubiquitin-proteasome system by protein aggregation. Science 292, 1552–1555 (2001).
Tsai, B., Ye, Y. & Rapoport, T. A. Retro-translocation of proteins from the endoplasmic reticulum into the cytosol. Nat. Rev. Mol. Cell Biol. 3, 246–255 (2002).
Wang, Q. et al. A ubiquitin ligase-associated chaperone holdase maintains polypeptides in soluble states for proteasome degradation. Mol. Cell 42, 758–770 (2011).
Lee, J. G., Baek, K., Soetandyo, N. & Ye, Y. Reversible inactivation of deubiquitinases by reactive oxygen species in vitro and in cells. Nat. Commun. 4, 1568 (2013).
Acknowledgements
We thank X. Wu (NHLBI) and J. Reece (NIDDK) for assistance with microscopy analyses, M. Davidson (Florida State University, USA), R. Pagano (Mayo Clinic, USA), T. Galli (Institut Jacques Monod, France) and F. Zhang (Broad Institute, USA) for plasmids, and W. Prinz, H. Bernstein and M. Gellert (NIDDK) for critical reading of the manuscript. This research is supported by the Intramural Research Program of the National Institute of Diabetes, Digestive & Kidney Diseases and of the National Eye Institute in the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
S.T. and S.I.T. made the initial key observation that overexpression of USP19 induces GFP secretion. Y.Y. conceived and oversaw the study. G.Z. performed the EM analyses. J.-G.L. and Y.Y. designed and performed the experiments, analysed the data, and generated the graphs. J.-G.L. and Y.Y. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 2 USP19 preferentially promotes secretion of misfolded proteins.
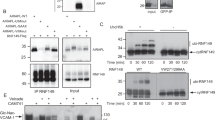
a, Secretion of GFP from HEK293T cells. HEK293T cells (0.5 × 106) were transfected with either 2 μg pcDNA3 (control) or 1 μg GFP together with 1 μg empty vector or 1 μg USP19. Conditioned mediums were harvested between 36 and 52 h post-transfection. Shown is the immunoblotting analysis of a fraction of the medium (15 μl) and different amount of lysate from cells co-transfected with GFP and USP19. The GFP protein level was used to normalize green fluorescence detected in the medium and lysate from cells co-transfected with GFP and USP19, as shown in Fig. 2a. b, GFP1-10 is more prone to aggregation in HEK293T cells. Cells (0.5 × 106) transfected with the indicated GFP variants for 48 h were lysed in a buffer (300 μl) containing the NP40 detergent. The insoluble materials (NP40 P) were re-solubilized in the Laemmli buffer (300 μl) and analysed together with the NP40 soluble fractions (NP40 S) by IB. The chromatin protein H2A is a loading control for the insoluble fractions whereas p97 for the soluble fractions. c, d, The misfolded GFP1-10 mutant is preferentially secreted from cells. c, Immunoblotting (IB) analyses of conditioned mediums and lysates from HEK293T cells transfected with the indicated constructs. Where indicated (+11), cells were co-transfected with a plasmid expressing the last β-strand of GFP. The quantification result of 4 independent experiments is shown in Fig. 2c. d, The green fluorescence intensity in cells from the experiments shown in c was measured by a FACS instrument (mean ± s.e.m., n = 4 independent experiments, ∗∗, P < 0.01 as determined by paired student t-test). e, Secretion of GFP1-10 is not sensitive to Brefeldin A (BFA). HEK293T cells transfected with GFP1-10 were treated with DMSO (control) or Brefeldin A (10 μg ml−1). Mediums collected at the indicated time points were analysed by IB. Statistics source data for d can be found in Supplementary Table 2.
Supplementary Figure 3 The functional interplay between MAPS and proteasome-mediated PQC.
a, Cycloheximide chase analysis of GFP1-10 degradation in HEK293T cells. Cells transfected with GFP1-10 for 24 h were incubated in a medium containing cycloheximide (50 μg ml−1). Where indicated, cells were also treated with MG132 (20 μM). b, The proposed functional interplay between MAPS and proteasome-mediated PQC. c, Ubl4A in the cell, but not secreted Ubl4A is ubiquitinated. FLAG-Ubl4A was immunoprecipitated under denaturing conditions from cell lysate or medium from cells co-transfected with Ubl4A-FLAG and hemagglutinin (HA)-tagged ubiquitin. Samples were analysed by IB. Asterisks indicate non-specific IgG bands. d, USP7 does not significantly reduce GFP1-10 ubiquitination. HEK293T cells transfected with the indicated constructs were treated with MG132 (20 μM) for 4 h. GFP1-10 immunoprecipitated from cell extracts was analysed by IB (top panels). A fraction of the lysate was directly analysed by IB (bottom panel). e, Overexpressed Ubl4A interacts with endogenous USP19. Control (ctrl.) or USP19 null CRISPR cells were transfected with either pcDNA3 (control) or a Ubl4A-FLAG-encoding plasmid for 24 h. Cells treated with formaldehyde were lysed in a NP40-containing lysis buffer. Cell lysates were subject to immunoprecipitation with FLAG beads. ∗, non-specific band. USP19* indicates a USP19 truncated species. f, Inhibition of the proteasome increases GFP1-10 secretion. Where indicated, cells transfected with GFP1-10 were pre-treated with MG132 (20 μM) for 1 h. g, h, Quantitative immunoblotting analysis of GFP1-10 and Ubl4A-FLAG secretion. g, Cells transfected with GFP1-10 and USP19 were incubated with fresh medium for 16 h. A fraction of the conditioned medium (15 μl) was harvested and analysed together with the indicated amount of cell lysate. The numbers indicate the amount of lysate loaded relative to the medium. h, As in g, except that cells were transfected with Ubl4A-FLAG together with USP19. i, USP19 knockout cells are not sensitive to ER stress inducers Tunicamycin (Tm) or DTT. Control and USP19 CRISPR cells treated with Tm or DTT were re-seeded in inhibitor-free medium. The plates were stained and imaged 14 days after re-seeding.
Supplementary Figure 4 USP19 uses the USP domain to recognize unfolded protein independent of Hsp90.
a, A Coomassie blue-stained gel shows that purified USP19 and USP19Δ1-493 both contain stoichiometric amount of Hsp90. b, c, The USP19 USP domain can bind unfolded luciferase independent of Hsp90. b, The USP domain purified in the presence of ADP (2 mM) or salt (500 mM NaCl) contains significantly reduced amount of Hsp90. c, The USP19 USP domain as shown in b were incubated with luciferase either on ice or at 42 °C (heat treatment) for 10 min. The soluble fractions were subject to immunoprecipitation by FLAG beads. Note that the USP19 USP domain binds denatured luciferase equally efficiently regardless of whether or not Hsp90 is present. Asterisk indicates the non-specific IgG band. d, The Hsp90 inhibitor genetespid has no effect on the chaperone activity of USP19. Luciferase was incubated with increased concentration of purified USP19 either in the presence or absence of the Hsp90 inhibitor genetespid. A fraction of the samples were directly analysed by IB (input, bottom panels). The remaining samples were heat-treated at 42 °C for 15 min. Proteins remaining in the supernatant fraction after centrifugation were analysed by IB (upper panels).
Supplementary Figure 5 USP19 recruits MAPS cargo to the ER membrane and ER-associated vesicles.
a, The permeabilized cell assay. COS7 cells expressing GFP1-10 were either fixed and stained (no digitonin treatment) or permeabilized with digitonin first before fixation and staining (digitonin treated). Digitonin treatment removed soluble cytosolic GFP1-10, and thus revert the ratio of nuclear and cytosolic GFP1-10 signals. Note that the digitonin-treated cell was scanned using a higher laser intensity. N, nucleus, Scale bar, 5 μm. b, USP19 overexpression increases membrane association of GFP1-10. COS7 cell transfected with GFP1-10 together with either a control or FLAG-USP19-expressing plasmid were permeabilized, fixed and stained with GFP and FLAG antibodies. Cells were imaged and grouped according to whether or not membrane-associated GFP1-10 was clearly visible. The graph summarizes the counting result of 59 cells without USP19 expression and 72 cells with USP19 expression. c, d, Confocal microscopy analysis of COS7 cells co-transfected with GFP1-10 and FLAG-USP19. Shown is a representative cell expressing USP19 (arrow) with more GFP1-10 on the membranes in a peri-nuclear region (circled area) than a cell with no USP19 overexpression (yellow arrowhead). N, nucleus. e–h, Confocal microscopy analysis of the localization of membrane-associated GFP1-10. Shown are representative COS7 cells transfected with GFP1-10 and FLAG-USP19, and stained with GFP (green) and PDI (red) antibodies after permeabilization. Nuclei were stained with DAPI in blue. i–k, COS7 cells transfected with mCherry-GFP1-10 (mCh-GFP1-10) together with mCitrine-USP19 (mCi-USP19) were permeabilized and imaged directly by confocal microscopy.
Supplementary Figure 6 Localization of MAPS cargos to Rab9 positive late endosomes.
a, mCh-GFP1-10 was secreted similarly as GFP1-10. Mediums and lysates from HEK293T cells expressing either GFP1-10 or mCh-GFP1-10 together with USP19 were analysed by IB. b, Live confocal microscopy analyses of cells transfected with mCh-GFP1-10 and mCi-USP19 after photobleaching show that mCh-GFP1-10-containing punctae are vesicles associated with the ER (top panels). Note the fission of the vesicle between t = 2 min and t = 3 min. The trace lines show the relative localization of the MAPS vesicles to the ER (green). The bottom panels show an example of tubulation of a GFP1-10-containing MAPS vesicle (arrow). Scale bar, 5 μm. c, The MAPS cargo GFP1-10 is not co-localized with Rab5. COS7 cells co-transfected with mCh-GFP1-10, mCi-USP19 and mCe-Rab5 were photobleached and then analysed by confocal microscopy. d, Co-localization of Ubl4A-Venus with mCh-Rab9. COS7 cells co-transfected with Ubl4A-Venus, mCe-USP19 and mCh-Rab5 were photobleached and then analysed by live confocal microscopy. Shown are two time points. The arrows show late endosomes that form tubules. e, f, A small fraction of GFP1-10-containing MAPS vesicles are stained by Lysotracker. e, COS7 cells co-transfected with mCh-CFP1-10 and mCi-USP19 were stained by a Lysotracker blue dye and then photobleached and analysed by live confocal microscopy. The arrows indicate MAPS vesicles that are not labelled by Lysotracker. f, Quantification of co-localization of MAPS vesicles with Rab9 or Lysotracker. Where indicated, cells were treated with an inhibitor cocktail that blocks lysosomal degradation for 1 h before imaging. g, Structure illuminated microscopy analysis of MAPS vesicles. Cells transfected with mCe-USP19, mCi-Rab9 and mCh-GFP1-10 were permeabilized, fixed and imaged. Shown are vesicles representing two distinct localizations observed for GFP1-10 in Rab9-bearing LEs. Scale bars, 1 μm. h, Transmission EM shows the luminal localization of GFP1-10. COS7 cells expressing FLAG-tagged GFP1-10 were permeabilized, fixed, and stained with anti-FALG antibody and immunogold-labelled secondary antibody. Yellow arrowheads show examples of luminal GFP1-10 signal. Note the tube (blue arrow) growing out of a vesicle reminiscent of that observed by live cell imaging.
Supplementary Figure 7 Autophagy is not involved in MAPS.
a, Secretion of GFP1-10 is not affected by starvation. Cells transfected with GFP1-10 together with either control or FLAG-USP19-encoding plasmid were either incubated in complete medium or starvation medium (EBSS). Mediums collected at the indicated time points were analysed by IB together with lysates prepared at the end of the chase. The graph shows the quantification results (mean ± s.e.m., n = 3 independent experiments, N.S. not significant, as analysed by paired student t-test). b, Secretion of GFP1-10 is not affected by 3-Methyladenine (3-MA). Cells transfected with FLAG-GFP1-10 and USP19 were incubated in a complete medium in the presence or absence of 3-MA (10 mM). Mediums collected at the indicated time points were analysed by IB together with lysates prepared at the end of the chase. c, Secretion of Ubl4A-FLAG is not affected by 3-MA. As in b, except that cells were transfected with Ubl4A-FLAG together with USP19. Statistics source data can be found in Supplementary Table 2.
Supplementary Figure 8 Secretion of misfolded proteins through late endosomes.
a, VAMP3 is not co-localized with Rab9. COS7 cells transfected with EGFP-VAMP3 and mCh-Rab9 were imaged by confocal microscopy. b, A model illustrates the encapsulation of MAPS cargos (green dots) by LEs. c, Association of GFP1-10 with the cell surface before secretion. COS7 cells transfected with GFP1-10 together with the indicated constructs were stained with GFP antibodies without fixing and permeabilization. Scale bar, 10 μm.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1693 kb)
Supplementary Table 2
Supplementary Information (XLSX 90 kb)
Permeabilization of COS7 cells.
A cell transfected with mCh-GFP1-10 together with mCe-USP19 were imaged directly by confocal microscopy. Cells were treated with 0.01% digitonin during the imaging process. (AVI 610 kb)
A photobleaching-based assay reveals MAPS vesicles containing mCh-GFP1-10.
A COS7 cell co-transfected with mCh-GFP1-10 and mCi-USP19 were subjected to repeated photobleaching with a 568 nm laser in the indicated area. The mCh-GFP1-10 signal was scanned right before each round of photobleaching. Note the appearance of mCh-GFP1-10-containing vesicles as the cytoplasmic mCh-GFP1-10 signal was dimed by photobleaching. (AVI 464 kb)
MAPS vesicles can undergo fission.
An example of a mCh-GFP1-10-containing vesicle undergoing fission in a COS7 cell expressing mCh-GFP1-10 (magenta) and mCi-USP19 (green) that has been subject to photobleaching. The arrow indicates the time point when the vesicle is about to divide. Note the extensive interaction between the GFP1-10 vesicles and the ER (labelled by USP19). The duration of the recording is 10 min. (AVI 8062 kb)
MAPS vesicles can form tubules
An example of mCh-GFP1-10 vesicles undergoing tubulation in COS7 cells expressing mCh-GFP1-10 (red) and mCi-USP19 (not shown) after photobleaching. Arrows indicate narrow tubes that are formed from the mCh-GFP1-10 vesicles. The duration of the recording time is 6 min. (AVI 18379 kb)
MAPS vesicles are tightly associated with the ER and moves along the ER tracks.
An example of mCh-GFP1-10 vesicles (green) that form tight association with the ER (magenta) in a COS7 cells expressing mCh-GFP1-10 and mCi-USP19 after photobleaching. Note that the yellow arrowhead labels a mCh-GFP1-10 vesicle that is almost immobile as it appears to be ‘glued’ on a mesh of ER. The white arrow indicates a mCh-GFP1-10-containing vesicle that is ‘hoping’ along an ER track. The duration of the recording is 3 min. (AVI 6619 kb)
Co-localization mCh-GFP1-10 with Rab9 at the late endosomes.
An example of mCh-GFP1-10 vesicles (magenta) that co-migrate with LEs labelled with mCi-Rab9 (green) in a COS7 expressing mCh-GFP1-10, mCi-Rab9 and mCe-USP19 (not shown) after photobleaching. Note that in some vesicles, mCh-GFP1-10 seems to be localized to the limiting membrane whereas in other, it appears to fill the lumen of LE vesicles. The duration of the recording is 1 min. (AVI 2187 kb)
Co-localization Ubl4A-Venus with Rab9.
An example showing that Ubl4A-Venus-containing vesicles (green) co-migrate with LEs labelled with mCh-Rab9 (magenta) in a COS7 cell expressing Ubl4A-Venus, mCe-USP19 (not shown) and mCh-Rab9 after photobleaching. The duration of the recording is 2 min. (AVI 9496 kb)
Rights and permissions
About this article
Cite this article
Lee, JG., Takahama, S., Zhang, G. et al. Unconventional secretion of misfolded proteins promotes adaptation to proteasome dysfunction in mammalian cells. Nat Cell Biol 18, 765–776 (2016). https://doi.org/10.1038/ncb3372
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3372
This article is cited by
-
LRP10 and α-synuclein transmission in Lewy body diseases
Cellular and Molecular Life Sciences (2024)
-
Opposing USP19 splice variants in TGF-β signaling and TGF-β-induced epithelial–mesenchymal transition of breast cancer cells
Cellular and Molecular Life Sciences (2023)
-
Deubiquitinating enzymes (DUBs): decipher underlying basis of neurodegenerative diseases
Molecular Psychiatry (2022)
-
Effects of oligomer toxicity, fibril toxicity and fibril spreading in synucleinopathies
Cellular and Molecular Life Sciences (2022)
-
G392E neuroserpin causing the dementia FENIB is secreted from cells but is not synaptotoxic
Scientific Reports (2021)