Abstract
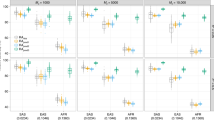
The genetic architecture of human diseases governs the success of genetic mapping and the future of personalized medicine. Although numerous studies have queried the genetic basis of common disease, contradictory hypotheses have been advocated about features of genetic architecture (for example, the contribution of rare versus common variants). We developed an integrated simulation framework, calibrated to empirical data, to enable the systematic evaluation of such hypotheses. For type 2 diabetes (T2D), two simple parameters—(i) the target size for causal mutation and (ii) the coupling between selection and phenotypic effect—define a broad space of architectures. Whereas extreme models are excluded by the combination of epidemiology, linkage and genome-wide association studies, many models remain consistent, including those where rare variants explain either little (<25%) or most (>80%) of T2D heritability. Ongoing sequencing and genotyping studies will further constrain the space of possible architectures, but very large samples (for example, >250,000 unselected individuals) will be required to localize most of the heritability underlying T2D and other traits characterized by these models.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Collins, F.S. & McKusick, V. Implications of the Human Genome Project for medical science. J. Am. Med. Assoc. 285, 540–544 (2001).
Jostins, L. & Barrett, J.C. Genetic risk prediction in complex disease. Hum. Mol. Genet. 20, R182–R188 (2011).
Thanassoulis, G. & Vasan, R. Genetic cardiovascular risk prediction—will we get there? Circulation 122, 2323–2334 (2010).
Grant, R.W., Moore, A.F. & Florez, J.C. Genetic architecture of type 2 diabetes: recent progress and clinical implications. Diabetes Care 32, 1107–1114 (2009).
Altshuler, D., Daly, M.J. & Lander, E.S. Genetic mapping in human disease. Science 322, 881–888 (2008).
Hirschhorn, J.N. & Daly, M.J. Genome-wide association studies for common diseases and complex traits. Nat. Rev. Genet. 6, 95–108 (2005).
McCarthy, M.I. et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat. Rev. Genet. 9, 356–369 (2008).
Cirulli, E.T. & Goldstein, D.B. Uncovering the roles of rare variants in common disease through whole-genome sequencing. Nat. Rev. Genet. 11, 415–425 (2010).
Kryukov, G.V., Shpunt, A., Stamatoyannopoulos, J.A. & Sunyaev, S.R. Power of deep, all-exon resequencing for discovery of human trait genes. Proc. Natl. Acad. Sci. USA 106, 3871–3876 (2009).
Price, A.L. et al. Pooled association tests for rare variants in exon-resequencing studies. Am. J. Hum. Genet. 86, 832–838 (2010).
Fisher, R.A. The genesis of twins. Genetics 4, 489–499 (1919).
Neale, M.C. & Maes, H.H.M. Methodology for Genetic Studies of Twins and Families (Kluwer Academic Publishers, Dordrecht, The Netherlands, 1992).
Martin, N., Boomsma, D.I. & Machin, G. A twin-pronged attack on complex traits. Nat. Genet. 17, 387–392 (1997).
Silventoinen, K. et al. Heritability of adult body height: a comparative study of twin cohorts in eight countries. Twin Res. 6, 399–408 (2003).
Lander, E.S. & Green, P. Construction of multilocus genetic linkage maps in humans. Proc. Natl. Acad. Sci. USA 84, 2363–2367 (1987).
Risch, N. Linkage strategies for genetically complex traits. II. The power of affected relative pairs. Am. J. Hum. Genet. 46, 229–241 (1990).
Botstein, D. & Risch, N. Discovering genotypes underlying human phenotypes: past successes for mendelian disease, future approaches for complex disease. Nat. Genet. 33, 228–237 (2003).
Purcell, S.M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Stahl, E.A. et al. Bayesian inference analyses of the polygenic architecture of rheumatoid arthritis. Nat. Genet. 44, 483–489 (2012).
Romeo, S. et al. Population-based resequencing of ANGPTL4 uncovers variations that reduce triglycerides and increase HDL. Nat. Genet. 39, 513–516 (2007).
Johansen, C.T. et al. Excess of rare variants in genes identified by genome-wide association study of hypertriglyceridemia. Nat. Genet. 42, 684–687 (2010).
Rivas, M.A. et al. Deep resequencing of GWAS loci identifies independent rare variants associated with inflammatory bowel disease. Nat. Genet. 43, 1066–1073 (2011).
Raychaudhuri, S. et al. A rare penetrant mutation in CFH confers high risk of age-related macular degeneration. Nat. Genet. 43, 1232–1236 (2011).
Bonnefond, A. et al. Rare MTNR1B variants impairing melatonin receptor 1B function contribute to type 2 diabetes. Nat. Genet. 44, 297–301 (2012).
Fearnhead, N.S. et al. Multiple rare variants in different genes account for multifactorial inherited susceptibility to colorectal adenomas. Proc. Natl. Acad. Sci. USA 101, 15992–15997 (2004).
Manolio, T.A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Eichler, E.E. et al. Missing heritability and strategies for finding the underlying causes of complex disease. Nat. Rev. Genet. 11, 446–450 (2010).
Maher, B. The case of the missing heritability. Nature 456, 18–21 (2008).
Lee, S.H. et al. Estimating the proportion of variation in susceptibility to schizophrenia captured by common SNPs. Nat. Genet. 44, 247–250 (2012).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
Coventry, A. et al. Deep resequencing reveals excess rare recent variants consistent with explosive population growth. Nat. Commun. 1, 131 (2010).
Li, Y. et al. Resequencing of 200 human exomes identifies an excess of low-frequency non-synonymous coding variants. Nat. Genet. 42, 969–972 (2010).
Keinan, A. & Clark, A.G. Recent explosive human population growth has resulted in an excess of rare genetic variants. Science 336, 740–743 (2012).
Nelson, M.R. et al. An abundance of rare functional variants in 202 drug target genes sequenced in 14,002 people. Science 337, 100–104 (2012).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Lupski, J.R., Belmont, J.W., Boerwinkle, E. & Gibbs, R.A. Clan genomics and the complex architecture of human disease. Cell 147, 32–43 (2011).
McClellan, J. & King, M.-C. Genetic heterogeneity in human disease. Cell 141, 210–217 (2010).
Mitchell, K.J. What is complex about complex disorders? Genome Biol. 13, 237 (2012).
Dickson, S.P., Wang, K., Krantz, I., Hakonarson, H. & Goldstein, D.B. Rare variants create synthetic genome-wide associations. PLoS Biol. 8, e1000294 (2010).
Lappalainen, T., Montgomery, S.B., Nica, A.C. & Dermitzakis, E.T. Epistatic selection between coding and regulatory variation in human evolution and disease. Am. J. Hum. Genet. 89, 459–463 (2011).
Zuk, O., Hechter, E., Sunyaev, S.R. & Lander, E.S. The mystery of missing heritability: genetic interactions create phantom heritability. Proc. Natl. Acad. Sci. USA 109, 1193–1198 (2012).
King, C.R., Rathouz, P.J. & Nicolae, D.L. An evolutionary framework for association testing in resequencing studies. PLoS Genet. 6, e1001202 (2010).
Browning, S.R. & Thompson, E.A. Detecting rare variant associations by identity-by-descent mapping in case-control studies. Genetics 190, 1521–1531 (2012).
Thornton, K.R., Foran, A.J. & Long, A.D. Properties and modeling of GWAS when complex disease risk is due to non-complementing, deleterious mutations in genes of large effect. PLoS Genet. 9, e1003258 (2013).
Reich, D.E. & Lander, E.S. On the allelic spectrum of human disease. Trends Genet. 17, 502–510 (2001).
Pritchard, J.K. & Cox, N.J. The allelic architecture of human disease genes: common disease–common variant.or not? Hum. Mol. Genet. 11, 2417–2423 (2002).
Eyre-Walker, A. Genetic architecture of a complex trait and its implications for fitness and genome-wide association studies. Proc. Natl. Acad. Sci. USA 107, 1752–1756 (2010).
Lambert, B.W., Terwilliger, J.D. & Weiss, K.M. ForSim: a tool for exploring the genetic architecture of complex traits with controlled truth. Bioinformatics 24, 1821–1822 (2008).
Gravel, S. et al. Demographic history and rare allele sharing among human populations. Proc. Natl. Acad. Sci. USA 108, 11983–11988 (2011).
Schaffner, S.F. et al. Calibrating a coalescent simulation of human genome sequence variation. Genome Res. 15, 1576–1583 (2005).
Ahituv, N. et al. Medical sequencing at the extremes of human body mass. Am. J. Hum. Genet. 80, 779–791 (2007).
Ward, L.D. & Kellis, M. Evidence of abundant purifying selection in humans for recently acquired regulatory functions. Science 337, 1675–1678 (2012).
Cowie, C.C., Rust, K., Byrd-Holt, D. & Gregg, E. Prevalence of diabetes and high risk for population in 1988–2006. Diabetes Care 33, 562–568 (2010).
Almgren, P. et al. Heritability and familiality of type 2 diabetes and related quantitative traits in the Botnia Study. Diabetologia 54, 2811–2819 (2011).
Zhu, Q. et al. A genome-wide comparison of the functional properties of rare and common genetic variants in humans. Am. J. Hum. Genet. 88, 458–468 (2011).
Montgomery, S.B., Lappalainen, T., Gutierrez-Arcelus, M. & Dermitzakis, E.T. Rare and common regulatory variation in population-scale sequenced human genomes. PLoS Genet. 7, e1002144 (2011).
Lindblad-Toh, K. et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature 478, 476–482 (2011).
Park, J.-H. et al. Distribution of allele frequencies and effect sizes and their interrelationships for common genetic susceptibility variants. Proc. Natl. Acad. Sci. USA 108, 18026–18031 (2011).
Lyssenko, V. et al. Predictors of and longitudinal changes in insulin sensitivity and secretion preceding onset of type 2 diabetes. Diabetes 54, 166–174 (2005).
Weijnen, C.F., Rich, S.S., Meigs, J.B., Krolewski, A.S. & Warram, J.H. Risk of diabetes in siblings of index cases with Type 2 diabetes: implications for genetic studies. Diabet. Med. 19, 41–50 (2002).
Guan, W., Pluzhnikov, A., Cox, N.J. & Boehnke, M. Meta-analysis of 23 type 2 diabetes linkage studies from the International Type 2 Diabetes Linkage Analysis Consortium. Hum. Hered. 66, 35–49 (2008).
Zeggini, E. et al. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat. Genet. 40, 638–645 (2008).
Voight, B.F. et al. The Metabochip, a custom genotyping array for genetic studies of metabolic, cardiovascular, and anthropometric traits. PLoS Genet. 8, e1002793 (2012).
Morris, A.P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Yang, J., Lee, S.H., Goddard, M.E. & Visscher, P.M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Popper, K.R. Conjectures and Refutations: The Growth of Scientific Knowledge (Routledge Classics, London, 2002).
Wang, K. et al. Interpretation of association signals and identification of causal variants from genome-wide association studies. Am. J. Hum. Genet. 86, 730–742 (2010).
Goldstein, D.B. The importance of synthetic associations will only be resolved empirically. PLoS Biol. 9, e1001008 (2011).
Guan, W., Boehnke, M., Pluzhnikov, A., Cox, N.J. & Scott, L.J. Identifying plausible genetic models based on association and linkage results: application to type 2 diabetes. Genet. Epidemiol. 9, 1–9 (2012).
Chakravarti, A. Population genetics—making sense out of sequence. Nat. Genet. 21 (suppl. 1), 56–60 (1999).
Zaitlen, N. et al. Using extended genealogy to estimate components of heritability for 23 quantitative and dichotomous traits. PLoS Genet. 9, e1003520 (2013).
Lee, S.H., Wray, N.R., Goddard, M.E. & Visscher, P.M. Estimating missing heritability for disease from genome-wide association studies. Am. J. Hum. Genet. 88, 294–305 (2011).
Acknowledgements
We gratefully acknowledge B. Lambert and K. Weiss (authors of the simulation tool ForSim) for helpful conversation, encouragement and technical assistance. Without their software, this work would not have been possible. We thank B. Voight for tremendous help in matching simulated genetic studies to those empirically conducted for T2D. We also thank E. Lander, C. Hartl, P. Fontanillas, B. Neale, M. McCarthy, M. Boehnke, M. Daly, S. Purcell and E. Stahl for discussion and insightful critiques. This work was supported by grants from the Doris Duke Charitable Foundation (award 2006087 to D.A.), the National Institute of General Medical Sciences (NIGMS; award R01GM078598 to S.S.) and the National Institute of Mental Health (NIMH; grant R01MH084676 to S.S.). V.A. is also supported by US National Institutes of Health (NIH) Training grants T32GM007753 and T32GM008313. J.F. is supported in part by NIH Training grant T32GM007748-33 as well as by funding from Pfizer. The GoT2D Study is supported by grant 1RC2DK088389-01 from the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK).
Author information
Authors and Affiliations
Consortia
Contributions
V.A., J.F., S.S. and D.A. conceived and designed the simulation framework. V.A., J.F. and S.S. fit population genetic parameters to match simulated and empirical data. V.A. performed the simulation studies. V.A. and J.F. analyzed simulation data under each disease model. V.A., J.F. and D.A. wrote the manuscript. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Full lists of members and affiliations appear in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–3, Supplementary Figures 1–24 and Supplementary Note (PDF 5708 kb)
Rights and permissions
About this article
Cite this article
Agarwala, V., Flannick, J., Sunyaev, S. et al. Evaluating empirical bounds on complex disease genetic architecture. Nat Genet 45, 1418–1427 (2013). https://doi.org/10.1038/ng.2804
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2804
This article is cited by
-
Weighted kernels improve multi-environment genomic prediction
Heredity (2023)
-
The genetics of non-monogenic IBD
Human Genetics (2023)
-
The genetics of autoimmune Addison disease: past, present and future
Nature Reviews Endocrinology (2022)
-
Human loss-of-function variants in the serotonin 2C receptor associated with obesity and maladaptive behavior
Nature Medicine (2022)
-
Clinical relevance of targeted exome sequencing in patients with rare syndromic short stature
Orphanet Journal of Rare Diseases (2021)