Abstract
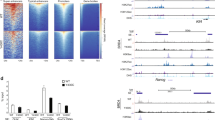
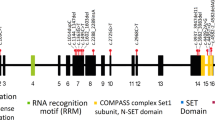
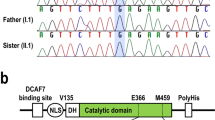
We show that haploinsufficiency of KANSL1 is sufficient to cause the 17q21.31 microdeletion syndrome, a multisystem disorder characterized by intellectual disability, hypotonia and distinctive facial features. The KANSL1 protein is an evolutionarily conserved regulator of the chromatin modifier KAT8, which influences gene expression through histone H4 lysine 16 (H4K16) acetylation. RNA sequencing studies in cell lines derived from affected individuals and the presence of learning deficits in Drosophila melanogaster mutants suggest a role for KANSL1 in neuronal processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Koolen, D.A. et al. Nat. Genet. 38, 999–1001 (2006).
Sharp, A.J. et al. Nat. Genet. 38, 1038–1042 (2006).
Shaw-Smith, C. et al. Nat. Genet. 38, 1032–1037 (2006).
Koolen, D.A. et al. J. Med. Genet. 45, 710–720 (2008).
Tan, T.Y. et al. J. Med. Genet. 46, 480–489 (2009).
Hutton, M. et al. Nature 393, 702–705 (1998).
Dubourg, C. et al. Eur. J. Med. Genet. 54, 144–151 (2011).
Cooper, G.M. et al. Nat. Genet. 43, 838–846 (2011).
Mendjan, S. et al. Mol. Cell 21, 811–823 (2006).
Li, X. et al. Mol. Cell 36, 290–301 (2009).
Dou, Y. et al. Cell 121, 873–885 (2005).
Ng, S.B. et al. Nat. Genet. 42, 790–793 (2010).
Raja, S.J. et al. Mol. Cell 38, 827–841 (2010).
Lone, M. et al. J. Cell Sci. 123, 2369–2374 (2010).
Acknowledgements
We thank the subjects and their parents for participation in this study and H. van Bokhoven for his comments on the manuscript. We thank M. Schepens, K. van de Donk and personnel from the Microarray Facility and Sequencing Facility Nijmegen for technical assistance. We thank K. Keleman (Institute of Molecular Pathology) and S. Sweeney (University of York) for Drosophila stocks. This study was financially supported by the Dutch Brain Foundation (KS 2012(1)-119 to D.A.K.), the Netherlands Organization for Health Research and Development (ZonMW; grants 917-66-363 and 911-08-025 to J.A.V., 916-86-016 to L.E.L.M.V., 917-96-346 to A.S. and 917-86-319 to B.B.A.d.V.), the European Union–funded TECHGENE project (Health-F5-2009-223143 to J.A.V. and H.S.), the AnEUploidy project (LSHG-CT-2006-37627 to H.G.B., J.A.V and B.B.A.d.V.) and the GENCODYS project (HEALTH-F4-2010-241995 to A.S. and B.B.A.d.V.). We obtained informed consent for participation in the study and to publish clinical photographs for individuals 1–4. The Medical Review Ethics Committee of the Radboud University Nijmegen Medical Center approved the study.
Author information
Authors and Affiliations
Contributions
D.A.K. and B.B.A.d.V. designed the study. K.N. performed expression profiling. W.M.N. designed and performed the genetics experiments. J.M.K. performed the experiments in Drosophila. C.G. and E.T.P.V. performed the bioinformatics experiments. S.M., P.d.V., L.E.L.M.V. and A.P.M.d.B. contributed to the expression profiling and genetics experiments. D.A.K., H.L.M.-B., F.V.E., A.T., J.A., V.M., A.C.-H.T., S.W.C., E.M.H.F.B., H.G.B. and B.B.A.d.V. recruited and evaluated the study subjects. T.F. performed statistical analysis. H.G.Y. supervised W.M.N. and S.M., J.A.V. and H.S. supervised K.N., C.G. and E.T.P.V., A.S. supervised J.M.K., and B.B.A.d.V. supervised D.A.K. D.A.K., J.M.K., K.N. and B.B.A.d.V. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Note, Supplementary Figure 1 and Supplementary Tables 1–4 and 8 (PDF 388 kb)
Supplementary Table 5
Differentially expressed genes in 17q21.31 microdeletion samples compared to controls. (XLSX 61 kb)
Supplementary Table 6
Differentially expressed genes in individual 4 compared to controls (XLSX 626 kb)
Supplementary Table 7
143 genes differentially expressed in both types of mutations (XLSX 35 kb)
Rights and permissions
About this article
Cite this article
Koolen, D., Kramer, J., Neveling, K. et al. Mutations in the chromatin modifier gene KANSL1 cause the 17q21.31 microdeletion syndrome. Nat Genet 44, 639–641 (2012). https://doi.org/10.1038/ng.2262
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2262
This article is cited by
-
A new blood DNA methylation signature for Koolen-de Vries syndrome: Classification of missense KANSL1 variants and comparison to fibroblast cells
European Journal of Human Genetics (2024)
-
Expanding the speech and language phenotype in Koolen-de Vries syndrome: late onset and periodic stuttering a novel feature
European Journal of Human Genetics (2023)
-
COX17 acetylation via MOF–KANSL complex promotes mitochondrial integrity and function
Nature Metabolism (2023)
-
H4K16ac activates the transcription of transposable elements and contributes to their cis-regulatory function
Nature Structural & Molecular Biology (2023)
-
GenIDA: an international participatory database to gain knowledge on health issues related to genetic forms of neurodevelopmental disorders
Journal of Neural Transmission (2023)