Abstract
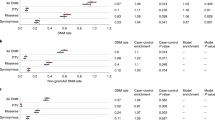
To evaluate evidence for de novo etiologies in schizophrenia, we sequenced at high coverage the exomes of families recruited from two populations with distinct demographic structures and history. We sequenced a total of 795 exomes from 231 parent-proband trios enriched for sporadic schizophrenia cases, as well as 34 unaffected trios. We observed in cases an excess of de novo nonsynonymous single-nucleotide variants as well as a higher prevalence of gene-disruptive de novo mutations relative to controls. We found four genes (LAMA2, DPYD, TRRAP and VPS39) affected by recurrent de novo events within or across the two populations, which is unlikely to have occurred by chance. We show that de novo mutations affect genes with diverse functions and developmental profiles, but we also find a substantial contribution of mutations in genes with higher expression in early fetal life. Our results help define the genomic and neural architecture of schizophrenia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Rodriguez-Murillo, L., Gogos, J.A. & Karayiorgou, M. The genetic architecture of schizophrenia: new mutations and emerging paradigms. Annu. Rev. Med. 63, 63–80 (2012).
Karayiorgou, M. et al. Schizophrenia susceptibility associated with interstitial deletions of chromosome 22q11. Proc. Natl. Acad. Sci. USA 92, 7612–7616 (1995).
Xu, B. et al. Strong association of de novo copy number mutations with sporadic schizophrenia. Nat. Genet. 40, 880–885 (2008).
Malhotra, D. et al. High frequencies of de novo CNVs in bipolar disorder and schizophrenia. Neuron 72, 951–963 (2011).
Kirov, G. et al. De novo CNV analysis implicates specific abnormalities of postsynaptic signalling complexes in the pathogenesis of schizophrenia. Mol. Psychiatry 17, 142–153 (2012).
Girard, S.L. et al. Increased exonic de novo mutation rate in individuals with schizophrenia. Nat. Genet. 43, 860–863 (2011).
Xu, B. et al. Exome sequencing supports a de novo mutational paradigm for schizophrenia. Nat. Genet. 43, 864–868 (2011).
Xu, B. et al. Elucidating the genetic architecture of familial schizophrenia using rare copy number variant and linkage scans. Proc. Natl. Acad. Sci. USA 106, 16746–16751 (2009).
Kryukov, G.V., Pennacchio, L.A. & Sunyaev, S.R. Most rare missense alleles are deleterious in humans: implications for complex disease and association studies. Am. J. Hum. Genet. 80, 727–739 (2007).
Neale, B.M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012).
Sanders, S.J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012).
O'Roak, B.J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012).
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012).
Kang, H.J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011).
Colantuoni, C. et al. Temporal dynamics and genetic control of transcription in the human prefrontal cortex. Nature 478, 519–523 (2011).
Barch, D.M. & Ceaser, A. Cognition in schizophrenia: core psychological and neural mechanisms. Trends Cogn. Sci. 16, 27–34 (2012).
Sobin, C., Roos, J.L., Pretorius, H., Lundy, L.S. & Karayiorgou, M. A comparison study of early non-psychotic deviant behavior in Afrikaner and US patients with schizophrenia or schizoaffective disorder. Psychiatry Res. 117, 113–125 (2003).
Christopherson, K.S. et al. Thrombospondins are astrocyte-secreted proteins that promote CNS synaptogenesis. Cell 120, 421–433 (2005).
Georges-Labouesse, E., Mark, M., Messaddeq, N. & Gansmuller, A. Essential role of a6 integrins in cortical and retinal lamination. Curr. Biol. 8, 983–986 (1998).
Jones, K.J. et al. The expanding phenotype of laminin a2 chain (merosin) abnormalities: case series and review. J. Med. Genet. 38, 649–657 (2001).
Van Kuilenburg, A.B. et al. Genotype and phenotype in patients with dihydropyrimidine dehydrogenase deficiency. Hum. Genet. 104, 1–9 (1999).
Tiedje, K.E., Stevens, K., Barnes, S. & Weaver, D.F. b-alanine as a small molecule neurotransmitter. Neurochem. Int. 57, 177–188 (2010).
Ben-David, E. et al. Identification of a functional rare variant in autism using genome-wide screen for monoallelic expression. Hum. Mol. Genet. 20, 3632–3641 (2011).
Carter, M.T. et al. Hemizygous deletions on chromosome 1p21.3 involving the DPYD gene in individuals with autism spectrum disorder. Clin. Genet. 80, 435–443 (2011).
Willemsen, M.H. et al. Chromosome 1p21.3 microdeletions comprising DPYD and MIR137 are associated with intellectual disability. J. Med. Genet. 48, 810–818 (2011).
Ripke, S. et al. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Arguello, P.A. & Gogos, J.A. Genetic and cognitive windows into circuit mechanisms of psychiatric disease. Trends Neurosci. 35, 3–13 (2012).
McGrath, J.J. & Susser, E.S. New directions in the epidemiology of schizophrenia. Med. J. Aust. 190, S7–S9 (2009).
Gilman, S.R. et al. Diverse types of genetic variation converge on functional gene networks involved in schizophrenia. Nat. Neurosci. (in the press).
Stark, K.L. et al. Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model. Nat. Genet. 40, 751–760 (2008).
Karayiorgou, M., Flint, J., Gogos, J.A. & Malenka, R.C. The best of times, the worst of times for psychiatric disease. Nat. Neurosci. 15, 811–812 (2012).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Grantham, R. Amino acid difference formula to help explain protein evolution. Science 185, 862–864 (1974).
Desmet, F.O. et al. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 37, e67 (2009).
Huang, D.W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Rossin, E.J. et al. Proteins encoded in genomic regions associated with immune-mediated disease physically interact and suggest underlying biology. PLoS Genet. 7, e1001273 (2011).
Acknowledgements
We are enormously grateful to all the families who participated in this research. We thank H. Pretorius and nursing sisters R. van Wyk, C. Botha and H. van den Berg for their assistance with subject recruitment, family history assessments and diagnostic evaluations. We thank S.L. Lundy for valuable assistance with clinical database maintenance and L. Rodriguez-Murillo for help with Supplementary Figure 9. We also thank B. Plummer and M. Robinson and the HudsonAlpha Genomics Services Laboratory for experimental support. Finally, we thank L.J. Mienie for the thymine loading test. This work was partially supported by National Institute of Mental Health (NIMH) grants MH061399 (to M.K.) and MH077235 (to J.A.G.) and the Lieber Center for Schizophrenia Research at Columbia University. B.X. was partially supported by a National Alliance for Research in Schizophrenia and Depression (NARSAD) Young Investigator Award.
Author information
Authors and Affiliations
Contributions
B.X., J.A.G. and M.K. designed the study, interpreted the data and prepared the manuscript. B.X. developed the analysis pipeline and had the primary role in the analysis and validation of sequence data. I.I.-L. performed statistical analysis of the sequence data. J.L.R. contributed to sample collection and clinical characterization. S.W. and Y.S. contributed to sample preparation and de novo mutation validation. B.B. performed exome library construction, capture and sequencing and initial analysis of SNV genotyping and indel variant calls. S.L. supervised the sequencing project at the HudsonAlpha Institute.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–9 and Supplementary Tables 1, 3–5, 7–9 and 11 (PDF 1422 kb)
Supplementary Table 2
De novo mutations identified in all three cohorts examined (XLS 63 kb)
Supplementary Table 6
List of prenatally-biased genes (XLS 33 kb)
Supplementary Table 10
Functional enrichment analysis of hsa-mir-367 and hsa-mir-1244 targets (XLS 32 kb)
Supplementary Table 12
HSF prediction results for splice site mutations in three genes with recurrent de novo mutations (XLS 25 kb)
Rights and permissions
About this article
Cite this article
Xu, B., Ionita-Laza, I., Roos, J. et al. De novo gene mutations highlight patterns of genetic and neural complexity in schizophrenia. Nat Genet 44, 1365–1369 (2012). https://doi.org/10.1038/ng.2446
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2446
This article is cited by
-
Association between urinary arsenic concentration and genetic polymorphisms in Korean adults
Toxicological Research (2024)
-
The molecular pathology of schizophrenia: an overview of existing knowledge and new directions for future research
Molecular Psychiatry (2023)
-
The genetic architecture of schizophrenia: review of large-scale genetic studies
Journal of Human Genetics (2023)
-
Impact of schizophrenia GWAS loci converge onto distinct pathways in cortical interneurons vs glutamatergic neurons during development
Molecular Psychiatry (2022)
-
Schizophrenia, autism spectrum disorders and developmental disorders share specific disruptive coding mutations
Nature Communications (2021)