Abstract
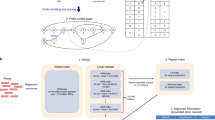
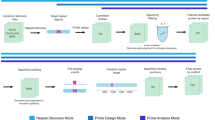
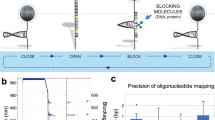
Complex DNA sequences are difficult to detect and profile, but are important contributors to human health and disease. Existing hybridization probes lack the capability to selectively bind and enrich hypervariable, long or repetitive sequences. Here, we present a generalized strategy for constructing modular hybridization probes (M-Probes) that overcomes these challenges. We demonstrate that M-Probes can tolerate sequence variations of up to 7 nt at prescribed positions while maintaining single nucleotide sensitivity at other positions. M-Probes are also shown to be capable of sequence-selectively binding a continuous DNA sequence of more than 500 nt. Furthermore, we show that M-Probes can detect genes with triplet repeats exceeding a programmed threshold. As a demonstration of this technology, we have developed a hybrid capture method to determine the exact triplet repeat expansion number in the Huntington's gene of genomic DNA using quantitative PCR.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
1000 Genomes Project Consortium. tA map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
1000 Genomes Project Consortium. tA global reference for human genetic variation. Nature 526, 68–74 (2015).
Mardis, E. R. Next-generation sequencing platforms. Annu. Rev. Anal. Chem. 6, 287–303 (2013).
Van Dijk, E. L., Auger, H., Jaszczyszyn, Y. & Thermes, C. Ten years of next-generation sequencing technology. Trends Genet. 30, 418–426 (2014).
Quail, M. A. et al. A tale of three next generation sequencing platforms: comparison of Ion Torrent, Pacific Biosciences and Illumina MiSeq sequencers. BMC Genomics 13, 341 (2012).
Kircher, M. & Kelso, J. High throughput DNA sequencing concepts and limitations. Bioessays 32, 524–536 (2010).
Gunderson, K. L., Steemers, F. J., Lee, G., Mendoza, L. G. & Chee, M. S. A genome-wide scalable SNP genotyping assay using microarray technology. Nat. Genet. 37, 549–554 (2005).
Higuchi, R., Fockler, C., Dollinger, G. & Watson, R. Kinetic PCR analysis: real-time monitoring of DNA amplification reactions. Biotechnology 11, 1026–1030 (1993).
Kim, T. M., Laird, P. W. & Park, P. J. The landscape of microsatellite instability in colorectal and endometrial cancer genomes. Cell 155, 858–868 (2013).
Duyao, M. et al. Trinucleotide repeat length instability and age of onset in Huntington's disease. Nat. Genet. 4, 387–392 (1993).
Iyer, R. R., Pluciennik, A., Napierala, M. & Wells, R. D. DNA triplet repeat expansion and mismatch repair. Annu. Rev. Biochem. 84, 199–226 (2015).
Freeman, J. D., Warren, R. L., Webb, J. R., Nelson, B. H. & Holt, R. A. Profiling the T-cell receptor beta-chain repertoire by massively parallel sequencing. Genome Res. 19, 1817–1824 (2009).
Benichou, J., Ben-Hamo, R., Louzoun, Y. & Efroni, S. Rep-Seq: uncovering the immunological repertoire through next-generation sequencing. Immunology 135, 183–191 (2012).
Feuk, L., Carson, A. R. & Scherer, S. W. Structural variation in the human genome. Nat. Rev. Genet. 7, 85–97 (2006).
Yang, L. et al. Diverse mechanisms of somatic structural variations in human cancer genomes. Cell 153, 919–929 (2013).
Wang, J. et al. CREST maps somatic structural variation in cancer genomes with base-pair resolution. Nat. Methods 8, 652–654 (2011).
Suggs, S. V., Wallace, R. B., Hirose, T., Kawashima, E. H. & Itakura, K. Use of synthetic oligonucleotides as hybridization probes: isolation of cloned cDNA sequences for human beta 2-microglobulin. Proc. Natl Acad. Sci. USA 78, 6613–6617 (1981).
Parkhurst, K. M. & Parkhurst, L. J. Kinetic studies by fluorescence resonance energy transfer employing a double-labeled oligonucleotide: hybridization to the oligonucleotide complement and to single-stranded DNA. Biochemistry 34, 285–292 (1995).
Tyagi, S. & Kramer, F. R. Molecular beacons: probes that fluoresce upon hybridization. Nat. Biotechnol. 14, 303–308 (1996).
Lanman, R. B. et al. Analytical and clinical validation of a digital sequencing panel for quantitative, highly accurate evaluation of cell-free circulating tumor DNA. PLoS ONE 10, e0140712 (2015).
Clark, M. J. et al. Performance comparison of exome DNA sequencing technologies. Nat. Biotechnol. 29, 908–914 (2011).
Zhang, D. Y. & Winfree, E. Control of DNA strand displacement kinetics using toehold exchange. J. Am. Chem. Soc. 131, 17303–17314 (2009).
Zhang, D. Y., Chen, S. X. & Yin, P. Thermodynamic optimization of nucleic acid hybridization specificity. Nat. Chem. 4, 208–214 (2012).
International HapMap Consortium et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Budworth, H. & McMurray, C. T. Problems and solutions for the analysis of somatic CAG repeat expansion and their relationship to huntington's disease toxicity. Rare Dis. 4, e1131885 (2016).
Jama, M., Millson, A., Miller, C. E. & Lyon, E. Triplet repeat primed PCR simplifies testing for Huntington disease. J. Mol. Diagn. 15, 255–262 (2016).
Bonifazi, E. et al. Use of RNA fluorescence in situ hybridization in the prenatal molecular diagnosis of myotonic dystrophy type I. Clin Chem. 52, 319–322 (2006).
Kern, A. & Seitz, O. Template-directed ligation on repetitive DNA sequences: a chemical method to probe the length of Huntington DNA. Chem. Sci. 6, 724–728 (2015).
Wu, L. R. et al. Continuously tunable nucleic acid hybridization probes. Nat. Methods 12, 1191–1196 (2015).
Wang, J. S. & Zhang, D. Y. Simulation-guided DNA probe design for consistently ultraspecific hybridization. Nat. Chem. 7, 545–553 (2015).
Kwok, S., Chang, S. Y., Sninsky, J. J. & Wang, A. A guide to the design and use of mismatched and degenerate primers. Genome Res. 3, S39–S47 (1994).
Ohtsuka, E., Matsuki, S., Ikehara, M., Takahashi, Y. & Matsubara, K. An alternative approach to deoxyoligonucleotides as hybridization probes by insertion of deoxyinosine at ambiguous codon positions. J. Biol. Chem. 260, 2605–2608 (1985).
Acknowledgements
The authors thank A. Pinto for discussions. This work was funded by the Cancer Prevention Research Institute of Texas (grant RP140132 to D.Y.Z.) and by the National Human Genome Research Institute (grant R01HG008752 to D.Y.Z.).
Author information
Authors and Affiliations
Contributions
J.S.W., Y.H.Y. and D.Y.Z. conceived the project. J.S.W. and Y.H.Y. performed experiments and data analysis. J.S.W., Y.H.Y. and D.Y.Z. wrote the manuscript. J.S.W. and Y.H.Y. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 8766 kb)
Rights and permissions
About this article
Cite this article
Wang, J., Yan, Y. & Zhang, D. Modular probes for enriching and detecting complex nucleic acid sequences. Nature Chem 9, 1222–1228 (2017). https://doi.org/10.1038/nchem.2820
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2820
This article is cited by
-
Encoding quantized fluorescence states with fractal DNA frameworks
Nature Communications (2020)
-
Expanding detection windows for discriminating single nucleotide variants using rationally designed DNA equalizer probes
Nature Communications (2020)