Abstract
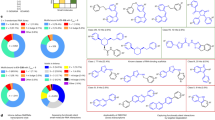
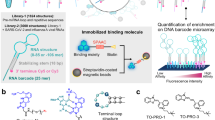
Oligonucleotides are designed to target RNA using base pairing rules, but they can be hampered by poor cellular delivery and nonspecific stimulation of the immune system. Small molecules are preferred as lead drugs or probes but cannot be designed from sequence. Herein, we describe an approach termed Inforna that designs lead small molecules for RNA from solely sequence. Inforna was applied to all human microRNA hairpin precursors, and it identified bioactive small molecules that inhibit biogenesis by binding nuclease-processing sites (44% hit rate). Among 27 lead interactions, the most avid interaction is between a benzimidazole (1) and precursor microRNA-96. Compound 1 selectively inhibits biogenesis of microRNA-96, upregulating a protein target (FOXO1) and inducing apoptosis in cancer cells. Apoptosis is ablated when FOXO1 mRNA expression is knocked down by an siRNA, validating compound selectivity. Markedly, microRNA profiling shows that 1 only affects microRNA-96 biogenesis and is at least as selective as an oligonucleotide.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Arambula, J.F., Ramisetty, S.R., Baranger, A.M. & Zimmerman, S.C. A simple ligand that selectively targets CUG trinucleotide repeats and inhibits MBNL protein binding. Proc. Natl. Acad. Sci. USA 106, 16068–16073 (2009).
Childs-Disney, J.L., Hoskins, J., Rzuczek, S.G., Thornton, C.A. & Disney, M.D. Rationally designed small molecules targeting the RNA that causes myotonic dystrophy type 1 are potently bioactive. ACS Chem. Biol. 7, 856–862 (2012).
Ofori, L.O., Hoskins, J., Nakamori, M., Thornton, C.A. & Miller, B.L. From dynamic combinatorial 'hit' to lead: in vitro and in vivo activity of compounds targeting the pathogenic RNAs that cause myotonic dystrophy. Nucleic Acids Res. 40, 6380–6390 (2012).
Kumar, A. et al. Chemical correction of pre-mRNA splicing defects associated with sequestration of muscleblind-like 1 protein by expanded r(CAG)-containing transcripts. ACS Chem. Biol. 7, 496–505 (2012).
Bose, D. et al. The tuberculosis drug streptomycin as a potential cancer therapeutic: inhibition of miR-21 function by directly targeting its precursor. Angew. Chem. Int. Edn Engl. 51, 1019–1023 (2012).
Stelzer, A.C. et al. Discovery of selective bioactive small molecules by targeting an RNA dynamic ensemble. Nat. Chem. Biol. 7, 553–559 (2011).
Parsons, J. et al. Conformational inhibition of the hepatitis C virus internal ribosome entry site RNA. Nat. Chem. Biol. 5, 823–825 (2009).
Davidson, A. et al. Simultaneous recognition of HIV-1 TAR RNA bulge and loop sequences by cyclic peptide mimics of Tat protein. Proc. Natl. Acad. Sci. USA 106, 11931–11936 (2009).
Guan, L. & Disney, M.D. Recent advances in developing small molecules targeting RNA. ACS Chem. Biol. 7, 73–86 (2012).
Thomas, J.R. & Hergenrother, P.J. Targeting RNA with small molecules. Chem. Rev. 108, 1171–1224 (2008).
Yonath, A. & Bashan, A. Ribosomal crystallography: initiation, peptide bond formation, and amino acid polymerization are hampered by antibiotics. Annu. Rev. Microbiol. 58, 233–251 (2004).
Mathews, D.H. et al. Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure. Proc. Natl. Acad. Sci. USA 101, 7287–7292 (2004).
Woese, C. & Pace, N. The RNA World 2nd edn. (eds. Gesteland, R.F., Cech, T.R. & Atkins, J.F.) 91–117 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1993).
Batey, R.T., Rambo, R.P. & Doudna, J.A. Tertiary motifs in RNA structure and folding. Angew. Chem. Int. Edn Engl. 38, 2326–2343 (1999).
Spahn, C.M. et al. Hepatitis C virus IRES RNA-induced changes in the conformation of the 40s ribosomal subunit. Science 291, 1959–1962 (2001).
Disney, M.D. et al. Two-dimensional combinatorial screening identifies specific aminoglycoside-RNA internal loop partners. J. Am. Chem. Soc. 130, 11185–11194 (2008).
Velagapudi, S.P., Seedhouse, S.J. & Disney, M.D. Structure-activity relationships through sequencing (StARTS) defines optimal and suboptimal RNA motif targets for small molecules. Angew. Chem. Int. Edn Engl. 49, 3816–3818 (2010).
Velagapudi, S.P., Seedhouse, S.J., French, J. & Disney, M.D. Defining the RNA internal loops preferred by benzimidazole derivatives via 2D combinatorial screening and computational analysis. J. Am. Chem. Soc. 133, 10111–10118 (2011).
Jiang, Q. et al. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic Acids Res. 37, D98–D104 (2009).
Griffiths-Jones, S., Saini, H.K., van Dongen, S. & Enright, A.J. miRBase: tools for microRNA genomics. Nucleic Acids Res. 36, D154–D158 (2008).
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Ambros, V. et al. A uniform system for microRNA annotation. RNA 9, 277–279 (2003).
Wu, M. & Turner, D.H. Solution structure of (rGCGGACGC)2 by two-dimensional NMR and the iterative relaxation matrix approach. Biochemistry 35, 9677–9689 (1996).
SantaLucia, J. Jr. & Turner, D.H. Structure of (rGGCGAGCC)2 in solution from NMR and restrained molecular dynamics. Biochemistry 32, 12612–12623 (1993).
Kozomara, A. & Griffiths-Jones, S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 39, D152–D157 (2011).
Krützfeldt, J. et al. Silencing of microRNAs in vivo with 'antagomirs'. Nature 438, 685–689 (2005).
Ebert, M.S. & Sharp, P.A. MicroRNA sponges: progress and possibilities. RNA 16, 2043–2050 (2010).
Obad, S. et al. Silencing of microRNA families by seed-targeting tiny LNAs. Nat. Genet. 43, 371–378 (2011).
Mathews, D.H. Using an RNA secondary structure partition function to determine confidence in base pairs predicted by free energy minimization. RNA 10, 1178–1190 (2004).
Childs-Disney, J.L., Wu, M., Pushechnikov, A., Aminova, O. & Disney, M.D. A small molecule microarray platform to select RNA internal loop-ligand interactions. ACS Chem. Biol. 2, 745–754 (2007).
Pilch, D.S. et al. Binding of a hairpin polyamide in the minor groove of DNA: sequence-specific enthalpic discrimination. Proc. Natl. Acad. Sci. USA 93, 8306–8311 (1996).
Pinto, I.G., Guilbert, C., Ulyanov, N.B., Stearns, J. & James, T.L. Discovery of ligands for a novel target, the human telomerase RNA, based on flexible-target virtual screening and NMR. J. Med. Chem. 51, 7205–7215 (2008).
Disney, M.D. et al. A small molecule that targets r(CGG)exp and improves defects in fragile X-associated tremor ataxia syndrome. ACS Chem. Biol. 7, 1711–1718 (2012).
Luzhkov, V.B. et al. Virtual screening and bioassay study of novel inhibitors for dengue virus mRNA cap (nucleoside-2′O)-methyltransferase. Bioorg. Med. Chem. 15, 7795–7802 (2007).
Xu, S., Witmer, P.D., Lumayag, S., Kovacs, B. & Valle, D. MicroRNA (miRNA) transcriptome of mouse retina and identification of a sensory organ–specific miRNA cluster. J. Biol. Chem. 282, 25053–25066 (2007).
Stenvang, J., Petri, A., Lindow, M., Obad, S. & Kauppinen, S. Inhibition of microRNA function by antimiR oligonucleotides. Silence. 3, 1 (2012).
Xie, L. et al. FOXO1 is a tumor suppressor in classical Hodgkin lymphoma. Blood 119, 3503–3511 (2012).
Guttilla, I.K. & White, B.A. Coordinate regulation of FOXO1 by miR-27a, miR-96, and miR-182 in breast cancer cells. J. Biol. Chem. 284, 23204–23216 (2009).
Dansen, T.B. & Burgering, B.M. Unravelling the tumor-suppressive functions of FOXO proteins. Trends Cell Biol. 18, 421–429 (2008).
Huang, H. & Tindall, D.J. FOXO factors: a matter of life and death. Future Oncol. 2, 83–89 (2006).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Tran, T. & Disney, M.D. Identifying the preferred RNA motifs and chemotypes that interact by probing millions of combinations. Nat. Commun. 3, 1125 (2012).
Hawkins, P.C., Skillman, A.G. & Nicholls, A. Comparison of shape-matching and docking as virtual screening tools. J. Med. Chem. 50, 74–82 (2007).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha–DGCR8 complex. Cell 125, 887–901 (2006).
Berezikov, E. et al. Phylogenetic shadowing and computational identification of human microRNA genes. Cell 120, 21–24 (2005).
Michlewski, G., Guil, S., Semple, C.A. & Caceres, J.F. Posttranscriptional regulation of miRNAs harboring conserved terminal loops. Mol. Cell 32, 383–393 (2008).
Lünse, C.E. et al. An aptamer targeting the apical-loop domain modulates pri-miRNA processing. Angew. Chem. Int. Edn Engl. 49, 4674–4677 (2010).
Zeng, Y. & Cullen, B.R. Sequence requirements for micro RNA processing and function in human cells. RNA 9, 112–123 (2003).
Wolkenberg, S.E. & Boger, D.L. Mechanisms of in situ activation for DNA-targeting antitumor agents. Chem. Rev. 102, 2477–2495 (2002).
Kramer, R. & Cohen, D. Functional genomics to new drug targets. Nat. Rev. Drug Discov. 3, 965–972 (2004).
Bevilacqua, J.M. & Bevilacqua, P.C. Thermodynamic analysis of an RNA combinatorial library contained in a short hairpin. Biochemistry 37, 15877–15884 (1998).
McKenna, S.A. et al. Purification and characterization of transcribed RNAs using gel filtration chromatography. Nat. Protoc. 2, 3270–3277 (2007).
Peyret, N., Seneviratne, P.A., Allawi, H.T. & SantaLucia, J. Jr. Nearest-neighbor thermodynamics and NMR of DNA sequences with internal A.A, C.C, G.G, and T.T mismatches. Biochemistry 38, 3468–3477 (1999).
SantaLucia, J. Jr. A unified view of polymer, dumbbell, and oligonucleotide DNA nearest-neighbor thermodynamics. Proc. Natl. Acad. Sci. USA 95, 1460–1465 (1998).
Puglisi, J.D. & Tinoco, I. Jr. Absorbance melting curves of RNA. Methods Enzymol. 180, 304–325 (1989).
Wang, Y. & Rando, R.R. Specific binding of aminoglycoside antibiotics to RNA. Chem. Biol. 2, 281–290 (1995).
Disney, M.D., Gryaznov, S.M. & Turner, D.H. Contributions of individual nucleotides to tertiary binding of substrate by a Pneumocystis carinii group I intron. Biochemistry 39, 14269–14278 (2000).
Landthaler, M., Yalcin, A. & Tuschl, T. The human DiGeorge syndrome critical region gene 8 and its D. melanogaster homolog are required for miRNA biogenesis. Curr. Biol. 14, 2162–2167 (2004).
Acknowledgements
We thank B. White (Department of Cell Biology, University of Connecticut Health Center) for the luciferase plasmids containing FOXO1 3′ UTRs; C. Haga for assistance designing qRT-PCR primers; K. Lowe for assistance with flow cytometry; S. Seedhouse for preliminary studies with Inforna; B. Liu, J. Sokolow and T. Tran for compiling the secondary structures of the miRNA precursors and the RNA motif–small molecule database; and J. Childs-Disney, T. Kodadek, J. Cleveland, B. Roush, J. Joyce, M. Burkard and M. Guo for critical review of the manuscript. This work was funded by the US National Institutes of Health (R01GM097455). M.D.D. is a Camille and Henry Dreyfus Teacher-Scholar.
Author information
Authors and Affiliations
Contributions
S.P.V. designed and completed all of the experiments and contributed to the writing of the manuscript; S.M.G. programmed Inforna; and M.D.D. conceived Inforna, designed experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Various aspects of this technology have copyright protection or are part of a provisional patent application.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1 and 2, Supplementary Figures 1–16, and Supplementary Note. (PDF 9192 kb)
Rights and permissions
About this article
Cite this article
Velagapudi, S., Gallo, S. & Disney, M. Sequence-based design of bioactive small molecules that target precursor microRNAs. Nat Chem Biol 10, 291–297 (2014). https://doi.org/10.1038/nchembio.1452
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1452
This article is cited by
-
Small molecule regulators of microRNAs identified by high-throughput screen coupled with high-throughput sequencing
Nature Communications (2023)
-
Broad-spectrum metastasis suppressing compounds and therapeutic uses thereof in human tumors
Scientific Reports (2023)
-
Design of a small molecule that stimulates vascular endothelial growth factor A enabled by screening RNA fold–small molecule interactions
Nature Chemistry (2020)
-
Inhibition of RNA-binding proteins with small molecules
Nature Reviews Chemistry (2020)