Abstract
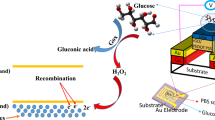
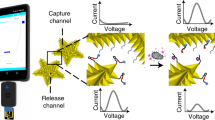
Portable, low-cost and quantitative detection of a broad range of targets at home and in the field has the potential to revolutionize medical diagnostics and environmental monitoring. Despite many years of research, very few such devices are commercially available. Taking advantage of the wide availability and low cost of the pocket-sized personal glucose meter—used worldwide by diabetes sufferers—we demonstrate a method to use such meters to quantify non-glucose targets, ranging from a recreational drug (cocaine, 3.4 µM detection limit) to an important biological cofactor (adenosine, 18 µM detection limit), to a disease marker (interferon-gamma of tuberculosis, 2.6 nM detection limit) and a toxic metal ion (uranium, 9.1 nM detection limit). The method is based on the target-induced release of invertase from a functional-DNA–invertase conjugate. The released invertase converts sucrose into glucose, which is detectable using the meter. The approach should be easily applicable to the detection of many other targets through the use of suitable functional-DNA partners (aptamers, DNAzymes or aptazymes).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Daar, A. S. et al. Top ten biotechnologies for improving health in developing countries. Nat. Genet. 32, 229–232 (2002).
Martinez, A. W., Phillips, S. T., Butte, M. J. & Whitesides, G. M. Patterned paper as a platform for inexpensive, low-volume, portable bioassays. Angew. Chem. Int. Ed. 46, 1318–1320 (2007).
Fan, C., Plaxco, K. W. & Heeger, A. J. Biosensors based on binding-modulated donor–acceptor distances. Trends Biotechnol. 23, 186–192 (2005).
Xia, F. et al. Colorimetric detection of DNA, small molecules, proteins, and ions using unmodified gold nanoparticles and conjugated polyelectrolytes. Proc. Natl Acad. Sci. USA 107, 10837–10841 (2010).
Zhang, J., Campbell, R. E., Ting, A. Y. & Tsien, R. Y. Creating new fluorescent probes for cell biology. Nat. Rev. Mol. Cell. Biol. 3, 906–918 (2002).
Giepmans, B. N. G., Adams, S. R., Ellisman, M. H. & Tsien, R. Y. Review–the fluorescent toolbox for assessing protein location and function. Science 312, 217–224 (2006).
Wegner, S. V., Okesli, A., Chen, P. & He, C. Design of an emission ratiometric biosensor from MerR family proteins: a sensitive and selective sensor for Hg2+ . J. Am. Chem. Soc. 129, 3474–3475 (2007).
Murray, R. W. Nanoelectrochemistry: metal nanoparticles, nanoelectrodes, and nanopores. Chem. Rev. 108, 2688–2720 (2008).
Wang, F. et al. Simultaneous phase and size control of upconversion nanocrystals through lanthanide doping. Nature 463, 1061–1065 (2010).
Favier, F., Walter, E. C., Zach, M. P., Benter, T. & Penner, R. M. Hydrogen sensors and switches from electrodeposited palladium mesowire arrays. Science 293, 2227–2231 (2001).
Clark, L. C. & Lyons, C. Electrode systems for continuous monitoring in cardiovascular surgery. Ann. NY Acad. Sci. 102, 29–45 (1962).
Montagnana, M., Caputo, M., Giavarina, D. & Lippi, G. Overview on self-monitoring of blood glucose. Clin. Chim. Acta 402, 7–13 (2009).
Carroll, A. E., Marrero, D. G. & Downs, S. M. The HealthPia GlucoPack (TM) diabetes phone: a usability study. Diabetes Technol. Ther. 9, 158–164 (2007).
Nie, Z., Deiss, F., Liu, X., Akbulut, O. & Whitesides, G. M. Integration of paper-based microfluidic devices with commercial electrochemical readers. Lab Chip 10, 3163–3169 (2010).
Robertson, D. L. & Joyce, G. F. Selection in vitro of an RNA enzyme that specifically cleaves single-stranded-DNA. Nature 344, 467–468 (1990).
Breaker, R. R. & Joyce, G. F. A DNA enzyme that cleaves RNA. Chem. Biol. 1, 223–229 (1994).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Breaker, R. R. Engineered allosteric ribozymes as biosensor components. Curr. Opin. Biotechnol. 13, 31–39 (2002).
Lee, J. F., Hesselberth, J. R., Meyers, L. A. & Ellington, A. D. Aptamer database. Nucleic Acids Res. 32, D95–D100 (2004).
Navani, N. K. & Li, Y. Nucleic acid aptamers and enzymes as sensors. Curr. Opin. Chem. Biol. 10, 272–281 (2006).
Song, S., Wang, L., Li, J., Fan, C. & Zhao, J. Aptamer-based biosensors. Trends Anal. Chem. 27, 108–117 (2008).
Liu, J., Cao, Z. & Lu, Y. Functional nucleic acid sensors. Chem. Rev. 109, 1948–1998 (2009).
Li, J. & Lu, Y. A highly sensitive and selective catalytic DNA biosensor for lead ions. J. Am. Chem. Soc. 122, 10466–10467 (2000).
Nutiu, R. & Li, Y. Structure-switching signaling aptamers. J. Am. Chem. Soc. 125, 4771–4778 (2003).
Yang, C. J., Jockusch, S., Vicens, M., Turro, N. J. & Tan, W. Light-switching excimer probes for rapid protein monitoring in complex biological fluids. Proc. Natl Acad. Sci. USA 102, 17278–17283 (2005).
Csordas, A. et al. Detection of proteins in serum by micromagnetic aptamer PCR (MAP) technology. Angew. Chem. Int. Ed. 49, 355–358 (2010).
Oh, S. S., Plakos, K., Lou, X. H., Xiao, Y. & Soh, H. T. In vitro selection of structure-switching, self-reporting aptamers. Proc. Natl Acad. Sci. USA 107, 14053–14058 (2010).
Liu, J. & Lu, Y. A colorimetric lead biosensor using DNAzyme-directed assembly of gold nanoparticles. J. Am. Chem. Soc. 125, 6642–6643 (2003).
Willner, I., Shlyahovsky, B., Zayats, M. & Willner, B. DNAzymes for sensing, nanobiotechnology and logic gate applications. Chem. Soc. Rev. 37, 1153–1165 (2008).
Xue, X., Wang, F. & Liu, X. One-step, room temperature, colorimetric detection of mercury (Hg2+) using DNA/nanoparticle conjugates. J. Am. Chem. Soc. 130, 3244–3245 (2008).
Willner, I. & Zayats, M. Electronic aptamer-based sensors. Angew. Chem. Int. Ed. 46, 6408–6418 (2007).
Lubin, A. A. & Plaxco, K. W. Folding-based electrochemical biosensors: the case for responsive nucleic acid architectures. Acc. Chem. Res. 43, 496–505 (2010).
Swensen, J. S. et al. Continuous, real-time monitoring of cocaine in undiluted blood serum via a microfluidic, electrochemical aptamer-based sensor. J. Am. Chem. Soc. 131, 4262–4266 (2009).
Yigit, M. V., Mazumdar, D. & Lu, Y. MRI detection of thrombin with aptamer functionalized superparamagnetic iron oxide nanoparticles. Bioconj. Chem. 19, 412–417 (2008).
Liu, J., Mazumdar, D. & Lu, Y. A simple and sensitive 'dipstick' test in serum based on lateral flow separation of aptamer-linked nanostructures. Angew. Chem. Int. Ed. 45, 7955–7959 (2006).
Niemeyer, C. M. Semisynthetic DNA–protein conjugates for biosensing and nanofabrication. Angew. Chem. Int. Ed. 49, 1200–1216 (2010).
Stojanovic, M. N., de Prada, P. & Landry, D. W. Aptamer-based folding fluorescent sensor for cocaine. J. Am. Chem. Soc. 123, 4928–4931 (2001).
Huizenga, D. E. & Szostak, J. W. A DNA aptamer that binds adenosine and ATP. Biochemistry 34, 656–665 (1995).
Vallon, V., Muhlbauer, B. & Osswald, H. Adenosine and kidney function. Physiol. Rev. 86, 901–940 (2006).
Sassanfar, M. & Szostak, J. W. An RNA motif that binds ATP. Nature 364, 550–553 (1993).
Lee, P. P., Ramanathan, M., Hunt, C. A. & Garovoy, M. R. An oligonucleotide blocks interferon-γ signal transduction. Transplantation 62, 1297–1301 (1996).
Balasubrananian, V., Nguyen, L. T., Balasubramanian, S. V. & Ramanathan, M. Interferon-γ-inhibitory oligodeoxynucleotides alter the conformation of interferon-γ. Mol. Pharmacol. 53, 926–932 (1998).
Tuleuova, N. et al. Development of an aptamer beacon for detection of interferon-γ. Anal. Chem. 82, 1851–1857 (2010).
Boehm, U., Klamp, T., Groot, M. & Howard, J. C. Cellular responses to interferon-γ. Annu. Rev. Immunol. 15, 749–795 (1997).
Pai, M., Riley, L. W. & Colford, J. M. Interferon-γ assays in the immunodiagnosis of tuberculosis: a systematic review. Lancet Infect. Dis. 4, 761–776 (2004).
Zhu, H. et al. A microdevice for multiplexed detection of T-cell-secreted cytokines. Lab Chip 8, 2197–2205 (2008).
Zhu, H. et al. Detecting cytokine release from single T-cells. Anal. Chem. 81, 8150–8156 (2009).
Liu, J. et al. A catalytic beacon sensor for uranium with parts-per-trillion sensitivity and millionfold selectivity. Proc. Natl Acad. Sci. USA 104, 2056–2061 (2007).
Acknowledgements
The authors thank the US Department of Energy (DE-FG02-08ER64568), National Institutes of Health (ES16865) and National Science Foundation (CTS-0120978) for financial support, and L.H. Tan and H.E. Ihms for the preparation of figures and proof reading of the manuscript.
Author information
Authors and Affiliations
Contributions
Y.L. and Y.X. conceived and designed the experiments. Y.X. performed the experiments and analysed the data. Y.L. and Y.X. co-wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 678 kb)
Rights and permissions
About this article
Cite this article
Xiang, Y., Lu, Y. Using personal glucose meters and functional DNA sensors to quantify a variety of analytical targets. Nature Chem 3, 697–703 (2011). https://doi.org/10.1038/nchem.1092
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1092
This article is cited by
-
Device integration of electrochemical biosensors
Nature Reviews Bioengineering (2023)
-
Glucometer-based biosensor for the determination of ractopamine in animal-derived foods using rolling circle amplification
Microchimica Acta (2023)
-
Acid-accelerated hydrolysis of NaBH4: a gas-generation reaction for diverse gas pressure biosensing
Microchimica Acta (2023)
-
Host-guest liquid gating mechanism with specific recognition interface behavior for universal quantitative chemical detection
Nature Communications (2022)
-
Batch-producible fibrous microelectrodes for enzyme-free electrochemical detection of glucose
Journal of Materials Science: Materials in Electronics (2022)