Abstract
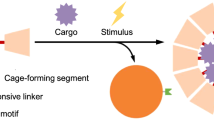
The interaction between a viral capsid and its genome governs crucial steps in the life cycle of a virus, such as assembly and genome uncoating. Tuning cargo–capsid interactions is also essential for successful design and cargo delivery in engineered viral systems. Here we investigate the interplay between cargo and capsid for the picorna-like Triatoma virus using a combined native mass spectrometry and atomic force microscopy approach. We propose a topology and assembly model in which heterotrimeric pentons that consist of five copies of structural proteins VP1, VP2 and VP3 are the free principal units of assembly. The interpenton contacts are established primarily by VP2. The dual role of the genome is first to stabilize the densely packed virion and, second, on an increase in pH to trigger uncoating by relaxing the stabilizing interactions with the capsid. Uncoating occurs through a labile intermediate state of the virion that reversibly disassembles into pentons with the concomitant release of protein VP4.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Speir, J. A. & Johnson, J. E. Nucleic acid packaging in viruses. Curr. Opin. Struct. Biol. 22, 65–71 (2012).
Douglas, T. & Young, M. Viruses: making friends with old foes. Science 312, 873–875 (2006).
Mateu, M. G. Mechanical properties of viruses analyzed by atomic force microscopy: a virological perspective. Virus Res. 168, 1–22 (2012).
Morton, V. L., Stockley, P. G., Stonehouse, N. J. & Ashcroft, A. E. Insights into virus capsid assembly from non-covalent mass spectrometry. Mass Spectrom. Rev. 27, 575–595 (2008).
Roos, W. H., Bruinsma, R. & Wuite, G. J. L. Physical virology. Nature Phys. 6, 733–743 (2010).
Uetrecht, C. & Heck, A. J. R. Modern biomolecular mass spectrometry and its role in studying virus structure, dynamics, and assembly. Angew. Chem. Int. Ed. 50, 8248–8262 (2011).
Uetrecht, C., Barbu, I. M., Shoemaker, G. K., Van Duijn, E. & Heck, A. J. R. Interrogating viral capsid assembly with ion mobility mass spectrometry. Nature Chem. 3, 126–132 (2011).
Bonning, B. C. & Miller, W. A. Dicistroviruses. Ann. Rev. Entomol. 55, 129–150 (2010).
Marti, G. A. et al. Prevalence and distribution of parasites and pathogens of Triatominae from Argentina, with emphasis on Triatoma infestans and Triatoma virus TrV. J. Invertebr. Pathol. 102, 233–237 (2009).
Muscio, O. A., La Torre, J. L., Bonder, M. A. & Scodeller, E. A. Triatoma virus pathogenicity in laboratory colonies of Triatoma infestans (Hemiptera: Reduviidae). J. Med. Entomol. 34, 253–256 (1997).
Rassi, A. Jr, Rassi, A. & Marin-Neto, J. A. Chagas disease. Lancet 375, 1388–1402 (2010).
Muscio, O. A., La Torre, J. L. & Scodeller, E. A. Characterization of Triatoma virus, a picorna-like virus isolated from the triatomine bug Triatoma infestans. J. Gen. Virol. 69, 2929–2934 (1988).
Czibener, C., La Torre, J. L., Muscio, O. A., Ugalde, R. A. & Scodeller, E. A. Nucleotide sequence analysis of Triatoma virus shows that it is a member of a novel group of insect RNA viruses. J. Gen. Virol. 81, 1149–1154 (2000).
Agirre, J. et al. Capsid protein identification and analysis of mature Triatoma virus (TrV) virions and naturally occurring empty particles. Virology 409, 91–101 (2011).
Tate, J. et al. The crystal structure of cricket paralysis virus: the first view of a new virus family. Nature Struct. Biol. 6, 765–774 (1999).
Kumar, S. & Bandyopadhyay, U. Free heme toxicity and its detoxification systems in human. Toxicol. Lett. 157, 175–188 (2005).
Schaub, G. A. in Advances in Insect Physiology (eds Simpson, S. J. and Casas, J) Ch. 4, 177–242 (Academic Press, 2009).
Oliveira, M. F. et al. Heme crystallization in the midgut of triatomine insects. Comp. Biochem. Physiol. C 146, 168–174 (2007).
Stiebler, R. et al. On the physico-chemical and physiological requirements of hemozoin formation promoted by perimicrovillar membranes in Rhodnius prolixus midgut. Insect Biochem. Mol. Biol. 40, 284–292 (2010).
Kollien, A. H., Grospietsch, T., Kleffmann, T., Zerbst-Boroffka, I. & Schaub, G. A. Ionic composition of the rectal contents and excreta of the reduviid bug Triatoma infestans. J. Insect Physiol. 47, 739–747 (2001).
Muscio, O., Bonder, M. A., La Torre, J. L. & Scodeller, E. A. Horizontal transmission of Triatoma virus through the fecal–oral route in Triatoma infestans (Hemiptera: Triatomidae). J. Med. Entomol. 37, 271–275 (2000).
Li, C., Wang, J. C., Taylor, M. W. & Zlotnick, A. In vitro assembly of an empty picornavirus capsid follows a dodecahedral path. J. Virol. 86, 13062–13069 (2012).
Benesch, J. L. P. Collisional activation of protein complexes: picking up the pieces. J. Am. Soc. Mass Spectrom. 20, 341–348 (2009).
Benesch, J. L. P., Ruotolo, B. T., Simmons, D. A. & Robinsons, C. V. Protein complexes in the gas phase: technology for structural genomics and proteomics. Chem. Rev. 107, 3544–3567 (2007).
Uetrecht, C., Rose, R. J., Van Duijn, E., Lorenzen, K. & Heck, A. J. R. Ion mobility mass spectrometry of proteins and protein assemblies. Chem. Soc. Rev. 39, 1633–1655 (2010).
Ruotolo, B. T., Benesch, J. L. P., Sandercock, A. M., Hyung, S. & Robinson, C. V. Ion mobility mass spectrometry analysis of large protein complexes. Nature Protocols 3, 1139–1152 (2008).
Evilevitch, A. et al. Effects of salts on internal DNA pressure and mechanical properties of phage capsids. J. Mol. Biol. 405, 18–23 (2011).
Carrasco, C. et al. DNA-mediated anisotropic mechanical reinforcement of a virus. Proc. Natl Acad. Sci. USA 103, 13706–13711 (2006).
Michel, J. P. et al. Nanoindentation studies of full and empty viral capsids and the effects of capsid protein mutations on elasticity and strength. Proc. Natl Acad. Sci. USA 103, 6184–6189 (2006).
Estrozi, L. F. et al. Phasing of the Triatoma virus diffraction data using a cryo-electron microscopy reconstruction. Virology 375, 85–93 (2008).
Castellanos, M., Pérez, R., Carrillo, P. J. P., De Pablo, P. J. & Mateu, M. G. Mechanical disassembly of single virus particles reveals kinetic intermediates predicted by theory. Biophys. J. 102, 2615–2624 (2012).
Ojosnegros, S. et al. Viral genome segmentation can result from a trade-off between genetic content and particle stability. PLoS Genetics 7, e1001344 (2011).
Fout, G. S., Medappa, K. C., Mapoles, J. E. & Rueckert, R. R. Radiochemical determination of polyamines in poliovirus and human rhinovirus 14. J. Biol. Chem. 259, 3639–3643 (1984).
Roos, W. H. et al. Squeezing protein shells: how continuum elastic models, molecular dynamics simulations, and experiments coalesce at the nanoscale. Biophys. J. 99, 1175–1181 (2010).
Bostina, M., Levy, H., Filman, D. J. & Hogle, J. M. Poliovirus RNA is released from the capsid near a twofold symmetry axis. J. Virol. 85, 776–783 (2011).
Agirre, J. et al. Cryo-TEM reconstructions of Triatoma virus particles: a clue to unravel genome delivery and capsid disassembly. J. Gen. Virol. http://dx.doi.org/10.1099/vir.0.048553-0 (2013).
Muscio, O. A., LaTorre, J. L. & Scodeller, E. A. Small nonoccluded viruses from triatomine bug Triatoma infestans (Hemiptera: Reduviidae). J. Invertebr. Pathol. 49, 218–220 (1987).
Kol, N. et al. A stiffness switch in human immunodeficiency virus. Biophys. J. 92, 1777–1783 (2007).
Van Den Heuvel, R. H. H. et al. Improving the performance of a quadrupole time-of-flight instrument for macromolecular mass spectrometry. Anal. Chem. 78, 7473–7483 (2006).
Tahallah, N., Pinkse, M., Maier, C. S. & Heck, A. J. R. The effect of the source pressure on the abundance of ions of noncovalent protein assemblies in an electrospray ionization orthogonal time-of-flight instrument. Rapid Commun. Mass Spectrom. 15, 596–601 (2001).
Lorenzen, K., Versluis, C., van Duijn, E., van den Heuvel, R. H. H. & Heck, A. J. R. Optimizing macromolecular tandem mass spectrometry of large non-covalent complexes using heavy collision gases. Int. J. Mass Spectrom. 268, 198–206 (2007).
Pringle, S. D. et al. An investigation of the mobility separation of some peptide and protein ions using a new hybrid quadrupole/travelling wave IMS/oa–ToF instrument. Int J. Mass Spectrom. 261, 1–12 (2007).
Bush, M. F. et al. Collision cross sections of proteins and their complexes: a calibration framework and database for gas-phase structural biology. Anal. Chem. 82, 9557–9565 (2010).
Mesleh, M. F., Hunter, J. M., Shvartsburg, A. A., Schatz, G. C. & Jarrold, M. F. Structural information from ion mobility measurements: effects of the long-range potential. J. Phys. Chem. 100, 16082–16086 (1996).
Shvartsburg, A. A. & Jarrold, M. F. An exact hard-spheres scattering model for the mobilities of polyatomic ions. Chem. Phys. Lett. 261, 86–91 (1996).
Uetrecht, C. et al. Stability and shape of hepatitis B virus capsids in vacuo. Angew. Chem. Int. Ed. 47, 6247–6251 (2008).
Roos, W. H. How to perform a nanoindentation experiment on a virus. Methods Mol. Biol. 783, 251–264 (2011).
Sader, J. E., Chon, J. W. M. & Mulvaney, P. Calibration of rectangular atomic force microscope cantilevers. Rev. Sci. Instrum. 70, 3967–3969 (1999).
Snijder, J., Ivanovska, I. L., Baclayon, M., Roos, W. H. & Wuite, G. J. L. Probing the impact of loading rate on the mechanical properties of viral nanoparticles. Micron 43, 1343–1350 (2012).
Horcas, I. et al. WSXM: a software for scanning probe microscopy and a tool for nanotechnology. Rev. Sci. Instrum. 78, 013705 (2007).
Acknowledgements
G.J.L.W. is supported by a NanoSci-E+ grant from FOM (Fundamenteel Onderzoek der Materie) under the ‘Physics of the Genome’ program and by a Vici grant from the Netherlands Organization for Scientific Research (NWO). J.S., C.U., R.J.R. and A.J.R.H. are supported by the Netherlands Proteomics Centre. A.J.R.H. is further supported by NWO under ALW-ECHO (819.02.10). R.S-E. is recipient of a predoctoral fellowship from the Basque Government (BG). G.A.M. is supported partially by PICT 2008-0035 and V188 (National University of La Plata). D.M.A.G. is supported by CYTED (Programa Internacional de Cooperación Científica y Tecnológica Multilateral) (209RT0364), by the BG (MV-2012-2-41; AE-2012-1-44, SPE1 1FB001) and by the Ministry of Science and Innovation (BFU2007-62062). We thank the Scientific Calculation Service of the UPV/EHU (SGIker) for computation resources. D.M.A.G. is visiting professor at the UPV/EHU.
Author information
Authors and Affiliations
Contributions
A.J.R.H., W.H.R., J.S. and D.M.A.G. conceptually conceived the research. G.A.M., J.A. and R.S-E. purified the virus samples. J.S., C.U., R.J.R. and A.J.R.H. performed and supervised the MS measurements. J.S. and W.H.R. performed and supervised the AFM measurements. R.S.E. and J.A. performed additional experiments. All authors contributed to interpreting the data and writing the paper. D.M.A.G., G.J.L.W. and A.J.R.H. supervised the work in the respective laboratories.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 1226 kb)
Rights and permissions
About this article
Cite this article
Snijder, J., Uetrecht, C., Rose, R. et al. Probing the biophysical interplay between a viral genome and its capsid. Nature Chem 5, 502–509 (2013). https://doi.org/10.1038/nchem.1627
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1627
This article is cited by
-
Complete and cooperative in vitro assembly of computationally designed self-assembling protein nanomaterials
Nature Communications (2021)
-
Physics of viral dynamics
Nature Reviews Physics (2021)
-
The effect of proteolytic enzymes and pH on GII.4 norovirus, during both interactions and non-interaction with Histo-Blood Group Antigens
Scientific Reports (2020)
-
Single-particle virology
Biophysical Reviews (2020)
-
Standard Proteoforms and Their Complexes for Native Mass Spectrometry
Journal of the American Society for Mass Spectrometry (2019)