Abstract
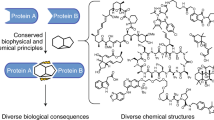
Supramolecular chemistry has recently emerged as a promising way to modulate protein functions, but devising molecules that will interact with a protein in the desired manner is difficult as many competing interactions exist in a biological environment (with solvents, salts or different sites for the target biomolecule). We now show that lysine-specific molecular tweezers bind to a 14-3-3 adapter protein and modulate its interaction with partner proteins. The tweezers inhibit binding between the 14-3-3 protein and two partner proteins—a phosphorylated (C-Raf) protein and an unphosphorylated one (ExoS)—in a concentration-dependent manner. Protein crystallography shows that this effect arises from the binding of the tweezers to a single surface-exposed lysine (Lys214) of the 14-3-3 protein in the proximity of its central channel, which normally binds the partner proteins. A combination of structural analysis and computer simulations provides rules for the tweezers' binding preferences, thus allowing us to predict their influence on this type of protein–protein interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Gale, P. A. (ed.) Supramolecular chemistry anniversary. Chem. Soc. Rev. 36, 141–142 (2007).
Gale, P. A. & Steed, J. W. (eds) Supramolecular Chemistry: From Molecules to Nanomaterials (Wiley, 2012).
Schrader, T. & Koch, S. Artificial protein sensors. Mol. BioSyst. 3, 241–248 (2007).
Peczuh, M. W. & Hamilton, A. D. Peptide and protein recognition by designed molecules. Chem. Rev. 100, 2479–2494 (2000).
Yin, H. & Hamilton, A. D. Strategies for targeting protein–protein interactions with synthetic agents. Angew. Chem. Int. Ed. 44, 4130–4163 (2005).
Martos, V., Castreño, P., Valero, J. & de Mendoza, J. Binding to protein surfaces by supramolecular multivalent scaffolds. Curr. Opin. Chem. Biol. 12, 698–706 (2008).
Gradl, S. N., Felix, J. P., Isacoff, E. Y., Garcia, M. L. & Trauner, D. Protein surface recognition by rational design: nanomolar ligands for potassium channels. J. Am. Chem. Soc. 125, 12668–12669 (2003).
Martos, V., Bell, S. C., Santos, E., Isacoff, E. Y., Trauner, D. & de Mendoza, J. Calix[4]arene-based conical-shaped ligands for voltage-dependent potassium channels. Proc. Natl Acad. Sci. USA 106, 10482–10486 (2009).
Gordo, S. et al. Stability and structural recovery of the tetramerization domain of p53–R337H mutant induced by a designed templating ligand. Proc. Natl Acad. Sci. USA 105, 16426–16431 (2008).
Nguyen, H. D., Dang, D. T., van Dongen, J. L. J. & Brunsveld, L. Supramolecular induced protein dimerization with cucurbit[8]uril. Angew. Chem. Int. Ed. 49, 895–898 (2010).
Ader, C. et al. A structural link between inactivation and block of a K+ channel. Nature Struct. Mol. Biol. 15, 605–612 (2008).
Chinai, J. M. et al. Molecular recognition of insulin by a synthetic receptor. J. Am. Chem. Soc. 133, 8810–8813 (2011).
McGovern, R. E., Fernandes, H., Khan, A. R., Powera, N. P. & Crowley, P. B. Protein camouflage in cytochrome c–calixarene complexes. Nature Chem. 4, 527–533 (2012).
Fokkens, M., Schrader, T. & Klärner, F-G. A molecular tweezer for lysine and arginine. J. Am. Chem. Soc. 127, 14415–14421 (2005).
Talbiersky, P., Bastkowski, F., Klärner, F-G. & Schrader, T. Molecular clip and tweezer introduce new mechanisms of enzyme inhibition. J. Am. Chem. Soc. 130, 9824–9828 (2008).
Klärner, F-G. et al. Molecular tweezer and clip in aqueous solution: unexpected self-assembly, powerful host–guest complex formation and quantum chemical 1H NMR shift calculation. J. Am. Chem. Soc. 128, 4831–4841 (2006).
Klärner, F-G. et al. The effect of molecular clips and tweezers on enzymatic reactions by binding coenzymes and basic amino acids. Pure Appl. Chem. 82, 991–999 (2010).
Wells, J. A. & McClendon, C. L. Reaching for high-hanging fruit in drug discovery at protein–protein interfaces. Nature 450, 1001–1009 (2007).
Morrison, D. K. The 14-3-3 proteins: integrators of diverse signaling cues that impact cell fate and cancer development. Trends Cell Biol. 19, 16–23 (2009).
Fantl, W. J. et al. Activation of Raf-1 by 14-3-3 proteins. Nature 371, 612–614 (1994).
Molzan, M. et al. Impaired binding of 14-3-3 to C-RAF in Noonan syndrome suggests new approaches in diseases with increased Ras signaling. Mol. Cell Biol. 19, 4698–4711 (2010).
Vassilev, A., Kaneko, K. J., Shu, H., Zhao, Y. & DePamphilis, M. L. TEAD/TEF transcription factors utilize the activation domain of YAP65, a Src/Yes-associated protein localized in the cytoplasm. Genes Dev. 15, 1229–1241 (2001).
Schumacher, B., Skwarczynska, M., Rose, R. & Ottmann, C. Structure of a 14-3-3σ-YAP phosphopeptide complex at 1.15 Å resolution. Acta Crystallogr. F 66, 978–984 (2010).
Rajagopalan, S., Sade, R. S., Townsley, F. M. & Fersht, A. R. Mechanistic differences in the transcriptional activation of p53 by 14-3-3 isoforms. Nucleic Acids Res. 38, 893–906 (2010).
Schumacher, B., Mondry, J., Thiel, P., Weyand, M. & Ottmann, C. Structure of the p53 C-terminus bound to 14-3-3: implications for stabilization of the p53 tetramer. FEBS Lett. 584, 1443–1448 (2010).
Fu, H., Coburn, J. & Collier, R. J. The eukaryotic host factor that activates exoenzyme S of Pseudomonas aeruginosa is a member of the 14-3-3 protein family. Proc. Natl Acad. Sci. USA 90, 2320–2324 (1993).
Ottmann, C. et al. Phosphorylation-independent interaction between 14-3-3 and exoenzyme S: from structure to pathogenesis. EMBO J. 26, 902–913 (2007).
Hermeking, H. The 14-3-3 cancer connection. Nature Rev. Cancer 3, 931–943 (2003).
Berg, D., Holzmann, C. & Riess, O. 14-3-3 proteins in the nervous system. Nature Rev. Neurosci. 9, 752–762 (2003).
Morrison, D. K. The 14-3-3 proteins: integrators of diverse signaling cues that impact cell fate and cancer development. Trends Cell Biol. 19, 16–23 (2009).
Andrews, R. K., Du, X. & Berndt, M. C. The 14-3-3zeta-GPIb-IX-V complex as an antiplatelet target. Drug News Perspect. 20, 285–292 (2007).
Wu, H., Ge, J. & Yao, S. Q. Microarray-assisted high-throughput identification of a cell-permeable small-molecule binder of 14-3-3 proteins. Angew. Chem. Int. Ed. 49, 6528–6532 (2010).
Corradi, V. et al. Identification of the first non-peptidic small molecule inhibitor of the c-Abl/14-3-3 protein–protein interactions able to drive sensitive and Imatinib-resistant leukemia cells to apoptosis. Bioorg. Med. Chem. Lett. 20, 6133–6137 (2010).
Rose, R. et al. Identification and structure of small-molecule stabilizers of 14-3-3 protein–protein interactions. Angew. Chem. Int. Ed. 49, 4129–4132 (2010).
Zhao, J. et al. Discovery and structural characterization of a small molecule 14-3-3 protein–protein interaction inhibitor. Proc. Natl Acad. Sci. USA 108, 16212–16216 (2011).
Richter, A., Rose, R., Hedberg, C., Waldmann, H. & Ottmann, C. An optimised small-molecule stabiliser of the 14-3-3–PMA2 protein–protein interaction. Chem. Eur. J. 21, 6520–6527 (2012).
Wang, B. et al. Isolation of high-affinity peptide antagonists of 14-3-3 proteins by phage display. Biochemistry 38, 12499–12504 (1999).
Wu, H., Ge, J. & Yao, S. Q. Microarray-assisted high-throughput identification of a cell-permeable small-molecule binder of 14-3-3 proteins. Angew. Chem. Int. Ed. 49, 6528–6532 (2010).
Corradi, V. et al. Identification of the first non-peptidic small molecule inhibitor of the c-Abl/14-3-3 protein–protein interactions able to drive sensitive and imatinib-resistant leukemia cells to apoptosis. Bioorg. Med. Chem. Lett. 20, 6133–6137 (2010).
Brooks, B. R. et al. CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J. Comput. Chem. 4, 187–217 (1983).
Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual Molecular Dynamics. J. Mol. Graphics 14, 33–38 (1996).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Sherwood, P. et al. QUASI: a general purpose implementation of the QM/MM approach and its application to problems in catalysis. J. Mol. Struct. (Teochem) 632, 1–28 (2003).
Grimme, S. Semiempirical GGA-type density functional constructed with a long-range dispersion correction. J. Comput. Chem. 27, 1787–1799 (2006).
Becke, A. D. Density functional thermochemistry. II. The effect of the Perdew–Wang generalized gradient correlation correction. J. Chem. Phys. 97, 9173–9177 (1992).
Schäfer, A., Horn, H. & Ahlrichs, R. Fully optimized contracted Gaussian basis sets for atoms Li to Kr. J. Chem. Phys. 97, 2571–2577 (1992).
Mackerell, A. D., Feig, M. Jr & Brooks, C. L. III . Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004).
Acknowledgements
C.O. thanks AstraZeneca, Bayer CropScience, Bayer Healthcare, Boehringer Ingelheim and Merck KGaA for support. E.S-G acknowledges a Liebig stipend and K.B-R. a pre-doctoral stipend from the Fonds der Chemischen Industrie. E.S-G thanks the Cluster of Excellence RESOLV (EXC 1069) funded by the Deutsche Forschungsgemeinschaft for financial support and W. Thiel for useful suggestions. The authors thank E. Hofmann, F. Syberg, I. Vetter and the SLS beamline staff for data collection at the Swiss Light Source, beamline PXII-X10SA, Paul Scherrer Institute, Villigen, Switzerland.
Author information
Authors and Affiliations
Contributions
D.B., R.R. and M.B. carried out the experiments. K.B.-R. and J.M.R.-A. performed the QM/MM and MD calculations. S.D. and C.W. synthesized the tweezers. F.-G.K. designed their synthesis. C.O., T.S. and E.S.-G. designed the experiments and calculations and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 7702 kb)
Supplementary Movie 1
Supplementary Movie 1 (WMV 6833 kb)
Supplementary Movie 2
Supplementary Movie 2 (WMV 5005 kb)
Rights and permissions
About this article
Cite this article
Bier, D., Rose, R., Bravo-Rodriguez, K. et al. Molecular tweezers modulate 14-3-3 protein–protein interactions. Nature Chem 5, 234–239 (2013). https://doi.org/10.1038/nchem.1570
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1570
This article is cited by
-
Domain-agnostic predictions of nanoscale interactions in proteins and nanoparticles
Nature Computational Science (2023)
-
A self-complementary macrocycle by a dual interaction system
Nature Communications (2022)
-
A molecular dynamics study of the complexation of tryptophan, phenylalanine and tyrosine amino acids with cucurbit[7]uril
Journal of Inclusion Phenomena and Macrocyclic Chemistry (2022)
-
Specific inhibition of the Survivin–CRM1 interaction by peptide-modified molecular tweezers
Nature Communications (2021)
-
Fluorescent artificial receptor-based membrane assay (FARMA) for spatiotemporally resolved monitoring of biomembrane permeability
Communications Biology (2020)