Abstract
Metabolic reprogramming is a hallmark of cancer. Herein we discover that the key glycolytic enzyme pyruvate kinase M2 isoform (PKM2), but not the related isoform PKM1, is methylated by co-activator-associated arginine methyltransferase 1 (CARM1). PKM2 methylation reversibly shifts the balance of metabolism from oxidative phosphorylation to aerobic glycolysis in breast cancer cells. Oxidative phosphorylation depends on mitochondrial calcium concentration, which becomes critical for cancer cell survival when PKM2 methylation is blocked. By interacting with and suppressing the expression of inositol-1,4,5-trisphosphate receptors (InsP3Rs), methylated PKM2 inhibits the influx of calcium from the endoplasmic reticulum to mitochondria. Inhibiting PKM2 methylation with a competitive peptide delivered by nanoparticles perturbs the metabolic energy balance in cancer cells, leading to a decrease in cell proliferation, migration and metastasis. Collectively, the CARM1–PKM2 axis serves as a metabolic reprogramming mechanism in tumorigenesis, and inhibiting PKM2 methylation generates metabolic vulnerability to InsP3R-dependent mitochondrial functions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
13 November 2017
In the version of this Article originally published, an amino acid (aa) range in Fig. 2a incorrectly read 390–53 aa. The correct range is 390–531 aa. In addition, two labels from Fig. 8j were displaced during production and instead appeared over Fig. 8g. These errors have now been corrected in the online version of the Article.
References
Cairns, R. A., Harris, I. S. & Mak, T. W. Regulation of cancer cell metabolism. Nat. Rev. Cancer 11, 85–95 (2011).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Lunt, S. Y. & Vander Heiden, M. G. Aerobic glycolysis: meeting the metabolic requirements of cell proliferation. Annu. Rev. Cell Dev. Biol. 27, 441–464 (2011).
Guo, J. Y. et al. Activated Ras requires autophagy to maintain oxidative metabolism and tumorigenesis. Genes Dev. 25, 460–470 (2011).
Weinberg, F. et al. Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc. Natl Acad. Sci. USA 107, 8788–8793 (2010).
Fogal, V. et al. Mitochondrial p32 protein is a critical regulator of tumor metabolism via maintenance of oxidative phosphorylation. Mol. Cell. Biol. 30, 1303–1318 (2010).
Weinberg, S. E. & Chandel, N. S. Targeting mitochondria metabolism for cancer therapy. Nat. Chem. Biol. 11, 9–15 (2015).
Noguchi, T., Inoue, H. & Tanaka, T. The M1- and M2-type isozymes of rat pyruvate kinase are produced from the same gene by alternative RNA splicing. J. Biol. Chem. 261, 13807–13812 (1986).
Christofk, H. R., Vander Heiden, M. G., Wu, N., Asara, J. M. & Cantley, L. C. Pyruvate kinase M2 is a phosphotyrosine-binding protein. Nature 452, 181–186 (2008).
Tamada, M., Suematsu, M. & Saya, H. Pyruvate kinase M2: multiple faces for conferring benefits on cancer cells. Clin. Cancer Res. 18, 5554–5561 (2012).
Keller, K. E., Tan, I. S. & Lee, Y. S. SAICAR stimulates pyruvate kinase isoform M2 and promotes cancer cell survival in glucose-limited conditions. Science 338, 1069–1072 (2012).
Li, Z., Yang, P. & Li, Z. The multifaceted regulation and functions of PKM2 in tumor progression. Biochim. Biophys. Acta 1846, 285–296 (2014).
Christofk, H. R. et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452, 230–233 (2008).
Xie, J., Dai, C. & Hu, X. Evidence that does not support pyruvate kinase M2 (PKM2)-catalyzed reaction as a rate-limiting step in cancer cell glycolysis. J. Biol. Chem. 291, 8987–8999 (2016).
Hosios, A. M., Fiske, B. P., Gui, D. Y. & Vander Heiden, M. G. Lack of evidence for PKM2 protein kinase activity. Mol. Cell 59, 850–857 (2015).
Yang, Y. & Bedford, M. T. Protein arginine methyltransferases and cancer. Nat. Rev. Cancer 13, 37–50 (2013).
Kawabe, Y., Wang, Y. X., McKinnell, I. W., Bedford, M. T. & Rudnicki, M. A. Carm1 regulates Pax7 transcriptional activity through MLL1/2 recruitment during asymmetric satellite stem cell divisions. Cell Stem Cell 11, 333–345 (2012).
Hein, K. et al. Site-specific methylation of Notch1 controls the amplitude and duration of the Notch1 response. Sci. Signal. 8, ra30 (2015).
Shishkova, E. et al. Global mapping of CARM1 substrates defines enzyme specificity and substrate recognition. Nat. Commun. 8, 15571 (2017).
Frietze, S., Lupien, M., Silver, P. A. & Brown, M. CARM1 regulates estrogen-stimulated breast cancer growth through up-regulation of E2F1. Cancer Res. 68, 301–306 (2008).
Cheng, H. et al. Overexpression of CARM1 in breast cancer is correlated with poorly characterized clinicopathologic parameters and molecular subtypes. Diagn. Pathol. 8, 129 (2013).
Wang, L. et al. CARM1 methylates chromatin remodeling factor BAF155 to enhance tumor progression and metastasis. Cancer Cell 25, 21–36 (2014).
Cárdenas, C. et al. Essential regulation of cell bioenergetics by constitutive InsP3 receptor Ca2+ transfer to mitochondria. Cell 142, 270–283 (2010).
Lee, J. & Bedford, M. T. PABP1 identified as an arginine methyltransferase substrate using high-density protein arrays. EMBO Rep. 3, 268–273 (2002).
Anastasiou, D. et al. Pyruvate kinase M2 activators promote tetramer formation and suppress tumorigenesis. Nat. Chem. Biol. 8, 839–847 (2012).
Lunt, S. Y. et al. Pyruvate kinase isoform expression alters nucleotide synthesis to impact cell proliferation. Mol. Cell 57, 95–107 (2015).
Patra, K. C. & Hay, N. The pentose phosphate pathway and cancer. Trends Biochem. Sci. 39, 347–354 (2014).
Rizzuto, R., De Stefani, D., Raffaello, A. & Mammucari, C. Mitochondria as sensors and regulators of calcium signalling. Nat. Rev. Mol. Cell Biol. 13, 566–578 (2012).
Patel, M. S. & Korotchkina, L. G. Regulation of the pyruvate dehydrogenase complex. Biochem. Soc. Trans. 34, 217–222 (2006).
Qattan, A. T., Radulovic, M., Crawford, M. & Godovac-Zimmermann, J. Spatial distribution of cellular function: the partitioning of proteins between mitochondria and the nucleus in MCF7 breast cancer cells. J. Proteome Res. 11, 6080–6101 (2012).
Koboldt, D. C. et al. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Mertins, P. et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 534, 55–62 (2016).
Sanchez, Y. et al. Genome-wide analysis of the human p53 transcriptional network unveils a lncRNA tumour suppressor signature. Nat. Commun. 5, 5812 (2014).
Thundimadathil, J. Cancer treatment using peptides: current therapies and future prospects. J. Amino Acids 2012, 967347 (2012).
Minn, A. J. et al. Genes that mediate breast cancer metastasis to lung. Nature 436, 518–524 (2005).
Hitosugi, T. et al. Tyrosine phosphorylation inhibits PKM2 to promote the Warburg effect and tumor growth. Sci. Signal 2, ra73 (2009).
Anastasiou, D. et al. Inhibition of pyruvate kinase M2 by reactive oxygen species contributes to cellular antioxidant responses. Science 334, 1278–1283 (2011).
Territo, P. R., Mootha, V. K., French, S. A. & Balaban, R. S. Ca2+ activation of heart mitochondrial oxidative phosphorylation: role of the F0/F1-ATPase. Am. J. Physiol. Cell Physiol. 278, C423–C435 (2000).
Foskett, J. K., White, C., Cheung, K. H. & Mak, D. O. Inositol trisphosphate receptor Ca2 + release channels. Physiol. Rev. 87, 593–658 (2007).
Cardenas, C. et al. Selective vulnerability of cancer cells by inhibition of Ca(2 +) transfer from endoplasmic reticulum to mitochondria. Cell Rep. 14, 2313–2324 (2016).
Christofk, H. R. et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452, 230–233 (2008).
Clayton, D. A. & Shadel, G. S. Isolation of mitochondria from tissue culture cells. Cold Spring Harb. Protoc. 2014, pdb.prot080002 (2014).
Kanaan, Y. M. et al. Metabolic profile of triple-negative breast cancer in African-American women reveals potential biomarkers of aggressive disease. Cancer Genom. Proteom. 11, 279–294 (2014).
Bai, R. K. & Wong, L. J. Simultaneous detection and quantification of mitochondrial DNA deletion(s), depletion, and over-replication in patients with mitochondrial disease. J. Mol. Diagn. 7, 613–622 (2005).
Prabaharan, M., Grailer, J. J., Pilla, S., Steeber, D. A. & Gong, S. Amphiphilic multi-arm-block copolymer conjugated with doxorubicin via pH-sensitive hydrazone bond for tumor-targeted drug delivery. Biomaterials 30, 5757–5766 (2009).
Chen, G., Wang, Y., Xie, R. & Gong, S. Tumor-targeted pH/redox dual-sensitive unimolecular nanoparticles for efficient siRNA delivery. J. Control. Release 259, 105–114 (2017).
Lee, Y. et al. Charge-conversion ternary polyplex with endosome disruption moiety: a technique for efficient and safe gene delivery. Angew. Chem. 120, 5241–5244 (2008).
Pittella, F. et al. Enhanced endosomal escape of siRNA-incorporating hybrid nanoparticles from calcium phosphate and PEG-block charge-conversional polymer for efficient gene knockdown with negligible cytotoxicity. Biomaterials 32, 3106–3114 (2011).
Chang, K.-L., Higuchi, Y., Kawakami, S., Yamashita, F. & Hashida, M. Efficient gene transfection by histidine-modified chitosan through enhancement of endosomal escape. Bioconjug. Chem. 21, 1087–1095 (2010).
Ziegler, A., Nervi, P., Durrenberger, M. & Seelig, J. The cationic cell-penetrating peptide CPP(TAT) derived from the HIV-1 protein TAT is rapidly transported into living fibroblasts: optical, biophysical, and metabolic evidence. Biochemistry 44, 138–148 (2005).
Vizcaino, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Haug, K. et al. MetaboLights–an open-access general-purpose repository for metabolomics studies and associated meta-data. Nucleic Acids Res. 41, D781–D786 (2013).
Acknowledgements
We thank M. G. Vander Heiden for kindly providing the immortalized MEF (PKM2fl/fl, Cre-oestrogen receptor) cells, J. Massagué for kindly providing the LM2 cells, N. Sherer for fluorescence microscopy, C. Coriano and J. Fan for comments, and P. Ahlquist for editing. This project is supported by NCI RO1 CA213293 to W.X. and R21 CA196653 to W.X. and S.G., and supported in part by the NIH/NCI P30CA014520 -UW Carbone Cancer Center Grant, and NIH R01 DK071801 to L.L., S10RR029531 and P41GM108538.
Author information
Authors and Affiliations
Contributions
W.X. and F.L. conceived the project, designed the experiments, analysed the data and wrote the manuscript. F.L. performed the experiments with assistance from H.Z., B.L., J.J. and Yidan W. F.M., L.H. and C.J. performed the mass spectrometry experiments; Yuyuan W., G.C. and S.G. designed the nanoparticles. P.L. and I.M.O. performed bioinformatics analyses; W.X., L.L. and S.G. directed and supervised all aspects of the study; all authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 CARM1 KO decreases EdU incorporation and increases OCR in MCF7 cells.
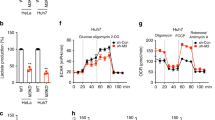
(a) EdU incorporation assays in parental MCF7 and CARM1 KO cells (n = 3 independent experiments). Cells were incubated with 10 μM EdU for 1 hour prior to flow cytometric analysis. (b,c) Basal OCR values normalized to cell numbers in parental MCF7 and CARM1 KO cells (b) or in parental MDA-MB-231 and CARM1 KO cells (c) (n = 6 independent experiments). (d,e) Relative lactate production in parental MCF7 and CARM1 KO cells (d) or in parental MDA-MB-231 and CARM1 KO cells (e) (n = 3 independent experiments). (f) Relative glucose uptake in parental MCF7 and CARM1 KO cells (n = 3 independent experiments). In a–f, data are shown as Mean ± SD and statistics source data are available in Supplementary Table 7. Statistical significance was assessed using two-tailed t-test. ∗∗p < 0.01, ns: not significant.
Supplementary Figure 2 TEPP-46 promotes PKM2 tetramer formation whereas R445/447/455K mutations on neither PKM1 nor PKM2 alter their di-/tetra-merization status.
(a) PDB structure of PKM2 tetramer (PDB ID: 3SRH) showing the positions of R445, 447 and 455 residues. Neither of the three R methylation sites is localized to the tetrameric interface. (b) Size exclusion chromatography and western blot analyses of His-tagged PKM2 in the presence and absence of TEPP-46 treatment. Wild type PKM2 and PKM2 R445/447/455K mutant peaks are completely overlapped. (c) Size exclusion chromatography and western blot analyses of His-tagged PKM1 and the corresponding R445/447/455K mutant. Mutations at R methylation sites do not alter PKM1 tetramer status. In b,c, data represent one of the two independent experiments with similar results. Unprocessed original scans of blots are shown in Supplementary Figure 9.
Supplementary Figure 3 Characterization of PKM2 KO clones.
(a) Genomic DNA sequencing results of selected PKM2 KO clones shows frame-shifts in PKM2 specific exon, resulting in knockout of PKM2 in MCF7 and MDA-MB-231 cells. (b) The relative pyruvate kinase activity in parental and MCF7 PKM2 KO clones (n = 3 independent experiments). (c) The relative pyruvate kinase activity in parental MCF7, MCF7 CARM1 KO, parental MDA-MB-231 or MAD-MB-231 CARM1 KO cells (n = 3 independent experiments). (d) Cell growth measured by MTT assays in parental MCF7 and MCF7 cells overexpressing PKM1 (n = 10 independent experiments). (e) Western blot analysis of PKM1 in MCF7 PKM2 KO or MDA-MB-231 PKM2 KO cells expressing ctrl shRNA or PKM1 shRNA (#1 and 2). (f) Cell growth measured by MTT assays in MCF7 PKM2 KO cells with ctrl shRNA or PKM1 shRNA (#1) knockdown (n = 6 independent experiments). (g) Basal OCR values normalized to cell numbers in MCF7 PKM2 KO cells with ctrl shRNA or PKM1 shRNA (#1) knockdown (n = 3 independent experiments). (h) Western blot analysis of methylated PKM2 in immunoprecipitated PKM2 from parental and CARM1 KO cells. (i) Colony formation assays in parental MCF7, PKM2 KO, PKM2wt/shPKM1 and PKM2mut/shPKM1 cells. (j) Cell apoptosis measured by Annexin V and propidium iodide (PI) staining in parental MCF7, PKM2 KO, PKM2wt/shPKM1 and PKM2mut/shPKM1 cells. In b–d,f,h, data are shown as Mean ± SD and statistics source data are available in Supplementary Table 7. Statistical significance was assessed using two-tailed t-test (b,c,g) and ANOVA (d,f). ns: not significant. In e,h–j, data represent one of the three independent experiments with similar results. Unprocessed original scans of blots are shown in Supplementary Figure 9.
Supplementary Figure 4 Inhibition of PKM2 methylation leads to increase of mitochondrial ROS levels.
(a–c) The ROS levels in parental MCF7 and MCF7 PKM2 KO cells (a); MCF7 expressing PKM2wt/shPKM1 and PKM2mut/shPKM1 (b); parental MCF7 and CARM1 KO cells (c) (n = 3 independent experiments). (d,f) Relative NADPH/NADP + ratio (d) and GSH concentration (f) in parental MCF7, PKM2 KO, PKM2wt/shPKM1 and PKM2mut/shPKM1 cells (n = 3 independent experiments). (e,g) Relative NADPH/NADP + ratios (e) and GSH concentrations (g) in parental MCF7 and CARM1 KO cells (n = 3). (h,i) Relative NADPH/NADP + ratios (h) and GSH concentrations (i) in parental MDA-MB-231, PKM2 KO, PKM2wt/shPKM1 and PKM2mut/shPKM1 cells (n = 3 independent experiments). (j–l) Cell growth measured by MTT assays in MCF7 PKM2mut/shPKM1 (j) or MDA-MB-231 PKM2mut/shPKM1 (k) or MCF7 CARM1 KO (l) cells treated with mitoTEMPO (n = 10 independent experiments). (m) Images of migrated MDA-MB-231 PKM2mut/shPKM1 cells treated with mitoTEMPO. Scale bars, 50 μm. (n) Cell growth in MCF7 PKM2mut/shPKM1 or MDA-MB-231 PKM2mut/shPKM1 or MCF7 CARM1 KO cells treated with glutathione (1mM) (n = 3 independent experiments). (o) Images of migrated MDA-MB-231 PKM2mut/shPKM1 cells treated with glutathione. Scale bars, 50 μm. In a–l,n, data are shown as Mean ± SD and statistics source data are available in Supplementary Table 7. Statistical significance was assessed using two-tailed t-test (a–c,e,g) and ANOVA (d,f,h–l,n). In m,o, data represent one of the two independent experiments with similar results. ∗p < 0.05, ∗∗p < 0.01 ∗∗∗p < 0.001, ns: not significant.
Supplementary Figure 5 PKM2 methylation suppresses mitochondrial membrane potential and mitochondrial DNA content.
(a) Mitochondrial membrane potential (ΔΨ) measured by the incorporation of TMRE dye in MDA-MB-231 cells (n = 3 independent experiments). (b) Mitochondrial DNA (mtDNA) content in parental MCF7, PKM2 KO, PKM2wt/shPKM1 and PKM2mut/shPKM1 cells (n = 3 independent experiments). In a,b, data are shown as Mean ± SD and statistics source data are available in Supplementary Table 7. Statistical significance was assessed using ANOVA (a,b). ∗p < 0.05, ∗∗p < 0.01 ∗∗∗p < 0.001, ns: not significant.
Supplementary Figure 6 MAM localized PKM2 interacts with and suppresses IP3Rs expression in methylation-dependent manner.
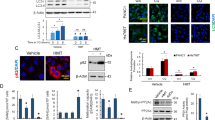
(a) Western blot analyses of PKM1 and PKM2 in cytosolic and mitochondria fractions derived from parental MCF7 or PKM2 KO cells. VDAC and tubulin serve as mitochondria and cytoplasm markers, respectively. (b) Confocal images of PKM2 localization in mitochondria. HSPA9 serves as a positive control which largely overlap with PKM2 staining. (c) Western blot analyses of wild type or mutant PKM2 in cytosolic and mitochondria fractions from MCF7 PKM2wt/shPKM1 and PKM2mut/shPKM1 cells. (d) List of selected ER and mitochondrial proteins that interact with wild type PKM2 or methylation-defective PKM2. Flag-tagged wild type or mutant PKM2 were transiently transfected into HEK293T cells. Flag-tagged PKM2 was pulled down from cell lysates and the interacting proteins were analyzed by mass spectrometry. The numbers of the detected peptides for each protein are indicated. (e) Venn diagram of PKM2 interacting proteins identified in Supplementary Fig. 6d overlapped with the altered proteins in response to PKM2 KO in MCF7 cells (Fig. 3c). 22 PKM2 interacting proteins were upregulated and 13 PKM2 interacting proteins were downregulated. ITPRs are also known as IP3Rs. (f) Western blot analysis of IP3R1 and IP3R3 in MCF7 cells overexpressing Flag-PKM1. (g) Co-immunoprecipitation of IP3R3 with PKM2 from MCF7 and MDA-MB-231 cell lysates. (h) Western blot analyses of IP3R3 protein levels in parental MCF7, CARM1 KO, or parental MCF7 treated with DMSO or TEPP-46. (i) Q-PCR analyses of mRNA levels of IP3R1, IP3R2 and IP3R3 in parental MCF7 and MDA-MB-231 cells and their respective PKM2 KO clones (n = 3 independent experiments). Data are shown as Mean ± SD and statistics source data are available in Supplementary Table 7. In a–c,f–h, data represent one of the three independent experiments with similar results. Unprocessed original scans of blots are shown in Supplementary Figure 9.
Supplementary Figure 7 Methylated PKM2 restrains mitochondrial addiction to Ca2+ through IP3Rs.
(a) Western blot analysis of relative IP3R3 in MCF7 and MDA-MB-231 cells. (b) Western blot analysis of IP3R3 knockdown efficiency in MCF7 PKM2 KO and MDA-MB-231 cells. (c) The gating strategy of flow cytometry. (d) Representative images of parental MDA-MB-231, PKM2 KO, PKM2wt/shPKM1 and PKM2mut/shPKM1 cells after treatment with 5 μM XeB for 24 hours. Scale bars, 50 μm. In a,b, data represent one of the three independent experiments with similar results. Unprocessed original scans of blots are shown in Supplementary Figure 9.
Supplementary Figure 8 Cellular PKM2 methylation can be inhibited by unimolecular nanoparticle (UMNP) loaded with non-methyl-PKM2 peptide.
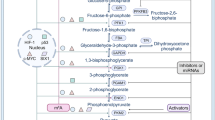
(a) Assessing the proportion of the endogenous methylated PKM2 by immunoprecipitation using the excess amount of methyl-PKM2 antibody. The amount of precipitated methyl-PKM2 is estimated by subtracting the PKM2 left in the flow-through (FT) fraction from the input following detection with PKM2 antibody. The western blot bands were quantified using ImageJ software. (b) The chemical structure of the unique unimolecular nanoparticles (UMNP) for PKM2 peptide delivery. (c) Illustration of the UMNP used for PKM2 peptide delivery. (d) Synthesis scheme of the multi-arm star block copolymer poly(amidoamine)–poly(aspartate diethyltriamine-aconitic acid-r-imidazole)-poly(ethylene glycol) (PAMAM-PAsp(DET-Aco-r-Im)-PEG-TAT). (e) 1H NMR spectrum of the multi-arm star block copolymer PAMAM-PAsp(DET-Aco-r-Im)-PEG-TAT. The * represents the solvent residual peak. (f) Dynamic light scattering (DLS) histogram of the UMNPs. (g) The relative pyruvate kinase activity of PKM2 in MDA-MB-231 cells treated with UMNP-methyl-peptide or UMNP-non-methyl-peptide (n = 3 independent experiments). Data are shown as Mean ± SD and statistics source data are available in Supplementary Table 7. Statistical significance was assessed using two-tailed t-test. ns: not significant. (h) The schematic diagram of energy homeostasis regulated by PKM2 methylation in cancer cells. CARM1 methylates dimeric PKM2 which associates with IP3Rs to inhibit Ca2+ influx from ER to mitochondria, resulting in increased PDH phosphorylation and decreased oxidative phosphorylation. Inhibiting PKM2 methylation by knocking out CARM1 or PKM2 or with a competitive PKM2 peptide increases IP3Rs expression, consequently increased [Ca2+]mito, de-phosphorylated PDH, and increased oxidative phosphorylation. The cell survival depends on [Ca2+]mito and is sensitive to IP3R inhibition. In a, data represent one of the three independent experiments with similar results. Unprocessed original scans of blots are shown in Supplementary Figure 9.
Supplementary information
Supplementary Information
Supplementary Information (PDF 38794 kb)
Supplementary Table 1
Supplementary Information
Supplementary Table 2
Supplementary Information
Supplementary Table 3
Supplementary Information
Supplementary Table 4
Supplementary Information
Supplementary Table 5
Supplementary Information
Supplementary Table 6
Supplementary Information
Supplementary Table 7
Supplementary Information
Rights and permissions
About this article
Cite this article
Liu, F., Ma, F., Wang, Y. et al. PKM2 methylation by CARM1 activates aerobic glycolysis to promote tumorigenesis. Nat Cell Biol 19, 1358–1370 (2017). https://doi.org/10.1038/ncb3630
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3630
This article is cited by
-
Transcriptional regulation and post-translational modifications in the glycolytic pathway for targeted cancer therapy
Acta Pharmacologica Sinica (2024)
-
Tyrosine phosphorylation of CARM1 promotes its enzymatic activity and alters its target specificity
Nature Communications (2024)
-
CIP2A induces PKM2 tetramer formation and oxidative phosphorylation in non-small cell lung cancer
Cell Discovery (2024)
-
The role of PKM2 in cancer progression and its structural and biological basis
Journal of Physiology and Biochemistry (2024)
-
Exploring Epigenetic and Genetic Modulation in Animal Responses to Thermal Stress
Molecular Biotechnology (2024)