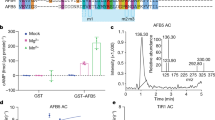
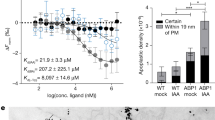
Abstract
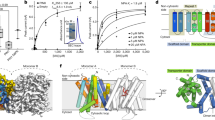
The plant hormone auxin regulates virtually every aspect of plant growth and development. Auxin acts by binding the F-box protein transport inhibitor response 1 (TIR1) and promotes the degradation of the AUXIN/INDOLE-3-ACETIC ACID (Aux/IAA) transcriptional repressors. Here we show that efficient auxin binding requires assembly of an auxin co-receptor complex consisting of TIR1 and an Aux/IAA protein. Heterologous experiments in yeast and quantitative IAA binding assays using purified proteins showed that different combinations of TIR1 and Aux/IAA proteins form co-receptor complexes with a wide range of auxin-binding affinities. Auxin affinity seems to be largely determined by the Aux/IAA. As there are 6 TIR1/AUXIN SIGNALING F-BOX proteins (AFBs) and 29 Aux/IAA proteins in Arabidopsis thaliana, combinatorial interactions may result in many co-receptors with distinct auxin-sensing properties. We also demonstrate that the AFB5–Aux/IAA co-receptor selectively binds the auxinic herbicide picloram. This co-receptor system broadens the effective concentration range of the hormone and may contribute to the complexity of auxin response.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Weijers, D., Ljung, K., Leyser, O. & Estelle, M. Auxin Signaling: From Synthesis to Systems Biology (ed. Estelle, M.) (Cold Spring Harbor Laboratory Press, 2011).
Hamann, T., Benkova, E., Baurle, I., Kientz, M. & Jurgens, G. The Arabidopsis BODENLOS gene encodes an auxin response protein inhibiting MONOPTEROS-mediated embryo patterning. Genes Dev. 16, 1610–1615 (2002).
Weijers, D. et al. Auxin triggers transient local signaling for cell specification in Arabidopsis embryogenesis. Dev. Cell 10, 265–270 (2006).
Holland, J.J., Roberts, D. & Liscum, E. Understanding phototropism: from Darwin to today. J. Exp. Bot. 60, 1969–1978 (2009).
Scarpella, E., Barkoulas, M. & Tsiantis, M. Control of leaf and vein development by auxin. Cold Spring Harb. Perspect. Biol. 2, a001511 (2010).
Vernoux, T., Besnard, F. & Traas, J. Auxin at the shoot apical meristem. Cold Spring Harb. Perspect. Biol. 2, a001487 (2010).
Overvoorde, P., Fukaki, H. & Beeckman, T. Auxin control of root development. Cold Spring Harb. Perspect. Biol. 2, a001537 (2010).
Sundberg, E. & Østergaard, L. Distinct and dynamic auxin activities during reproductive development. Cold Spring Harb. Perspect. Biol. 1, a001628 (2009).
Woodward, A.W. & Bartel, B. Auxin: regulation, action, and interaction. Ann. Bot. 95, 707–735 (2005).
Abel, S. & Theologis, A. Early genes and auxin action. Plant Physiol. 111, 9–17 (1996).
Overvoorde, P.J. et al. Functional genomic analysis of the AUXIN/INDOLE-3-ACETIC ACID gene family members in Arabidopsis thaliana. Plant Cell 17, 3282–3300 (2005).
Tiwari, S.B., Wang, X.J., Hagen, G. & Guilfoyle, T.J. AUX/IAA proteins are active repressors and their stability and activity are modulated by auxin. Plant Cell 13, 2809–2822 (2001).
Long, J.A., Ohno, C., Smith, Z.R. & Meyerowitz, E.M. TOPLESS regulates apical embryonic fate in Arabidopsis. Science 312, 1520–1523 (2006).
Szemenyei, H., Hannon, M. & Long, J.A. TOPLESS mediates auxin-dependent transcriptional repression during Arabidopsis embryogenesis. Science 319, 1384–1386 (2008).
Okushima, Y. et al. Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: unique and overlapping functions of ARF7 and ARF19. Plant Cell 17, 444–463 (2005).
Weijers, D. et al. Developmental specificity of auxin response by pairs of ARF and Aux/IAA transcriptional regulators. EMBO J. 24, 1874–1885 (2005).
Ulmasov, T., Murfett, J., Hagen, G. & Guilfoyle, T.J. Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 9, 1963–1971 (1997).
Ramos, J.A., Zenser, N., Leyser, O. & Callis, J. Rapid degradation of auxin/indoleacetic acid proteins requires conserved amino acids of domain II and is proteasome dependent. Plant Cell 13, 2349–2360 (2001).
Chapman, E.J. & Estelle, M. Mechanism of auxin-regulated gene expression in plants. Annu. Rev. Genet. 43, 265–285 (2009).
Dreher, K.A., Brown, J., Saw, R.E. & Callis, J. The Arabidopsis Aux/IAA protein family has diversified in degradation and auxin responsiveness. Plant Cell 18, 699–714 (2006).
Worley, C.K. et al. Degradation of Aux/IAA proteins is essential for normal auxin signalling. Plant J. 21, 553–562 (2000).
Dharmasiri, N. et al. Plant development is regulated by a family of auxin receptor F box proteins. Dev. Cell 9, 109–119 (2005).
Gray, W.M., Kepinski, S., Rouse, D., Leyser, O. & Estelle, M. Auxin regulates SCFTIR1-dependent degradation of AUX/IAA proteins. Nature 414, 271–276 (2001).
Parry, G. et al. Complex regulation of the TIR1/AFB family of auxin receptors. Proc. Natl. Acad. Sci. USA 106, 22540–22545 (2009).
Dharmasiri, N., Dharmasiri, S. & Estelle, M. The F-box protein TIR1 is an auxin receptor. Nature 435, 441–445 (2005).
Kepinski, S. & Leyser, O. The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 435, 446–451 (2005).
Calderón-Villalobos, L.I., Tan, X., Zheng, N. & Estelle, M. Auxin perception–structural insights. Cold Spring Harb. Perspect. Biol. 2, a005546 (2010).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Greenham, K. et al. The AFB4 auxin receptor is a negative regulator of auxin signaling in seedlings. Curr. Biol. 21, 520–525 (2011).
Vidal, E.A. et al. Nitrate-responsive miR393/AFB3 regulatory module controls root system architecture in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 107, 4477–4482 (2010).
Dharmasiri, N., Dharmasiri, S., Jones, A.M. & Estelle, M. Auxin action in a cell-free system. Curr. Biol. 13, 1418–1422 (2003).
Timpte, C., Wilson, A.K. & Estelle, M. The axr2–1 mutation of Arabidopsis thaliana is a gain-of-function mutation that disrupts an early step in auxin response. Genetics 138, 1239–1249 (1994).
Abel, S., Oeller, P.W. & Theologis, A. Early auxin-induced genes encode short-lived nuclear proteins. Proc. Natl. Acad. Sci. USA 91, 326–330 (1994).
Prigge, M.J., Lavy, M., Ashton, N.W. & Estelle, M. Physcomitrella patens auxin-resistant mutants affect conserved elements of an auxin-signaling pathway. Curr. Biol. 20, 1907–1912 (2010).
Vernoux, T. et al. The auxin signalling network translates dynamic input into robust patterning at the shoot apex. Mol. Syst. Biol. 7, 508 (2011).
Alonso, J.M. et al. Five components of the ethylene-response pathway identified in a screen for weak ethylene-insensitive mutants in Arabidopsis. Proc. Natl. Acad. Sci. USA 100, 2992–2997 (2003).
Ruegger, M. et al. The TIR protein of Arabidopsis function in auxin response and is related to human SKP2 and yeast Grr1p. Genes Dev. 12, 198–207 (1998).
Hayashi, K. et al. Small-molecule agonists and antagonists of F-box protein-substrate interactions in auxin perception and signaling. Proc. Natl. Acad. Sci. USA 105, 5632–5637 (2008).
Zhao, Y. & Hasenstein, K.H. Physiological interactions of antiauxins with auxin in roots. J. Plant Physiol. 167, 879–884 (2010).
Hao, G.F. & Yang, G.F. The role of Phe82 and Phe351 in auxin-induced substrate perception by TIR1 ubiquitin ligase: a novel insight from molecular dynamics simulations. PLoS ONE 5, e10742 (2010).
Walsh, T.A. et al. Mutations in an auxin receptor homolog AFB5 and in SGT1b confer resistance to synthetic picolinate auxins and not to 2,4-dichlorophenoxyacetic acid or indole-3-acetic acid in Arabidopsis. Plant Physiol. 142, 542–552 (2006).
Prusty, R., Grisafi, P. & Fink, G.R. The plant hormone indoleacetic acid induces invasive growth in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 101, 4153–4157 (2004).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Evans, M.L., Ishikawa, H. & Estelle, M. Responses of Arabidopsis roots to auxin studied with high temporal resolution—comparison of wild-type and auxin-response mutants. Planta 194, 215–222 (1994).
Jenik, P.D., Jurkuta, R.E. & Barton, M.K. Interactions between the cell cycle and embryonic patterning in Arabidopsis uncovered by a mutation in DNA polymerase epsilon. Plant Cell 17, 3362–3377 (2005).
Prime, V. 2.1. (Schrödinger, LLC, 2010).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. Procheck-A program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Sippl, M.J. Calculation of conformational ensembles from potentials of mean force. An approach to the knowledge-based prediction of local structures in globular proteins. J. Mol. Biol. 213, 859–883 (1990).
Molecular Operating Environment 2008.10. (Chemical Computing Group Inc., 2009).
Korb, O., Monecke, P., Hessler, G., Stutzle, T. & Exner, T.E. pharmACOphore: multiple flexible ligand alignment based on ant colony optimization. J. Chem. Inf. Model. 50, 1669–1681 (2010).
Acknowledgements
We thank E.J. Chapman for helpful discussions and comments to the manuscript and R. Shao and I. Kim for technical assistance. We also thank A. McCammon for hosting part of the computational analyses. We gratefully acknowledge financial support from the US National Institutes of Health (NIH) (R01 CA107134 to N.Z. and T32 GM07270 to L.B.S.), HHMI (M.E. and N.Z.) and the UK Biotechnology and Biological Sciences Research Council (BB/F013981/1 to S.K. and BB/F014651/1 to R.N.). The McCammon group, including A.I. and C.D.O., is supported by the NIH, US National Science Foundation and HHMI. We dedicate this work to the memory of L.B. Sheard, a talented young scientist who made key contributions at the early stage of this work. Her life was tragically cut short while this manuscript was under review.
Author information
Authors and Affiliations
Contributions
L.I.A.C.V. and M.E. prepared the manuscript. L.I.A.C.V. designed and performed the experiments and analyzed the data. X.T. and H.M. purified TIR1–ASK1 complex. L.I.A.C.V. expressed and purified AFB5–ASK1 complex. G.P. contributed to the generation of Y2H clones; C.D.O., A.I. and W.B. carried out homology modeling of AFB5 and docking experiments of picloram; and IAA. S.L., L.A., R.N. and S.K. expressed TIR1–ASK1 and AFB5–ASK1 constructs. S.L., S.K. and R.N. designed and performed SPR experiments. L.B.S. and N.Z. helped in the expression and purification of TIR1–ASK1 and AFB5–ASK1 complexes and the initial radioligand binding experiments. S.K., R.N., L.S. and N.Z. provided input to the manuscript. M.E. oversaw the project and approved the intellectual content.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 1157 kb)
Rights and permissions
About this article
Cite this article
Calderón Villalobos, L., Lee, S., De Oliveira, C. et al. A combinatorial TIR1/AFB–Aux/IAA co-receptor system for differential sensing of auxin. Nat Chem Biol 8, 477–485 (2012). https://doi.org/10.1038/nchembio.926
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.926
This article is cited by
-
Auxin and carbohydrate control flower bud development in Anthurium andraeanum during early stage of sexual reproduction
BMC Plant Biology (2024)
-
Overexpression of Pseudomonas putida-responsive miR393a-5p attunes morpho-physiological and biochemical parameters in Arabidopsis
Plant Cell, Tissue and Organ Culture (PCTOC) (2024)
-
Different evolutionary patterns of TIR1/AFBs and AUX/IAAs and their implications for the morphogenesis of land plants
BMC Plant Biology (2023)
-
Genome-wide identification and expression analysis of the AUX/IAA gene family in turnip (Brassica rapa ssp. rapa)
BMC Plant Biology (2023)
-
NLR surveillance of pathogen interference with hormone receptors induces immunity
Nature (2023)