Abstract
A key component of genetic architecture is the allelic spectrum influencing trait variability. For autism spectrum disorder (herein termed autism), the nature of the allelic spectrum is uncertain. Individual risk-associated genes have been identified from rare variation, especially de novo mutations1,2,3,4,5,6,7,8. From this evidence, one might conclude that rare variation dominates the allelic spectrum in autism, yet recent studies show that common variation, individually of small effect, has substantial impact en masse9,10. At issue is how much of an impact relative to rare variation this common variation has. Using a unique epidemiological sample from Sweden, new methods that distinguish total narrow-sense heritability from that due to common variation and synthesis of results from other studies, we reach several conclusions about autism's genetic architecture: its narrow-sense heritability is ∼52.4%, with most due to common variation, and rare de novo mutations contribute substantially to individual liability, yet their contribution to variance in liability, 2.6%, is modest compared to that for heritable variation.
Similar content being viewed by others
Main
Autism is a neurodevelopmental disorder typified by striking deficits in social communication and genetically by a mixture of de novo and inherited variation contributing to liability. Rare variants clearly have a role in liability, with the contribution of de novo variation being the most obvious and easy to characterize, but inherited rare variation also has a role11,12. The contribution from inherited common variation is less substantiated. A handful of genome-wide association studies (GWAS) have been conducted in which significant findings have been few and specific to a single GWAS13,14,15. The results mirror those from the early GWAS analyses for schizophrenia, which, in retrospect, were underpowered, as evidenced by replicable associations involving common variants found in studies of tens of thousands of subjects10. In another parallel with early GWAS for schizophrenia, one of the first rays of hope for understanding how common variants affect liability came from the use of genetic scores, which were built from a large number of common variants and were shown to predict liability reliably16. For autism, scores also predict risk13. Some of the SNPs conferring risk for schizophrenia appear to also confer risk for autism17, a result that complements those for copy number variants (CNVs)18,19.
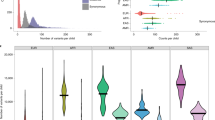
A natural complement to genetic scores from common variants is the estimation of narrow-sense heritability from the same variants. Two recent studies estimate heritability attributable to common variation to be substantial9,10, yet one estimates heritability at roughly 50% (Fig. 1) and the other estimates heritability at 17%. As described in the reports9,10, there are several technical reasons for these differences, one being quite different study designs and another being differences in ascertainment. As we shall show here, 50% seems more realistic.
(a) Heritability estimate (95% confidence interval) compared across study designs and analytical methods. The horizontal reference (dashed) is the PAGES estimate of heritability from SNP genotypes. Twin studies: (1) Californian twins for strictly defined autism (95% confidence interval = 8–84%), the largest twin study thus far using diagnosis only20; (2) Swedish twins of 9–12 years of age (95% confidence interval = 29–91%)32 and (3) Swedish twins of 9–12 years of age characterized for a quantitative measure of autism (most extreme cutoff; 95% confidence interval = 44–74%)33. SNP-based estimates of heritability: (4) Swedish family study (95% confidence interval = 44–64%)21; (5) simplex cases versus population controls (95% confidence interval = 26–73%)9 and (6) multiplex autism cases versus population controls (95% confidence interval = 38–93%)9. SNP-based estimates from the PAGES study, assuming prevalence (K) = 0.3%: (7) heritability due to common variants using autism cases versus population controls (95% confidence interval = 31–69%) and (8) total narrow-sense heritability due to both common and rare variation using smoothed estimates of relatedness (95% confidence interval = 35–71%). (b) Heritability per chromosome versus chromosome length. (c) Prevalence by county for all 21 counties in Sweden tallied by birth year cohort. Each box plot has a lower tail that extends from the minimum county-level prevalence to the 25th percentile; a central box that begins at the 25th percentile and ends at the 75th percentile, with a line demarcating the median prevalence; and an upper tail that extends from the 75th percentile to either (i) the maximum county-level prevalence (in the absence of any outliers) or (ii) a value of the 75th percentile + 1.5 times the vertical distance covered by the box—in this case, any outliers that exceed this end of the tail are noted by circular points on the plot. (d) PAGES heritability versus the population prevalence of autism for two estimators of heritability: case-control contrast using SNP genotypes (green) and total heritability from smoothed relationships among subjects, based on SNP genotypes (blue). Beyond the analysis of the PAGES study, we applied meta-analysis of selected h2 estimates (Online Methods) to obtain h2 = 51.4% (s.e.m. = 5.2%), which corresponds to a 95% confidence interval of 41.0—61.8%. Contrasting this with the comprehensive estimate of h2 obtained from the Swedish family study (h2 = 54%, s.e.m. = 5%) produces an estimate of h2 due to rare variants of 2.6% (s.e.m. = 7.2%, 95% confidence interval = 0–17%). Hence, we conclude that common variants explain the bulk of the heritability for autism, at least 41% of the variability, and rare variants explain at most 17%, on the basis of the upper and lower bounds of the respective 95% confidence intervals.
In any case, neither of these values approaches those from early twin studies, which place autism heritability close to 100% (see Supplementary Table 1 for data and discussion). Still, results from early studies could be compatible with estimates from common variants (Fig. 2). The key issue is that early twin studies of autism assumed that the genetic covariance of monozygotic twins was determined solely by additive effects and that non-additive and de novo effects on monozygotic similarity could be ignored. These are dubious assumptions, creating ample room for the discrepancy observed between study designs. In contrast, a recent large study of twins places heritability at 38% (ref. 20) (Figs. 1a and 2) under the same assumptions.
Variance in autism liability is determined by genetic and environmental factors. The genetic factors include additive effects (A), non-additive effects (D; dominant, recessive, epistatic) and de novo mutations (N). Environmental factors are split between common or shared environment (C) and stochastic or unique environment (E). (a) Early-autism twin studies estimate additive effects from the contrast of monozygotic (MZ) and dizygotic (DZ) correlations while assuming that non-additive effects and de novo mutations are zero. These are common assumptions for ACE (additive genetics, common environment, unique environment) heritability models but are unlikely to be appropriate for autism. (b) Applying the ACE model to the largest autism twin study thus far yields a lower estimate of additive heritability. (c) Heritability results using a more extensive set of family relationships and based on much of the population of Sweden. (d) Results from the PAGES study (see Fig. 1). (e) Contribution of the various factors to the variance in autism liability according to family relationship. De novo variation should not be shared in dizygotic twins, and, when it appears to be, it is almost surely inherited variation from a parent with gonadal mosaicism because the chance of the same mutation appearing de novo in the dizygotic twins is negligible. Most twin studies assume that common or shared environment is the same for monozygotic and dizygotic twins, although this approximation has been debated. Of note, the excess covariance of monozygotic twins relative to dizygotic twins is 1/2A + 3/4D + N as opposed to the 1/2A value assumed in the ACE model. Sibs, siblings. (f) Synthesis of results for the genetic architecture of autism (ASD).
To resolve this conundrum, we are evaluating a population sample by a variety of genetic analyses to estimate the relative contributions of rare, common, inherited and de novo variation to overall liability. We have ascertained subjects with strictly defined autism ('autistic disorder') from a Swedish epidemiological sample (Population-Based Autism Genetics and Environment Study, or PAGES).
Concurrently, a comprehensive study of autism in Sweden has been ongoing, and it recently reported the largest study of familial risk thus far21. This Swedish family study, a population-based cohort of all Swedish children born from 1982 to 2007 and a registry of all diagnoses before 2010, includes more than 1.6 million families with at least 2 children, yielding 5,799,875 cousin pairs, 2,642,064 full-sibling pairs, 432,281 maternal half-sibling pairs, 445,531 paternal half-sibling pairs and 37,570 twins. Of the 14,516 cases of broadly defined autism, 5,689 (39%) have a strict diagnosis. This massive homogeneous sample permits the precise estimation of relative recurrence risk for autism, given the diagnosis in relatives from monozygotic twins to first cousins and after modeling covariates such as sex, birth year, parental psychiatric history and parental age at birth. By analyzing these recurrence risk rates for additive and non-additive genetic effects and shared and non-shared environmental effects, the best model consists of only additive genetic and non-shared environmental effects and yields quite precise estimates of the narrow-sense heritability of autism (h2 = 54%, s.e.m. = 5%).
The Swedish family study provides a sound foundation from which to address other questions about the genetic architecture of autism using PAGES. There are no major differences in the population and samples underlying both studies. To estimate heritability for PAGES, controls were sampled from the Swedish population, and both cases and controls were genotyped on a common genotyping platform. After genotyping and quality control, we analyzed data from 531,906 SNPs characterized for 3,046 subjects, 466 with autism and 2,580 subjects10 not known to be affected.
We used the software package GCTA22 to estimate the heritability due to common variants, that is, SNP-based heritability. To ensure that all cases and controls were essentially unrelated (no pairs with kinship greater than 5th-degree relatives), 151 individuals were excluded. The resulting estimate of total variance in liability explained by measured SNPs was 49.4% (s.e.m. = 9.6%) (Fig. 1a). This heritability estimate compared remarkably well with findings based on independent data from population samples and similar methods9. The common variation imparting this heritability was distributed roughly uniformly across the chromosomes (Fig. 1b), an expectation of polygenic inheritance that is reflected in the significant correlation between heritability per chromosome and chromosome size (r = 0.49, P = 0.018). Prevalence of strictly defined autism, required to calculate heritability, was set to 0.3% (Fig. 1c and Supplementary Fig. 1) for these heritability calculations. The estimate is a lower bound for total narrow-sense heritability because it includes contributions from causal variants not tagged by the measured SNPs. Although synthetic association23—a pileup of rare risk variants in linkage disequilibrium with a common variant—could account for a small fraction of this heritability, this fraction cannot be large, as described below and previously24,25.
To obtain an estimate of the heritability due to both common and rare variation, we next included more closely related individuals. In a traditional analysis of heritability, for example, the Swedish family study, the relationship matrix is given. Instead, we estimated relationships from the SNPs genotyped for PAGES. Because estimates of relationship from SNP genotypes tend to be noisy, we used treelet covariance smoothing26 to improve estimates of pairwise relationship, especially for more distantly related individuals, and thereby refine estimates of heritability26. When we included relatives, albeit mostly distant (Supplementary Fig. 2), the estimated total heritability was 52.4% (s.e.m. = 9.5%). Although estimates were somewhat sensitive to prevalence, the differences between SNP-based heritability and heritabilities based on estimated relatedness were insensitive to prevalence (Fig. 1d): at any prevalence, the difference was approximately 3%. To evaluate how successful this approach would be at partitioning sources of heritability, we performed a simple simulation experiment that demonstrated that we could successfully partition heritability into the portions explained by common and rare variants (Online Methods).
Previous work has shown that autism heritability could fluctuate substantially when the bulk of the sample comprised simplex families, which lower heritability, versus multiplex families, which increase heritability9. These observations are consistent with liability being a classical quantitative genetic trait. Because PAGES is population based, it has no obvious simplex/multiplex ascertainment bias. Still there could be sources of more subtle bias. On the basis of the conjecture that subjects with intellectual disability and autism have a greater fraction of liability determined by de novo variants, one obvious bias would be misclassification of individuals who are comorbid for intellectual disability and autism as having one or the other disorder. To evaluate this issue, we first determined the diagnostic classification of Swedes, according to governmental records. In this population, 43.6% of subjects with autism also had intellectual disability, a rate comparable to those in other populations (note that strictly defined autism had a higher rate of comorbidity with intellectual disability than broadly defined autism). Next, to determine whether IQ has a substantial impact on heritability, we contrasted two estimates based on data from the Autism Genome Project (AGP): using the full sample, heritability equaled 51.1% (s.e.m. = 4.8%; n = 2,097), whereas, for subjects with IQ > 80, heritability was slightly but not significantly larger (59.3%, s.e.m. = 7.8%; n = 871). Subjects in the sample met broad criteria for an autism diagnosis; for the subset of individuals given a strict diagnosis, heritability was 52.3% (s.e.m. = 6.2%; n = 1,242).
Next, we asked how much of the variance in liability for autism could be explained by de novo mutations, applying the standard liability model to reported rates for de novo CNVs and loss-of-function mutations in autistic subjects and their siblings from the Simons Simplex Collection (SSC). In this sample, structured to enrich for de novo CNVs and loss-of-function mutations, the contribution of these variants to the variance in liability was 2.6% (Supplementary Tables 2 and 3, and Supplementary Note). Yet, de novo events can have a large impact on liability, and 14% of subjects carried such mutations: roughly 80% of subjects who were carriers of a de novo CNV would not be affected if they were not carriers; likewise, for carriers of de novo loss-of-function mutations, 57% would not be affected (Supplementary Note).
The estimate of heritability could indirectly include dominant or non-additive effects but should not include the impact of recessive inheritance. A recent study estimated the contribution from rare, recessive variation to be about 3% (ref. 27), a contribution similar to that from the additive effects of rare variants. Rare hemizygous loss-of-function mutations accounted for another 2% of liability.
We conclude that inherited rare variation explains a smaller fraction of total heritability than common variation (Fig. 2). Although uncertainty is inherent in all of these estimates (Fig. 1), the results converge on a total heritability in the range of 50–60%, with common variants explaining the bulk of it. Our analyses illustrate an approach to identify the contribution of rare and common variation to the heritability of any phenotype28. Estimating the total contribution of genetic variation to variation in liability, which includes non-additive effects and de novo variation, is more challenging. If the only non-additive effects of genes were due solely to recessive inheritance, roughly 5% would be added to the total, but that estimate could be low on the basis of both theoretical and empirical grounds29,30. And, although 14% of affected subjects carry de novo CNV and loss-of-function mutations, the contribution of these mutations to the variance in liability is only 2.6%. Summing these estimates suggests that genetic variation accounts for roughly 60% of the variation in risk for autism in Sweden, implying that the majority of risk is due to genetic variation.
By contrast, a recent twin study found that shared twin environment accounts for the majority of the variation in risk, 55%, on the basis of a population sample of Californians from the United States. These different populations could have different genetic architecture or there could be an unknown sampling bias. Alternatively, the California study fits many parameters to a relatively small data set—concordance rates on 54 monozygotic pairs and 138 dizygotic pairs—from which the study selects the best model on the basis of statistical criteria. For small samples, however, the correct model, the one truly generating the data, can be quite different in structure from the selected model, and yet the two can have only small differences in likelihood. It is possible that the different conclusion of the California study in comparison to others of its design is due to a modest stochastic difference that altered model selection. In this regard, a cautionary note for all such studies, including ours, is worthwhile: although we assume here a simple model structure, ours is but one of many possible models that could underlie trait covariance (for example, see ref. 31). The assumed model can alter inference, sometimes substantially, and many of these models can fit the data almost equally well. Nonetheless, the finding that all Swedish studies, regardless of design, converge on similar estimates of heritability lends strong support for our conclusion that the bulk of risk for autism arises from genetic variation.
Methods
Ascertainment of subjects.
We developed an epidemiological sample of autism or, more precisely, autistic disorder, taking advantage of the detailed birth and medical registries and universal access to healthcare in Sweden. Our sample frame was the medical birth register including all births in Sweden, where there is mandatory screening of all children at age 4 years for neurodevelopmental disorders. The medical registries included all individuals diagnosed with autistic disorder at any time. Cases with autistic disorder (International Statistical Classification of Diseases (ICD)-9 code 299A or ICD-10 codes F84.0–F84.1), henceforth termed autism, were identified from the Swedish National Patient Register (NPR). Controls free from schizophrenia and bipolar disease were recruited from the general Swedish population and matched by county, sex and birth year. Prevalence was 30 cases per 10,000 individuals for autism and approximately 100 cases per 10,000 individuals for the more inclusive, broadly defined autism diagnosis (Supplementary Fig. 1). Inclusion criteria were a diagnosis of autism in NPR; birth in Sweden; parents who were both born in a Nordic country; age of 10–65 years; and signed consent by a parent or a legal guardian (or by the subject, when possible and appropriate). Exclusion criteria were a diagnosis of autism but the presence of a genetic disorder also known to be associated with autistic features (for example, Fragile X, Down and Klinefelter syndromes) and medical or psychiatric history that could mitigate confident diagnosis with autism. In this way, 536 subjects with autism were recruited from 12 counties in Sweden. This study has been reviewed and approved by institutional review boards at the Karolinska Institutet, Icahn School of Medicine at Mount Sinai and Carnegie Mellon University.
Genetic characterization.
Samples were genotyped on the Illumina HumanOmniExpressExome BeadChip. Here we analyzed only the OmniExpress content of >715,000 SNPs across the genome. Duplicate samples and samples with genotype completion rates of <98% were removed, resulting in a final sample of 3,046 individuals, of whom 466 were autism cases and 2,580 were controls. We controlled for more subtle population structure using 7 significant dimensions of ancestry (P < 0.05) as covariates in all subsequent analyses (n = 3,044, omitting 1 individual from each set of twins).
Heritability.
To estimate heritability, the Swedish family study relied on an extended sibling design, which included full siblings, half-siblings, cousins and twins. The design facilitated the estimation of additive and non-additive genetic sources of variance, as well as shared and non-shared environmental sources of variance.
For all genetic analyses of heritability, SNPs with minor allele frequency (MAF) of >0.05 were evaluated using the program GCTA22 to produce an estimated genetic relationship matrix (GRM). As described further in the Supplementary Note, we then modeled case-control status via the mixed linear model y = Xβ + g + e, where y is the vector of case-control status, β is the vector of coefficients for the fixed effects (seven ancestry dimensions) with associated design matrix X, g is the vector of random additive genetic effects associated with SNPs and e is a vector of random errors, which were assumed to be independent. To obtain estimates of heritability, variance in phenotype was expressed as  (where A is the GRM, I is an identity matrix and
(where A is the GRM, I is an identity matrix and  and
and  partition the total phenotypic variation into pieces attributable to additive genetic effects and random error, respectively), and heritability was calculated as
partition the total phenotypic variation into pieces attributable to additive genetic effects and random error, respectively), and heritability was calculated as

on the observed scale, which was transformed to the liability scale as a function of the population prevalence (K).
To estimate the heritability due to common SNPs, we used the genetic analysis software GCTA to calculate a GRM and then obtain a heritability estimate based on a sample of essentially unrelated individuals (A < 0.025). To estimate the total narrow-sense heritability, we included all sampled individuals, computed the GRM, smoothed this matrix using Treelet Covariance smoothing26 (TCS) and then computed heritability from the GCTA package. See the Supplementary Note for details on the implementation of TCS. We used simulations to assess the accuracy of this procedure of estimating heritability (see the Supplementary Note for complete details). We started with the phased genomes (haplotypes) of individuals from the HapMap 3 database, selecting two populations of European ancestry (CEU and TSI), and, using the available haplotypes, we generated a large sample of haplotypes, representative of those that might be sampled from the unrelated founders of a population. After generating haplotype pairs, we randomly assigned chromosomes to founders in each of 100 families, and the founder chromosomes were dropped through a 5-generation pedigree. We combined 100 sets of independent pedigrees, including 20 individuals sampled per pedigree, to generate the full genotype sample of size 2,000. For the given set of genotypes, 50 independent vectors of phenotypes were simulated. For each simulation, a random set of causal variants was chosen: 1,000 rare (MAF < 0.01) and 1,000 common variants. These two classes of SNPs generated 25% and 50% of the heritability (h2), respectively, for total h2 = 75%. Using GCTA to estimate h2 solely from common variant genotypes—after removing relatives—we obtained mean h2 = 50.7% (s.e.m. = 3.5%). The finding that this estimate is close to the simulated value for common variants suggests that the impact of synthetic association is minimal. Next, applying GCTA to genotypes from common variants and the full sample, including relatives, produced mean h2 = 72.4% (s.e.m. = 1.2%) with TCS and mean h2 = 70.6% (s.e.m. = 1.2%) without TCS. Both captured most h2 due to rare variation.
Impact of clinical features on estimates of heritability, exemplified by diagnosis and intellectual function.
Consistent with quantitative genetics theory, it has already been shown that families who are multiplex for autism carry a larger load of liability-associated alleles than simplex families (defined as families with only one affected subject within the set of first- and second-degree relatives). Clinical phenotypes could also affect heritability/genetic load, although how much impact they might have is an open question. To address this question, we evaluated two phenotypes thought to have major effects on the genetics of autism, namely, diagnosis per se and higher versus lower functioning, as measured by IQ. First, by linking registry data from Sweden, we obtained an estimate of the fraction of subjects with autism and intellectual disability (IQ < 70) to determine its comparability with values in other population samples. To assess the impact of diagnosis, we used AGP data and followed up the AGP analysis by examining strict autism diagnosis, as defined by the meeting of criteria for autism on the Autism Diagnostic Interview-Revised and Autism Diagnostic Observation Schedule, versus broadly defined autism, which includes autism disorder and subjects who meet looser criteria for a spectrum diagnosis (see the Supplementary Note). For IQ, we targeted subjects with IQ ≥ 80, beyond the bound for intellectual disability. After quality control, there were 2,097 AGP cases13 and 1,663 Health, Aging and Body Composition controls9 genotyped for 828,352 markers. After analysis using GCTA, we observed heritabilities of 51.1 ± 4.8%, 52.3 ± 6.2% and 59.3 ± 7.8% for broadly defined autism, strictly defined autism and autism with IQ ≥ 80, respectively.
Meta-analysis of heritability.
A meta estimate of h2 due to common variants could be derived by taking a weighted average of two estimates of this quantity obtained from two independent samples: the PAGES study (h2 = 49.4%, s.e.m. = 9.5%) and the other 1,242 strictly defined autism subjects from AGP data (h2 = 52.3%, s.e.m. = 6.2%). We did not use the estimate based on the SSC sample (provided in Fig. 1) because SSC ascertainment of only simplex families introduces a negative bias on the estimate. Meta-analysis produced h2 = 51.4% (s.e.m. = 5.2%) and the corresponding 95% confidence interval of 41.0–61.8%. Contrasting this value with the total h2 value obtained from the Swedish family study (h2 = 54%, s.e.m. = 5%) produced an estimate of h2 due to rare variants  = 0.2.6% (s.e.m. = 7.2%, 95% confidence interval = 0–17%).
= 0.2.6% (s.e.m. = 7.2%, 95% confidence interval = 0–17%).
Estimating the contribution of de novo mutations and heritable variation to liability and variation in liability for autism.
For computational methods, see the Supplementary Note. To estimate the variance in liability explained by de novo variation, results from the SSC sample were analyzed, contrasting the rates of de novo CNVs, loss-of-function mutations and missense mutations. All three classes of variation have been shown to be significantly in excess in probands with autism relative to their unaffected siblings, although not all studies found de novo missense variation to be in excess1,2,3,4,5,7,34. For inference, we assumed that the excess proportion of cases carrying de novo mutations, relative to control siblings, is the fraction of de novo mutations that conferred liability.
De novo copy number variants. As described further in the Supplementary Note, 75 de novo CNVs were found in 858 probands and 19 de novo CNVs were found in 863 sibling controls34 (relative risk = 4.25). Assuming an 'exposure rate' of 0.022 (= 19/863), the classical liability model determined that de novo CNVs accounted for 1.46% of variability in the liability scale.
De novo loss-of-function mutations. Of the 599 probands with autism, 72 had a de novo loss-of-function mutation in comparison to 32 of the 599 sibling controls1,4,35 (relative risk = 2.42). With an exposure rate of 0.053, de novo loss-of-function mutations accounted for 1.11% of the variance in liability.
De novo missense mutations. Of the 599 probands, 253 had at least 1 de novo missense mutation in comparison to 238 of 599 sibling controls (relative risk = 1.11). With an exposure rate of 0.397, de novo missense mutations accounted for negligible variance in liability (0.04%).
Accession codes.
Data used in the preparation of this article reside in the US National Institutes of Health (NIH)-supported National Database for Autism Research (NDAR) under NDAR study 346.
References
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012).
Neale, B.M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012).
O'Roak, B.J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012).
Sanders, S.J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012).
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science 316, 445–449 (2007).
Glessner, J.T. et al. Autism genome-wide copy number variation reveals ubiquitin and neuronal genes. Nature 459, 569–573 (2009).
Sanders, S.J. et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron 70, 863–885 (2011).
Pinto, D. et al. Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Am. J. Hum. Genet. 94, 677–694 (2014).
Klei, L. et al. Common genetic variants, acting additively, are a major source of risk for autism. Mol. Autism 3, 9 (2012).
Lee, S.H. et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013).
State, M.W. & Levitt, P. The conundrums of understanding genetic risks for autism spectrum disorders. Nat. Neurosci. 14, 1499–1506 (2011).
Devlin, B. & Scherer, S.W. Genetic architecture in autism spectrum disorder. Curr. Opin. Genet. Dev. 22, 229–237 (2012).
Anney, R. et al. Individual common variants exert weak effects on the risk for autism spectrum disorderspi. Hum. Mol. Genet. 21, 4781–4792 (2012).
Wang, K. et al. Common genetic variants on 5p14.1 associate with autism spectrum disorders. Nature 459, 528–533 (2009).
Weiss, L.A. et al. A genome-wide linkage and association scan reveals novel loci for autism. Nature 461, 802–808 (2009).
International Schizophrenia Consortium. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Cross-Disorder Group of the Psychiatric Genomics Consortium. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013).
Stefansson, H. et al. CNVs conferring risk of autism or schizophrenia affect cognition in controls. Nature 505, 361–366 (2014).
Talkowski, M.E. et al. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell 149, 525–537 (2012).
Hallmayer, J. et al. Genetic heritability and shared environmental factors among twin pairs with autism. Arch. Gen. Psychiatry 68, 1095–1102 (2011).
Sandin, S. et al. The familial risk of autism. J. Am. Med. Assoc. (in the press) (2014).
Yang, J., Lee, S.H., Goddard, M.E. & Visscher, P.M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Dickson, S.P., Wang, K., Krantz, I., Hakonarson, H. & Goldstein, D.B. Rare variants create synthetic genome-wide associations. PLoS Biol. 8, e1000294 (2010).
Orozco, G., Barrett, J.C. & Zeggini, E. Synthetic associations in the context of genome-wide association scan signals. Hum. Mol. Genet. 19, R137–R144 (2010).
Wray, N.R., Purcell, S.M. & Visscher, P.M. Synthetic associations created by rare variants do not explain most GWAS results. PLoS Biol. 9, e1000579 (2011).
Crossett, A., Lee, A.B., Klei, L., Devlin, B. & Roeder, K. Refining genetically inferred relationships using treelet covariance smoothing. Ann. Appl. Stat. 7, 669–690 (2013).
Lim, E.T. et al. Rare complete knockouts in humans: population distribution and significant role in autism spectrum disorders. Neuron 77, 235–242 (2013).
Agarwala, V., Flannick, J., Sunyaev, S. & Altshuler, D. Evaluating empirical bounds on complex disease genetic architecture. Nat. Genet. 45, 1418–1427 (2013).
Zuk, O., Hechter, E., Sunyaev, S.R. & Lander, E.S. The mystery of missing heritability: genetic interactions create phantom heritability. Proc. Natl. Acad. Sci. USA 109, 1193–1198 (2012).
Lynch, M. & Walsh, B. Genetics and Analysis of Quantitative Traits (Sinauer Associates, Sunderland, MA, 1998).
Devlin, B., Daniels, M. & Roeder, K. The heritability of IQ. Nature 388, 468–471 (1997).
Lichtenstein, P., Carlstrom, E., Rastam, M., Gillberg, C. & Anckarsater, H. The genetics of autism spectrum disorders and related neuropsychiatric disorders in childhood. Am. J. Psychiatry 167, 1357–1363 (2010).
Lundström, S. et al. Autism spectrum disorders and autistic like traits: similar etiology in the extreme end and the normal variation. Arch. Gen. Psychiatry 69, 46–52 (2012).
Levy, D. et al. Rare de novo and transmitted copy-number variation in autistic spectrum disorders. Neuron 70, 886–897 (2011).
Willsey, A.J. et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell 155, 997–1007 (2013).
Acknowledgements
This study was supported by National Institute of Mental Health (NIMH) grants MH057881 and MH097849 and also in part through the computational resources and staff expertise provided by the Scientific Computing Facility at the Icahn School of Medicine at Mount Sinai. We thank the Mount Sinai Genomics Core Facility for carrying out Illumina bead array genotyping. We thank D. Cutler, M. Daly and S. Purcell for comments on the manuscript and M. Daly and P. Sullivan for facilitating access to control samples, collected and genotyped by M. Daly, C.M.H., P.S. and P. Sullivan with support from NIMH grants MH095034 and MH077139. We also thank the nurses, A.-K. Sundberg and A.-B. Holmgren, for their hard work in collecting the samples. AGP: We used data from the Autism Genome Project (AGP) Consortium Whole-Genome Association and Copy Number Variation Study of over 2,600 parent-offspring trios (database of Genotypes and Phenotypes (dbGaP) study phs000267.v4.p2). Funding for AGP was provided from the US NIH (HD055751, HD055782, HD055784, HD35465, MH52708, MH55284, MH57881, MH061009, MH06359, MH066673, MH080647, MH081754, MH66766, NS026630, NS042165 and NS049261); the Canadian Institutes for Health Research (CIHR); Assistance Publique–Hôpitaux de Paris, France; Autism Speaks, UK; the Canada Foundation for Innovation/Ontario Innovation Trust; grant Po 255/17-4 from Deutsche Forschungsgemeinschaft, Germany; the European Community's Sixth Framework Programme AUTISM MOLGEN; Fundação Calouste Gulbenkian, Portugal; Fondation de France; Fondation FondaMental, France; Fondation Orange, France; Fondation pour la Recherche Médicale, France; Fundação para a Ciência e Tecnologia, Portugal; The Hospital for Sick Children Foundation and the University of Toronto, Canada; INSERM, France; Institut Pasteur, France; Convention 181 of 19.10.2001 from the Italian Ministry of Health; the John P. Hussman Foundation, USA; the McLaughlin Centre, Canada; Rubicon 825.06.031 from the Netherlands Organization for Scientific Research; TMF/DA/5801 from the Royal Netherlands Academy of Arts and Sciences; the Ontario Ministry of Research and Innovation, Canada; the Seaver Foundation, USA; the Swedish Science Council; the Centre for Applied Genomics, Canada; the Utah Autism Foundation, USA; and Core award 075491/Z/04 from the Wellcome Trust, UK. Genotype and phenotype data were obtained from dbGaP, as provided by AGP study investigators. HealthABC: These controls were obtained from dbGaP. Funding support for the CIDR Visceral Adiposity Study (dbGaP study phs000169.v1.p1) was provided through the Division of Aging Biology and the Division of Geriatrics and Clinical Gerontology, National Institute on Aging. The CIDR Visceral Adiposity Study includes a GWAS funded as part of the Division of Aging Biology and the Division of Geriatrics and Clinical Gerontology, National Institute on Aging. Assistance with phenotype harmonization and genotype cleaning, as well as with general study coordination, was provided by Health ABC study investigators. This manuscript reflects the views of the authors and does not necessarily reflect the opinions or views of the US NIH.
Author information
Authors and Affiliations
Contributions
A.R., C.M.H., B.D., K.R. and J.D.B. conceived the project and designed its components. A.R., C.M.H., B.D., K.R. and J.D.B. identified funding for the study. S.S., O.S. and C.M.H. were responsible for ascertaining case samples, and P.S. and C.M.H. were responsible for ascertaining control samples. J.R., D.M. and M.M. were responsible for genotyping the case samples. A.P.G. organized and managed the data files, and T.G. and A.P.G. carried out quality control for the SNP data. S.J.S. carried out simulations and additional analyses to assess the contribution of rare variation to variance in liability, and S.R. carried out imputation to 1000 Genomes Project data. T.G., C.A.B. and A.B.L. carried out statistical analyses under the guidance of B.D., L.K. and K.R., and Y.P., S.S., O.S., A.R. and C.M.H. carried out epidemiological analyses. B.D., K.R. and J.D.B. took the lead in writing the manuscript, and all authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Estimated prevalence of strict autism for the entire Swedish population as a function of individuals’ ages.
Prevalence was calculated by Kaplan-Meier function utilizing year of birth, age at autistic disorder diagnosis and censoring due to death, emigration or end of study period, whichever came first. (a) Prevalence pooled across cohorts and counties. (b) Prevalence by cohort.
Supplementary Figure 2 Distribution of relative pairs by degree of relatedness estimated using TCS.
On the basis of 3,044 subjects, there are 4,631,446 subject pairs, and 63% of the pairs are estimated to be unrelated (degree > 10). Of these pairs, 1,346 were estimated to be degree 6 or closer, with 12, 37, 104 and 1,193 of the pairs falling into the intervals 2–3, 3–4, 4–5 and 5–6, respectively.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–3 and Supplementary Figures 1 and 2. (PDF 40004 kb)
Rights and permissions
About this article
Cite this article
Gaugler, T., Klei, L., Sanders, S. et al. Most genetic risk for autism resides with common variation. Nat Genet 46, 881–885 (2014). https://doi.org/10.1038/ng.3039
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3039
This article is cited by
-
A cohort study of neurodevelopmental disorders and/or congenital anomalies using high resolution chromosomal microarrays in southern Brazil highlighting the significance of ASD
Scientific Reports (2024)
-
Marked neuropsychiatric involvement and dysmorphic features in nemaline myopathy
Neurological Sciences (2024)
-
Implications of Genetic Factors and Modifiers in Autism Spectrum Disorders: a Systematic Review
Review Journal of Autism and Developmental Disorders (2024)
-
Assessment of vulnerability dimensions considering Family History and environmental interplay in Autism Spectrum Disorder
BMC Psychiatry (2023)
-
Endophenotype trait domains for advancing gene discovery in autism spectrum disorder
Journal of Neurodevelopmental Disorders (2023)